FEDERAL COURT OF AUSTRALIA
Meat & Livestock Australia Limited v Cargill, Inc [2018] FCA 51
ORDERS
MEAT & LIVESTOCK AUSTRALIA LIMITED (ACN 081 678 364) First Appellant DAIRY AUSTRALIA LIMITED (ACN 105 227 987) Second Appellant | ||
AND: | First Respondent BRANHAVEN LLC Second Respondent | |
DATE OF ORDER: |
THE COURT ORDERS THAT:
1. Within 14 days of the date of these orders, each of the parties file and serve proposed minutes of orders and short submissions (limited to three pages) to give effect to these reasons, including on the question of any steps necessary to deal with any application to amend the claims of patent application no. 2010202253 and on the question of costs.
2. Costs reserved.
Note: Entry of orders is dealt with in Rule 39.32 of the Federal Court Rules 2011.
BEACH J:
1 The first appellant, Meat & Livestock Australia Limited, invests in research relevant to the Australian meat and livestock industry. It was established on 18 February 1998 as a declared industry marketing and research body under the Australian Meat and Live-stock Industry Act 1997 (Cth). The second appellant, Dairy Australia Limited, also an Australian company, invests in research relevant to the Australian dairy industry. For convenience, I will refer to the appellants as MLA.
2 The first respondent, Cargill, Inc. (Cargill), is a US corporation, one of whose major businesses is the raising of livestock. The second respondent, Branhaven LLC (Branhaven), is also a US corporation. Branhaven and Cargill are co-applicants of Australian patent application number 2010202253 (the 253 Application) titled “Compositions, methods and systems for inferring bovine traits”. The principal claims of the 253 Application involve method claims for identifying a trait of a bovine subject from a nucleic acid sample of that subject. The field of the invention relates generally to gene association analyses, specifically to single nucleotide polymorphisms and correlated traits of bovine species. The scientific disciplines that are relevant are molecular genetics and quantitative genetics, on which I will say a little more later.
3 The 253 Application was filed on 1 June 2010 as a divisional application of the parent Australian patent application number 2003303599 filed on 31 December 2003, which has now been withdrawn. The parent application claimed a priority date of 31 December 2002 based upon US application number 60/437,482. Accordingly, the co-applicants of the 253 Application assert that the earlier priority date of 31 December 2002 applies.
4 In proceedings before the Patent Office, MLA opposed the grant of the 253 Application. Subject to one matter, that opposition was unsuccessful before the delegate of the Commissioner of Patents. The proceeding before me is an appeal by MLA from the decision of the delegate made on 6 May 2016; only Branhaven has actively resisted the present appeal. The delegate decided that the opposition brought by MLA failed on all grounds, save for one ground of lack of clarity and an aspect of a “manner of manufacture” concern affecting one claim of the 253 Application (Meat & Livestock Australia Limited and Dairy Australia Limited v Cargill, Inc. and Branhaven LLC [2016] APO 26).
5 I would note one matter at this point. The 253 Application is to be considered pursuant to the provisions of the Patents Act 1990 (Cth) (the Act) prior to the amendments made by the Intellectual Property Laws Amendment (Raising the Bar) Act 2012 (Cth), as I will explain later. The appeal is pursuant to s 60(4) of the Act, but the amendment concerning s 60(3A) does not apply.
6 The appeal is a hearing de novo on the grounds advanced and evidence adduced before me (Commissioner of Patents v Sherman (2008) 172 FCR 394 at [18] to [21] per Heerey, Kenny and Middleton JJ). Evidence before the delegate is not able to be adduced before me without leave. Further, as this is a complete re-hearing, findings made by the delegate have little separate status although I am entitled to take them into account (as I have done) given the delegate’s significant technical expertise. In this case, the delegate was Dr Lexie Press, who has worked as a molecular geneticist with the CSIRO and Stanford University prior to joining IP Australia. But I should note that I have not given her findings substantial weight given that most of the expert evidence adduced before me had not been put before her.
7 The following principles, as synthesised by Moshinsky J in Merial Inc v Intervet International BV (No 3) (2017) 122 IPR 128; [2017] FCA 21 at [11] to [16], apply to the present appeal. First, opposition to the grant of a standard patent may be based on the grounds set out in s 59 of the Act. Second, the opponent bears the relevant onus both before the delegate and before me on appeal to establish the relevant ground(s). Third, for the opponent’s appeal to succeed it must be “clear” or “practically certain” that the patent, if granted, would not be valid; I will discuss any significance attaching to the different formulations in a moment. Fourth, the standard of proof is generally that prescribed by s 140 of the Evidence Act 1995 (Cth), but subject to the threshold that I have just mentioned.
8 The basis for the requirement that an opponent must show that it is “clear” or “practically certain” that a patent if granted would not be valid was illuminated by Emmett J in F Hoffman-La Roche AG v New England Biolabs Inc (2000) 99 FCR 56, following Genetics Institute Inc v Kirin-Amgen Inc (1999) 92 FCR 106 at [17] to [21]. Because there are two stages post acceptance at which validity might be challenged, namely, pre-grant opposition proceedings and post-grant revocation proceedings, there is a distinction between the two proceedings. As his Honour said, this is “consistent with the proposition that pre-grant opposition is intended to provide a relatively inexpensive mechanism for resolving third party disputes as to validity” and that “[t]he purpose of pre-grant opposition proceedings is to provide a swift and economical means of settling disputes that would otherwise need to be dealt with by the courts in more expensive and time consuming post-grant litigation” (at [47] and see also [66]).
9 In determining whether a ground of opposition was made out on appeal, his Honour adopted the test that it “should appear clear to the Court that no patent granted in respect of the specification would be valid”, and accepted in this respect that where the proposed patent was tricky the opponent may be entitled to adduce evidence of considerable complexity to establish to the requisite degree that such a patent would, if granted, be invalid. So his Honour said at [67]:
The language employed in the cases to which I have referred suggests that it should appear clear to the Court that no patent granted in respect of the specification would be valid. I consider that, before the Court would uphold an opposition to the grant of a patent, the Court should be clearly satisfied that the patent, if granted, would not be valid. That, however, is not to say that an opponent should not be permitted appropriate opportunity to lead evidence-in-chief as to the facts that are designed to demonstrate, with the requisite degree of clarity, that a patent, if granted, would not be valid. Where the subject matter of the patent is one of complexity, of necessity, the evidence that an opponent would be entitled to adduce would itself be of considerable complexity.
10 Further, in Austal Ships Pty Ltd v Stena Rederi Aktiebolag (2005) 66 IPR 420; [2005] FCA 805, Bennett J considered such principles in pre-grant opposition proceedings in the context of expert opinion evidence directed to the ultimate question of whether an asserted ground of opposition had been made out. Her Honour said (at [12]):
I can accept that a lower standard may apply to proof of evidence such as whether a document has been published or, indeed, whether a prior art vessel was well-known. I do not accept that it properly applies to the factual question that itself is the test for obviousness or lack of inventive step. Where the factual question is itself the legal test, as set out in s 7(3) of the Act, it seems to me that it should be determined at the higher standard. That means that where there are two opposing expert views that are conclusive on obviousness, both presented bona fide by witnesses of accepted expertise, unless one set of views can be rejected on proper grounds, the legal burden to establish a ground of opposition is not discharged; the court cannot be practically certain that obviousness or lack of inventive step is established.
11 Now there seems to be some difference in the authorities concerning the formulation of the relevant test. Some authorities use the language of “practically certain”, whilst other authorities use the language of “appears clear” or “clearly satisfied” that the patent, if granted, would be invalid. It has been suggested that the varying phraseology is to similar effect (Aspirating IP Ltd v Vision Systems Ltd (2010) 88 IPR 52; [2010] FCA 1061 at [33] per Besanko J) although I am not so convinced; there is something to be said for justifying the historical use of the former phrase in the earlier acceptance phase only, which is ex parte, rather than in the later opposition phase, which is inter partes. In any event, for my part I prefer the phraseology of Emmett J. But I have also adopted the approach of Bennett J in Austal Ships at [12] that where there are conflicting sets of expert opinions on an issue such as a lack of inventive step presented bona fide by witnesses of accepted expertise, then unless one set of views can be rejected on proper grounds, the legal burden on the opponent to establish to the requisite degree the relevant ground of opposition will not be discharged. In such a scenario, I could not be clearly satisfied that the ground of opposition had been made out. But notwithstanding these observations, I accept for present purposes that primary facts (as distinct from secondary conclusions or the ultimate ground of opposition question) that are relevant to and that are said to support any ground of opposition need only be established by the opponent on the balance of probabilities.
12 In summary, I have rejected MLA’s principal grounds of opposition and its corresponding grounds of appeal.
13 First, MLA’s central attack that the claims do not satisfy the “manner of manufacture” requirement fails. D’Arcy v Myriad Genetics Inc (2015) 258 CLR 334 is distinguishable, save for one claim to a particular isolated polynucleotide out of the 15 claims. But to demonstrate this I will need to discuss that case in more detail than is usual. The conclusion is clear, but the analysis is not without its complexity. I should note that my case, except in one minor aspect, does not concern whether product claims to nucleic acid molecules per se are patentable, even if isolated from their natural environment and artificially created in a chemical sense or, as so isolated, having different actual or potential chemical properties to the naturally occurring nucleic acid in its natural environment. The case that I have to resolve principally concerns method claims. Further, the case that I have to resolve does not just concern looking at a claim in relation to a nucleic acid molecule and considering whether one should, in the context of such a claim, characterise the invention in terms of its chemical structure and properties on the one hand or its genetic informational content on the other hand. Further, the case that I have to resolve does not just deal with claims that involve the discovery of an objectively observed statistically significant correlation between genotype and phenotype. In the context of the claims that I have to consider, that is only the starting point for the analysis rather than the finishing point to determining patentability.
14 Second, MLA’s other significant attacks, namely, a lack of novelty and lack of inventive step go nowhere close to satisfying the threshold MLA must demonstrate on this appeal given the genuine and substantial conflict in the expert opinion evidence.
15 Third, MLA has also advanced arguments concerning lack of utility, lack of sufficiency and lack of fair basis which I have also rejected, save as to an aspect of lack of utility which relates to two of my findings on construction. But contrastingly, MLA has had some success on some questions of construction and associated grounds dealing with lack of clarity and lack of definition. Several integers of the relevant claim(s) will need to be amended to deal with questions of linkage disequilibrium between relevant single nucleotide polymorphisms and also to address questions of statistical significance. And if appropriate amendments are made, then the conclusions of lack of clarity, lack of definition and the residual aspect of lack of utility that I have reservations about fall away.
16 It is convenient to divide the balance of my reasons into the following sections:
(a) Some scientific principles – [18] to [145];
(b) The 253 Application – [146] to [175];
(c) The steps of the claimed invention – [176] to [212];
(d) Construction of the claims – [213] to [385];
(e) Manner of manufacture – [386] to [516];
(f) Lack of novelty – [517] to [677];
(g) Lack of inventive step – [678] to [819];
(h) Lack of utility – [820] to [882];
(i) Lack of sufficiency – [883] to [914];
(j) Lack of fair basis – [915] to [931];
(k) Lack of clarity and definition – [932] to [946]; and
(l) Conclusion – [947] to [949].
17 The length of what follows is in part a reflection of the notable sophistication with which Ms Katrina Howard SC and Mr Tom Cordiner QC for MLA, and Mr Christian Dimitriadis SC and Mr Benjamin Fitzpatrick for Branhaven, have presented their cases.
Some scientific principles
18 Let me begin with a summary of some of the non-controversial scientific principles that will inform my later discussion. The summary has been drawn from the expert evidence including aspects of the technical summary provided by MLA to the extent that it accords with the evidence. The summary reflects common general knowledge as at the priority date of the relevant person skilled in the applicable art, unless I indicate otherwise.
(a) Glossary of key terms
19 To facilitate comprehension, the following glossary of key terms is necessary, although I will elaborate on some of the concepts in more detail later:
Allele
An allele is one of two or more alternative forms of the same gene or same genetic locus. An individual inherits two alleles for each gene, one from each parent. If the two alleles are the same, the individual is homozygous for that gene/loci. If the alleles are different, the individual is heterozygous as represented in the following diagram:
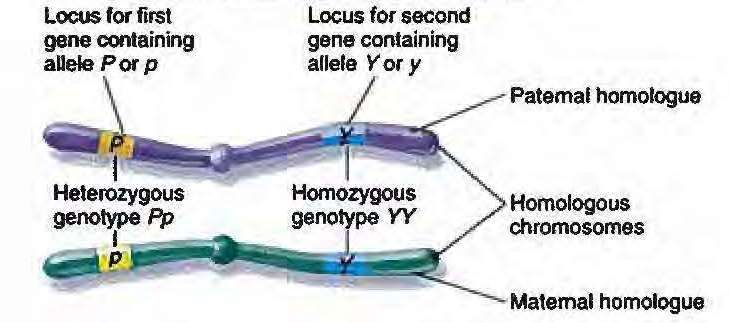
Chromosome
A coiled three dimensional structure of double helix DNA, coiled around support and scaffold proteins and containing many genes.
Codon
A codon is a trinucleotide sequence of DNA bases (A, C, G and T) which encode for a specific amino acid. There are 64 different codons. 61 codons specify for amino acids (although sometimes for the same amino acid), while the remaining three codons are used as stop signals.
Contig
A contig (short for contiguous) is a reference to overlapping DNA sequences.
DNA
Deoxyribonucleic acid (DNA) is the molecular code of inheritance. The DNA sequence is the arrangement of the 4 letters (representing DNA bases) of the genetic code into information.
Epistasis
The interaction between different genes (as opposed to different alleles) including suppression or inhibition of the phenotypic expression of one gene by the second non-allelic gene (gene at a different locus).
Exon
An exon is that portion of a gene that codes for amino acids. In mammals, most gene sequences are broken up by one or more DNA sequences called introns. The parts of the gene sequence that are expressed as a protein(s) are called exons.
Gamete
A germ cell (egg cell or sperm cell) that is typically, and for present purposes is, haploid.
Gene
Generally, this is the basic unit of heredity in organisms. It consists of a sequence of DNA in mammals coding ultimately for a protein. Proteins work together to contribute to traits. I will return to the definition of “gene” later and whether it includes not only exons but also introns and other regulatory but non-coding sequences. The definition of “gene” is one of the issues between the parties relating to the construction of claim 1 of the 253 Application.
Genetic marker
A genetic marker is a DNA sequence with a known physical location on a chromosome.
Genome
The genome is the entire set of genetic instructions found in a cell. The genome consists of pairs of chromosomes.
Genomic selection (GS)
I have explained the parameters of genomic selection (GS) in sub-section (f) below, which I have described as approach 5.
Genome wide association study (GWAS)
I have explained the parameters of a genome wide association study (GWAS) in sub-section (f) below, which I have described as approach 6.
Genotype
A genotype is an individual’s collection of genes. But the term can also refer to the two alleles inherited for a particular gene.
Haploid
A single set of unpaired chromosomes.
Haplotype
A collection of specific alleles, DNA variations, or polymorphisms, which tend to be inherited together due to their proximity to each other. For example, there are two simple haplotypes present in the diagram set out above in the discussion of an allele: PY and pY.
Homologous recombination
Homologous recombination is a type of genetic recombination that occurs during meiosis (the formation of egg and sperm cells). Paired chromosomes from the male and female parent align so that similar DNA sequences from the paired chromosomes cross over each other. Crossing over results in a shuffling of genetic material and is an important cause of the genetic variation seen among offspring.
Intron
An intron is a portion of a gene (a proposition that I will justify later) that does not code for amino acids. In mammals, most gene sequences include one or more introns.
Karyotype
The number and appearance of chromosomes in the cell nuclei of, say, a mammal.
Linkage
Linkage is the close association of genes or DNA markers on the same chromosome. The closer two genes/markers are to each other, the greater the probability that they will be inherited together.
Linkage disequilibrium (LD)
The concept of linkage disequilibrium (LD) is used in population genetics to describe a non-random association of alleles at two or more loci on the same chromosome reflecting haplotypes descended from a single ancestral chromosome. It is a measure of whether an allele at one locus tends to be found more often with an allele at another locus. It is a measure of combinations of alleles or genetic markers in a population that are more frequently found to be inherited together than would be expected from the random formation of haplotypes. The further apart two alleles or markers are, the less likely that they are to be “linked”.
Locus/Loci
A locus is the specific physical location of a gene or DNA marker on a chromosome. The plural of locus is “loci”. A variant of a similar DNA sequence located at a locus is called an allele.
Marker assisted selection
Use of genetic markers in methods that allows for selection of animals with desired genotypes.
Microsatellite
Microsatellite sequences are repetitive DNA sequences that are usually several base pairs in length (e.g. GCGCGCGCGC). They are used as genetic markers to follow the inheritance of genes. They are sometimes referred to as short tandem repeats. When microsatellite sequences are replicated, the repetitive nature of the sequence means that the copying mechanisms can make errors in the copy, typically by adding (in this example) an additional GC. The number of times that the unit is repeated in a given microsatellite can be highly variable, a characteristic that makes them useful as genetic markers. Microsatellites are markers suitable for QTL mapping. Sometimes an alternative description is given for microsatellites, being what are known as polymorphic repeat sequences. Microsatellites mainly occur in non-coding sequences, usually a 2 to 5 nucleotide repeating sequence e.g. GCGCGCGCGC, where the repeating unit is present in different numbers (5 GC repeats in this example).
Non-coding DNA
Non-coding DNA sequences do not code for amino acids. Most non-coding DNA lies between genes on a chromosome. Other non-coding DNA, called introns, is found within genes (on the definition of “gene” which I have accepted). Some non-coding DNA plays a role in the regulation of gene expression; whether such DNA is part of a “gene” is a matter for debate.
Phenotype
A phenotype is an individual’s observable physical characteristics or traits, whether generally or for a particular trait, that can vary from animal to animal. The genetic contribution to the phenotype can sometimes be referred to as the genotype.
Pleiotropic
The effect of an allele on multiple traits.
Polymorphism
A polymorphism involves one of two or more variants of a particular DNA sequence. The most common type of polymorphism involves variation at a single base pair, called a single nucleotide polymorphism or SNP. But polymorphisms can also be much larger in size and involve long stretches of DNA.
Qualitative trait
A qualitative trait is one in which a clear distinction between the presence of the trait, or the absence of the trait, can be determined (i.e. for cattle, horned or polled (no horns)).
Quantitative trait
A quantitative trait is one that can vary continuously from animal to animal. Quantitative traits can be attributed to one gene, but they are usually the result of the activity of a combination of genes on different chromosomes. In other words, a quantitative trait can be the sum effect attributable to two or more genes. Indeed, a large number of genes may each make a small contribution to the overall trait. Alternatively, a small number of genes may make a large contribution to the overall trait.
Quantitative trait locus (QTL)
Quantitative trait loci (QTLs) are stretches of DNA containing or linked to particular genes that correlate with a continuously variable trait. “Linked” means “associated with, or attributable to”.
Shotgun sequencing
Shotgun sequencing is a laboratory technique for determining the DNA sequence of an organism’s genome. The method involves breaking the genome into a collection of small DNA fragments that are sequenced individually. A computer program looks for overlaps in the DNA sequences and uses them to place the individual fragments in their correct order to reconstitute the genome.
Single nucleotide polymorphism (SNP)
As I have already touched on, single nucleotide polymorphisms (SNPs) are a type of polymorphism involving variation of a single base pair at specific loci. It is a DNA sequence variation in which a single nucleotide differs between members of a biological species or differs in the one member as between paired chromosomes. SNPs can act as biological markers to locate genes that are associated with disease or other traits of interest. When SNPs occur within a coding region of a gene or in a regulatory region for a gene, they may play a more direct role in the relevant trait by affecting the gene’s function.
SNP chip/SNP array
A SNP array is a solid substrate, typically glass, containing a specific series of short nucleic acid sequences immobilized at known locations. The array includes pairs of sequences, varying by a specific SNP. Sample DNA is incubated with the array to allow binding to the array. Probes are used to detect binding between the sample DNA and the DNA immobilized on the array, with binding or non-binding at a specific location on the array indicating the presence or absence (as the case may be) of a specific SNP in the sample DNA.
20 In order to explain the relativity of some of these concepts, the following analogy provided by Professor Graham Plastow (an expert called by Branhaven) should be of assistance:
The genome the entire world
A chromosome a country in the world
QTL a state within the country
A gene a city within the state
A SNP a street address in that state
(b) The genetic code
DNA
21 Genetic information is stored in the form of DNA. DNA consists of four different nucleotides, each consisting of an identical pentose sugar group (the sugar backbone), an identical phosphate group and one of four nitrogenous bases (adenine: A; guanine: G; thymine: T; and cytosine: C). The DNA sequence is the arrangement of unique combinations of these four bases.
22 DNA exists in the form of a double helix (see Figure 1 below) comprising two complementary strands which are bound together via hydrogen bonding between specific complementary (base paired) nucleotides (A bonds with T and C bonds with G). As such, each strand provides a complementary template of the other strand.
Figure 1 – Double helix structure of DNA
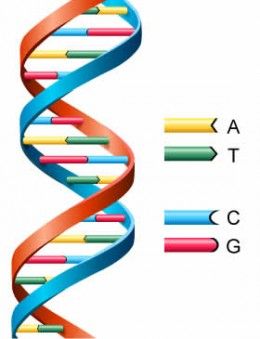
23 The standard format for writing and reading a DNA sequence is to list the “coding” strand in a 5’ (five prime) to 3’ (three prime) direction. The terminology 5’ and 3’ relates to the carbon numbers in the sugar backbone of nucleic acids, with a hydroxyl group on the third carbon of the sugar backbone of one nucleic acid bonding with a phosphate group on the fifth carbon of the sugar backbone of the subsequent nucleic acid (see Figure 2 below). This gives DNA conceptual directionality when synthesised and read.
Figure 2 – Directionality of nucleic acids in DNA
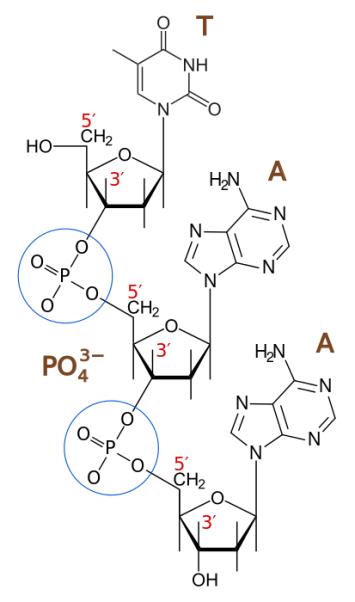
24 Consequently, and for example, a DNA sequence such as ATGGCGTGTGACGAAATAGGG denotes the coding strand (5’-ATGGCGTGTGACGAAATAGGG-3’), which when read from left to right can provide the code for the relevant amino acid(s) and ultimately the relevant protein. However, as DNA is a double stranded molecule, each coding sequence has a complementary strand, with each nucleotide in one strand being bound to its complementary nucleotide on the complementary strand (see Figure 3 below). This complementary strand can be written in a 5’ to 3’ direction. For example, the complement of the above sequence when written 5’ to 3’ (known as the reverse complement) is 5’-CCCTATTTCGTCACACGCCAT-3’. However, usually only the coding strand is written.
Figure 3 – Complementary DNA sequences
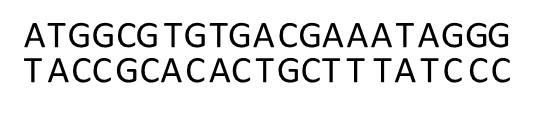
Genetic code
25 DNA stores genetic information and can be duplicated to permit the genetic information to be passed on to daughter cells, via a process known as mitosis. Specific portions of DNA (genes) are “read” by molecular structures in the cell and are converted (through the process of transcription) into another form of polynucleotide called ribonucleic acid (RNA).
Codons
26 To allow four nucleotides to encode for twenty different amino acids, DNA is “read” in blocks of three sequential nucleotides known as a “codons”. For example, ATGGCGTGTGATGAAATAGGG, can be read as ATG-GCG-TGT-GAT-GAA-ATA-GGG. Each codon codes for an amino acid.
27 As can be seen in the Table below, each amino acid (with the exception of Methionine and Tryptophan) is encoded by more than one codon.
Amino Acid | Single letter abbreviation | Three letter abbreviation | Codon |
Alanine | A | Ala | GCT, GCC, GCA, GCG |
Cysteine | C | Cys | TGT, TGC |
Aspartic acid | D | Asp | GAT, GAC |
Glutamic acid | E | Glu | GAA, GAG |
Phenylalanine | F | Phe | TTT, TTC |
Glycine | G | Gly | GGT, GGC, GGA, GGG |
Histidine | H | His | CAT, CAC |
Isoleucine | I | Ile | ATT, ATC, ATA |
Lysine | K | Lys | AAA, AAG |
Leucine | L | Leu | TTA, TTG, CTT, CTC, CTA, CTG |
Methionine | M | Met | ATG |
Asparagine | N | Asn | AAT, AAC |
Proline | P | Pro | CCT, CCC, CCA, CCG |
Glutamine | Q | Gln | CAA, CAG |
Arginine | R | Arg | CGT, CGC, CGA, CGG, AGA, AGG |
Serine | S | Ser | TCT, TCC, TCA, TCG, AGT, AGC |
Threonine | T | Thr | ACT, ACC, ACA, ACG |
Valine | V | Val | GTT, GTC, GTA, GTG |
Tryptophan | W | Trp | TGG |
Tyrosine | Y | Tyr | TAT, TAC |
Stop codon | - | Term | TAA, TAG, TGA |
Start codon
28 The codon for methionine (ATG) is referred to as the “start codon” as methionine is the first amino acid in the proteins of all eukaryotic cells (i.e. a cell with a nucleus containing the chromosomes). Therefore, ATG when read in the correct open reading frame will signify the start of a protein coding region in a gene.
Stop codon
29 As can be seen from the above Table, three codons (TAA, TAG and TGA) are referred to as “stop codons”. These stop codons function to indicate the end of a protein encoding region of DNA. The redundancy in DNA sequences and the presence of stop codons are important elements when considering how genetic variation can influence protein structure and function.
Genes
30 A gene, the definition of which I will discuss later, can be considered to include a span of DNA that ultimately encodes for a protein. The process by which a cell produces a protein from a gene is referred to as gene expression and invariably includes a messenger ribonucleic acid (mRNA) intermediate. The DNA is transcribed into mRNA through several steps that I will describe in a moment. The mRNA is then translated, ultimately, into a protein(s). Each codon is read with the corresponding amino acid formed. Chains of amino acids are formed with peptide bonds between each amino acid. The polypeptide so formed is the protein. For a further uncontroversial discussion of amino acids, peptide bonds and polypeptides, see Idenix Pharmaceuticals LLC v Gilead Sciences Pty Ltd [2017] FCAFC 196 at [50] to [53] per Nicholas, Beach and Burley JJ.
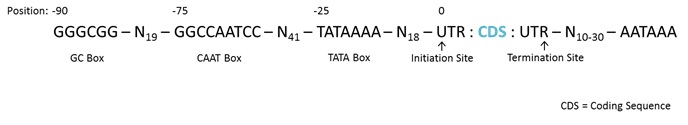
31 The transcription initiation site is generally determined by specific regions which are upstream (5’) of the transcription initiation site. The expression UTR refers to either of two untranslated regions either side of the coding sequence. The upstream regions are known as “promotor sites” and usually are defined by highly conserved regions of DNA that serve as binding sites for transcription factors which facilitate or activate transcription of DNA to RNA. Generally, there are three consensus sequences (sequences with similar structures) upstream of the transcription initiation site (position 0). The first of these is the TATA box (TATAAAA) which is found approximately 25 nucleotides upstream (position -25) of the transcription initiation site. The second of these is the CAAT box (GGCCAATCC) approximately 75 nucleotides upstream of the initiation site. The third of these is the GC box (GGGCGG) approximately 90 nucleotides upstream of the transcription initiation site (see Figure 4 below). Transcription continues beyond the translation termination site as shown below, and the generated RNA is then later cleaved.
Figure 4 – Simple gene structure including UTRs
Exons and introns
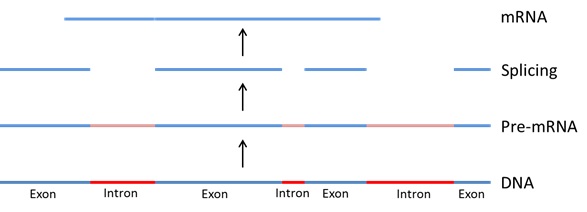
32 A gene, on the generally accepted view of what constitutes a “gene” that I will discuss in more detail later, also includes sections that are transcribed into RNA but are then excised before the resulting mRNA is translated into a protein (see Figure 5 below). These sequences that are excised are called introns, and do not contribute to coding for the amino acid sequence leading to the protein. The sections of the DNA that ultimately code for the protein are located within regions called exons.
Figure 5 – Transcription of DNA to mRNA
33 In general, exons only account for about 4% to 5% of the nucleotides that are transcribed. Furthermore, there are considerable spans of DNA, known as intergenic DNA, that sit between transcribed regions and which do not encode for RNA or proteins. As such there is only a relatively small portion of DNA in mammals that actually encodes for proteins.
34 Nevertheless these vast non-coding (intergenic and intronic) regions have important biochemical and biological functions. Portions of these non-coding regions play a critical role in regulation of the coding regions and can determine how much, or how little, mRNA (and consequently protein) is produced. In some instances, these regulatory regions act as binding sites for regulatory elements. Binding of these regulatory elements to the DNA influences the rate of transcription of the coding DNA into mRNA. Furthermore, non-coding DNA can have important structural properties that may play a role in chromosome structure, the function of centromeres (the central portion of a chromosome), the process of mitosis (cell division) and the process of meiosis (generation of gametes).
35 Due to the importance of non-coding DNA, especially the DNA in close proximity to coding DNA, on one view the term “gene” can also include the promoter and enhancer regions as well as the DNA regions in between these regions and the transcribed DNA. I will return to the construction question concerning “gene” later in my reasons.
36 Before proceeding further I should also note that in relation to the statements in the preceding three paragraphs, there is a debate between the parties as to whether such matters (or part thereof) were part of common general knowledge as at the priority date.
Chromosomes
37 In eukaryotic cells, genetic information is stored in coiled and condensed structures called chromosomes. Each chromosome consists of a complex three-dimensional structure of double helix DNA, which has been coiled around support and scaffold proteins, to form a condensed cluster of DNA.
38 Many genes can be physically clustered together on a single chromosome (indicated by the dark band in Figure 6 below). Depending upon their proximity to one another, they may have a propensity to be inherited together.
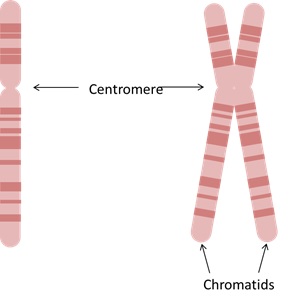
39 The structure of a chromosome varies depending on the stage of the cell cycle. Typically, a chromosome is a single supercoiled length of DNA which has a spindle of fibres in the middle called the centromere (see Figure 6(a) below). But in the later stage of the cell cycle, the chromosome is duplicated, to form two copies of the chromosome, which are connected at the centromere. In this form, each copy is referred to as a chromatid (see Figure 6(b) below). Each chromatid is then separated during duplication of the cell and provides the genetic information for one of the two cells (the original parent cell and the daughter cell).
Figure 6 – Chromosome structure
(a) (b)
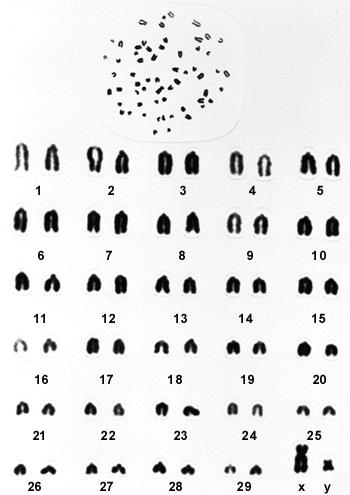
40 Chromosomes exist in pairs. A mammal inherits two copies of each chromosome, one from their mother and one from their father. The notable exception to this are the sex chromosomes in males, whereby a male has one X chromosome (inherited from his mother) and one Y chromosome (inherited from his father). In cattle there are thirty pairs of chromosomes (sixty chromosomes in total – see Figure 7 below).
Figure 7 – Bovine karyotype
Mutations and alleles
41 During cell replication, DNA is duplicated so that a copy of the genetic code can be incorporated into a daughter cell. During this process, double stranded DNA is separated, and each strand is used as a template to produce a second complementary strand. Cells have extensive proof-reading and error-checking abilities. Consequently, DNA is generally replicated, and passed onto the daughter cell, unchanged. However, despite this stringency, changes in the DNA sequence, known as mutations, can occur. Mutations can be caused by errors in the replication of DNA known as copy errors. Another cause of errors is exposure to external influences, for example exposure to chemicals, free radicals or radiation.
42 Gene mutations can be classified into either acquired (somatic) mutations or hereditary (germline) mutations.
43 Somatic mutations are mutations that only appear in a specific cell(s) during an animal’s life, and are then reproduced in the daughter cells from that specific cell during replication. Consequently, these mutations only exist in a subset of cells within an animal. The mutations that lead to cancer are examples of somatic mutations.
44 Contrastingly, germline mutations are mutations that are either formed in the fertilised egg shortly after fertilisation, or formed in the gametes (egg or spermatozoa) of the parents (in the same way described above for somatic mutations). Germline mutations will be present in every cell of the animal’s body. Accordingly, they may be present in the gametes that the animal produces and hence in his or her offspring.
45 Differences between individuals in DNA sequence are in part due to mutations that have occurred in their ancestors. These mutations provide the basis for genetic diversity within populations, and account for a significant portion of the differences in the physiology of individuals within a given population. Moreover, combinations of mutations are specific to an individual, and are useful in population studies to identify genetic differences between individuals, and assess how those differences may influence the physiology of that individual.
Types of mutations
46 The simplest form of mutation is called a point mutation. This type of mutation arises when a single nucleotide is substituted by another single nucleotide (e.g. a T mutates to a C). An individual carrying this mutation may pass it on to many descendants so that both the original form (T) and the mutated form (C) exist in the population. When a nucleotide at the same location (locus) in the genome varies amongst individuals, this is called a single nucleotide polymorphism (SNP or pronounced “snip”) (see the red boxes in Figure 8 below). SNPs can be identified by extracting a sample of DNA from individuals, sequencing it and then comparing the homologous sequence from several individuals within the same species.
Figure 8 – Identification of SNPs
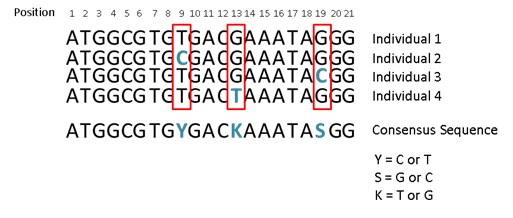
47 A SNP in that part of the DNA that codes for amino acids within a protein (a coding SNP) can be classified as one of the following three types:
(a) First, there is a type known as a “synonymous mutation”. A synonymous mutation occurs when the SNP does not result in a change in protein sequence (i.e. the mutated codon still encodes for the same amino acid). For example the sequence for individual 1 in Figure 8 above (assuming it is an in-frame coding region) will be read as ATG:GCG:TGT:GAC:GAA:ATA:GGG and will encode for the amino acid sequence Met, Ala, Cys, Asp, Glu, Ile, and Gly. Likewise, the sequence for individual 2 (ATG:GCG:TGC:GAC:GAA:ATA:GGG), which has a T/C mutation at position 9, will encode for the same amino acid sequence. This is due to the third codon of individual 2, spanning from position 7 to 9 (TGC), and the third codon of individual 1 (TGT), both encoding for the same amino acid, Cysteine (Cys). Consequently, despite a mutation in the DNA the resultant protein has not changed. As such it is likely that this mutation will have no or very little functional consequence on the animal.
(b) Second, there is a type known as a “missense mutation”. Contrastingly to a synonymous mutation, a missense mutation occurs when a SNP occurs inside of a coding region and results in a change in the protein sequence (i.e. the mutated codon encodes for a different amino acid). With reference to the sequences illustrated in Figure 8 above, the sequence for individual 3 (having a G/C mutation at position 19) would result in the last codon being CGG (instead of GGG for individuals 1, 2 and 4). The consequence of this is that the codon for individual 3 (CGG) encodes for Arginine, while the codon for individuals 1, 2 and 4 (GGG) encodes for Glycine. This change in amino acid sequence may change the function of the protein.
(c) Third, there is a type described as a “nonsense mutation”. A nonsense mutation occurs when a SNP occurs inside of a coding (exonic) region and results in a premature stop codon. As I have already indicated, three codons (TAA, TAG, TGA) encode for stop codons which signal the termination of translation of mRNA into a protein. As such, the premature introduction of a stop codon leads to a truncated protein, which potentially may have significantly reduced or no function. With reference to the sequences illustrated in Figure 8 above, the sequence for individual 4 (having a G/T mutation at position 13) would result in the fifth codon (spanning positions 13 to 15) being TAA (instead of GAA as per individuals 1 to 3). The consequence of this is that the codon for individual 4 (TAA) encodes a stop codon, while the codon for individuals 1 to 3 (GAA) encodes Glutamic acid. This change will result in the protein translated from the mRNA sequence of individual 4 to be prematurely stopped after four amino acids (Met, Ala, Cys, Asp, instead of Met, Ala, Cys, Asp, Glu, Ile, and Gly).
48 So far I have discussed coding SNPs. Let me turn to other types of SNPs. SNPs that occur outside the coding DNA are called non-coding SNPs and may have an effect on the regulation of a gene, for example, by altering the binding of a regulatory molecule to the DNA.
49 More generally, let me now turn to some other types of mutations being insertions/deletions, duplications and repeat expansion.
50 Insertion and deletion mutations occur when nucleotides are introduced or removed from a DNA sequence. Collectively these are known as “indels”. These indels can be as small as one nucleotide change or as large as 10,000 nucleotides. They can be even larger on a chromosomal level. Like SNPs, these indels can occur in coding or non-coding regions and their consequence will vary depending on their location as follows:
(a) If the indel is in an exon and its length is 3, 6, 9 … nucleotides (i.e. a multiple of 3), it will add or delete a number of amino acids to the protein.
(b) If the indel is in an exon and its length is not a multiple of 3 nucleotides, it will lead to a frame shift. This will generally result in significant alterations to the structure of the subsequently produced protein. As I have said, DNA sequences are read as groups of three nucleotides (codons). Consequently, if an indel is not a multiple of 3 nucleotides, then the amino acids encoded after the mutation are generally altered.
(c) An illustration of an insertion mutation is provided in Figure 9 below. As can be seen in Figure 9(a), a guanine (G) has been inserted after position 3 of the sequence for individual 2. Consequently, the nucleotides after the insertion are displaced one position and the codons after the insertion have been changed (as shown in Figure 9(b)) for individual 2. The insertion of a guanine results in a premature stop codon at codon number 4 (positions 10 to 12) for individual 2. This is typical of what happens when there is an insertion or deletion of nucleotides in a coding sequence, except for when nucleotides are inserted in multiples of 3. But when the indel is outside of the coding region, it is less likely that the indel will have a functional effect, although it may alter gene regulation.
Figure 9 – Insertion mutation
(a)
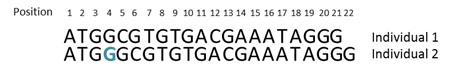
(b)

51 A duplication mutation consists of a piece of DNA that is abnormally copied one or more times. The duplication may be as small as two nucleotides, or it may involve the duplication of large chunks of DNA (10,000s – 100,000s nucleotides).
52 As for repeat expansion mutations, nucleotide repeats are short DNA sequences that are repeated a number of times in a row. For example, a trinucleotide repeat is made up of 3-base-pair sequences, and a tetranucleotide repeat is made up of 4-base-pair sequences. A repeat expansion is a mutation that increases the number of times that the short DNA sequence (i.e. the tri- or tetra-nucleotide) is repeated. When in a coding region, this type of mutation can cause the resulting protein to function improperly. However, when these mutations happen outside of a coding region, they are significantly less likely to have a functional effect.
Alleles
53 When a segment of DNA of one individual at a locus varies in comparison to the DNA at the same locus of another individual, these different forms of DNA are called alleles. The term allele can be used to refer to a variant form of a gene (i.e. an allele of the gene) or may refer to a variation at a particular locus (i.e. an allele of a SNP).
54 An individual site in the genome is usually either monomorphic (all sequences are the same at this site) or polymorphic (sequences vary) with only 2 alleles, being the non-mutated wild-type (WT) form (possessed by the majority of individuals in a population) or the mutated form. For example a gene corresponding to the growth hormone (GH) may have a A/T SNP at position 177. This implies a biallelic SNP and may also be described as two alleles of the GH gene, one with an A at position 177 and one with a T at position 177, which can be denoted as 117A>T.
55 But while in the case of SNPs there are usually only two different alleles at any one position or locus, that is WT or mutated at a specific position, there are many positions at which polymorphisms exist. For example in addition to the mutation at position 177, the growth hormone gene may also have a mutation (for example a C/G mutation) at position 208. In this example, there are then four possible alleles: 117A/208C (i.e. an A at position 117 and a C at position 208), 117A/208G, 117T/208C and 117T/208G. This is referred to as GH being multi-allelic. But an alternative nomenclature is to refer to a combination of alleles at different positions as a haplotype. In this example, there are relevantly 4 haplotypes.
56 As I have explained, mammals inherit two copies of each chromosome, one from each parent. Consequently, each individual has two copies of each gene, except in the case of male sex chromosomes. Consequently, it is possible for each individual to have different allelic forms of a gene. When an individual has inherited the same allele from their mother and father, they are called homozygous. When an individual has inherited different alleles from their mother and father, they are called heterozygous.
(c) Genotypes and phenotypes
57 The genotype of an animal is a description of the alleles that it carries at one or more loci. For example, a WT allele of a gene may be expressed as a D (one allelic version of the gene), while a mutated form of the gene (a second allelic version of the gene having a mutation) can be expressed as D’. As mammals inherit two copies of each gene, one from their father and one from their mother, consequently in the example of the D locus there can be three possible genotypes (DD, DD’ or D’D’).
58 The appearance of an organism is referred to as its phenotype. A phenotype may be a clearly visible trait such as coat colour or growth rate, or it may be a trait that is not observable with the naked eye such as milk fat content. In simple terms, an organism’s phenotype will be determined by a combination of both its genotype and its environment. Accordingly, two individuals may have differing phenotypes or differing observable traits even if they have the same genotype.
59 Now while the genotype of an individual may influence its phenotype, an alteration of a genotype does not guarantee an altered phenotype. There are several reasons for this.
60 First, a mutation may not alter an expressed protein or other functional factor. This may be because the mutation is not present in a coding or regulatory region of DNA. Alternatively, the mutation may be in one of these regions, but does not alter the function of these regions. For example a SNP in a coding region may not necessarily alter the amino acid sequence of the protein that is produced as I have explained earlier; one may refer to this as a “silent mutation”. Alternatively, even if the mutation does change the amino acid sequence of the produced protein, the substituted amino acid may not alter the functioning of the protein.
61 Now proteins exist in three-dimensional structures, with the structure partly determined by the amino acid sequence. Some amino acids are hydrophobic and will have a propensity for the centre of a protein, while others are hydrophilic and therefore will have a propensity for the outer portion of a protein. As such, a substitution of a hydrophilic amino acid for a hydrophobic amino acid can influence the three-dimensional structure of a protein. Further, only portions of the protein are active sites (for example a catalytic site or a site which interacts with another molecule), with the accessibility to these sites, and the activity of these sites, influenced by the three-dimensional structure of the protein. Typically, amino acid substitutions which result in a change in the three dimensional structure of a protein, or are in an active site, will result in a functional change in the protein. But not all amino acids in a sequence are critical. Some amino acid changes will not result in a change in the three-dimensional structure of the protein. Further, the substituted amino acid might not be in an active site of the protein. Additionally, the effect of the alteration in the protein might not be sufficient to alter a biological pathway and therefore may not present as an altered phenotype.
62 Second, a mutation in an allele might only be in one of the two chromosomes of an individual. If the other, non-mutant allele is a dominant allele (I will explain this in a moment), then the dominant allele will mask the effect of the mutated allele. For example, where one allele of a gene (a non-mutant/WT allele) is denoted D and the mutated allele is denoted as D’, as I have said there are three possible genotypes: DD, DD’ or D’D’. If this gene influences disease resistance of an individual and D is the dominant form of the gene (and has improved resistance) and D’ is the recessive form of the gene (having decreased resistance), then individuals having the DD and DD’ genotype will still have the same phenotype. The dominant allele masks the presence of the recessive allele i.e. neither will have impaired disease resistance. Consequently, only the animals having the D’D’ genotype (homozygous recessive) will have a change in phenotype with a decrease in disease resistance.
63 More generally, where a mutation does influence a phenotype, for example when a mutation is in the coding sequence and causes a mutated protein, which results in an alteration of a biological process, then this mutation is called a causative mutation.
64 Traits are typically divided into two different categories based on how they can be measured. The two main categories of traits are referred to as qualitative traits or quantitative traits.
65 A qualitative trait is a trait in which a clear distinction between the presence of the trait, or the absence of the trait, can be determined. In cattle, an example of a qualitative trait is coat colour (e.g. black or red) or being horned or polled. The presence or absence of these traits is easily determined by the presence or absence of a particular feature. Qualitative traits account for a small proportion of observable traits in an animal.
66 The majority of traits in animals are considered to be quantitative traits. A quantitative trait is a trait that can be measured in a continuously variable manner, rather than being categorised into two distinct categories. Examples of quantitative traits in cattle are milk production, height and weight.
(d) Sexual reproduction and inheritance
67 Most mammalian cells are referred to as diploid cells as they include pairs of two homologous (similarly structured) chromosomes, with one of the chromosomes inherited from the father, and one inherited from the mother.
68 Sexual reproduction in mammals occurs when two gametes (a sperm cell and an egg cell), which each carry a single copy of a chromosome, fuse together to form a diploid cell (a cell with two homologous copies of each chromosome). As such, each gamete (one from the father and one from the mother) contributes half of the genetic component of an individual. In most cases, each gamete produced by the father (sperm) and the mother (ovum) are genetically distinct to the other gametes. This is one reason why siblings are different.
69 Before elaborating further, it is appropriate to discuss some history.
Mendelian inheritance
70 The plant hybridisation studies of the Augustinian friar and natural philosopher Gregor Mendel in the mid-nineteenth century provided the genesis of modern genetics. He established various principles concerning dominant and recessive alleles of genes, the segregation of genes and the independence of sorting of genes, although not strictly correct.
71 Mendel crossed pea plants having either a pure white flower or a purple flower (a qualitative trait). He noted that when the two different phenotypes were crossed, they did not produce a flower which was a blend of the two colours, but rather only produced offspring (denoted as a first generation (F1)) with purple flowers. But when F1 plants, which all had purple flowers, were crossed, their offspring had an assortment of purple and white flowers in a ratio of 3:1 respectively. From this observation, Mendel hypothesised about how genetic information is inherited.
72 Mendel hypothesised that there were two alternative forms of genes (different alleles), one that encoded for the purple flower (P) and one that encoded for the white flower (P’), and that each of the plants had two copies of these alleles. He also hypothesised that each of the two alleles were separated during reproduction (the law of segregation) and that the purple flower allele was dominant over the white flower allele (the law of dominance). Generally speaking, these hypotheses became established as fundamental aspects of modern genetics.
73 The experiment performed by Mendel and the genotype and phenotype of the plant crosses is set out in Figure 10 below. As can be seen, all of the progeny from the cross (denoted as F1) of the purple flowered plants with the white flowered plants produced purple flowered offspring. This is caused by the dominant purple allele of the gene (P) masking the presence of the recessive white flower allele (P’). However, when the first generation (F1) are crossed with each other, the resulting progeny (denoted as F2) produce both purple and white flowered plants. As can be seen, there are four possible genotypes in the F2 generation PP, PP’, P’P and P’P’. Due to the dominant nature of P over P’, three of these genotypes have a purple phenotype, and only one (P’P’) has a white phenotype.
Figure 10 – Mendelian inheritance
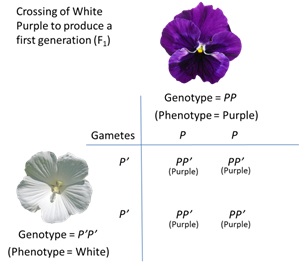
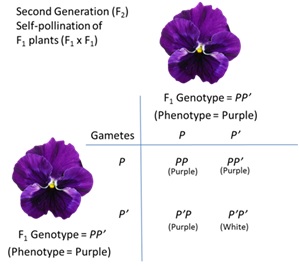
74 Mendel also set forth a third law, which has become known as the law of independent assortment. He noted that two distinct qualitative traits (for example, the presence/absence of lobed leaves and flower colour) were inherited separately of each other. That is, the presence of one trait was independent of the presence of the other trait. Generally speaking, this law has proven to be accurate when the genes for given traits are located on separate chromosomes or are spaced sufficiently far apart on the same chromosome. During sexual reproduction these loci are inherited independently of each other.
75 Now it has been accepted that there are limitations to Mendel’s three laws.
76 First, very few traits are monogenic (caused by a single gene). The majority of traits in an animal are polygenic traits caused by the interaction of many genes and environmental factors. Consequently, the simple pattern of inheritance observed in Mendel’s experiments is rarely seen in practice.
77 Second, alleles cannot always be classified as dominant or recessive. Many alleles can be incompletely dominant or incompletely recessive. Many alleles can be co-dominant. In the case of inheritance of incompletely dominant and incompletely recessive alleles, the resulting phenotype will typically be somewhere between the phenotype determined by each of the alleles alone, albeit closer to the incompletely dominant allele. While in the case of co-dominance, or additivity, the final phenotype will be halfway between the phenotype determined by each allele.
78 Third, not all alleles are independently assorted during reproduction. Many genes are clustered together on the same chromosome. Therefore particular alleles of those genes are more likely to be inherited together. As such, specific phenotypic traits can be linked.
79 Fourth, one gene can influence several traits and therefore inheritance of one allelic form of a gene may influence multiple traits.
80 Fifth, there are factors other than simple genetic mutations that can influence a trait, referred to as epigenetic effects. Epigenetic effects are genetic effects that are not related to changes in the DNA code, but can influence the production of specific proteins, and hence traits.
Chromosomal recombination
81 A consequence of sexual reproduction is genetic variation in offspring. Genetic variation is introduced during a process known as meiosis, whereby gametes are produced. As I have said, mammals are diploid species, having two chromosomes, one from their mother and one from their father. During meiosis, homologous chromosomes swap genetic material with each other to form a new, genetically unique, chromosome. This process is known as chromosomal recombination or chromosomal crossover. The new chromosome then forms the genetic material of the gamete.
82 The process of gamete formation (meiosis) follows the following steps and is detailed in Figure 11 below:
(a) Chromosomes are duplicated and form an X-shaped structure that consists of two chromatids, with each chromatid being a copy of the chromosome. This process happens for both the maternally inherited chromosome and the paternally inherited chromosome. In essence, at this stage, there are four copies of each chromosome in each cell.
(b) Each chromosome pairs up with its homologous counterpart so that a maternally inherited chromosome and its homologous paternally inherited chromosome are paired together.
(c) Once homologous chromosomes have paired, they exchange portions of genetic material. This process is random, and can happen at almost any position along the chromosome. Consequently, the likelihood that alleles at two loci on any one chromosome will be inherited together is partly dependent on the distance between the two loci. The closer they are, the more likely they are to be inherited together. The further apart the loci are, the more likely it is that a recombination event will happen between the two loci such that they are not inherited together.
(d) Following recombination, the chromatids separate from each other. Each of these chromatids then becomes a chromosome for a gamete. As seen below, each of the four chromatids is genetically distinct from the other. Each gamete possesses a single copy of each chromosome (i.e. each gamete is haploid).
Figure 11 – Meiosis (gamete formation)
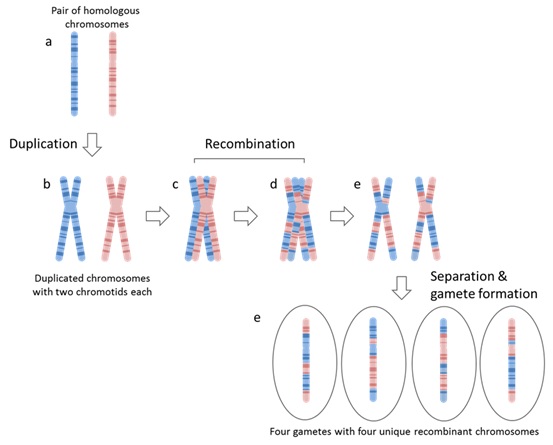
83 During sexual reproduction, haploid gametes fuse to form a diploid cell having one chromosome of each pair inherited from each of their parents.
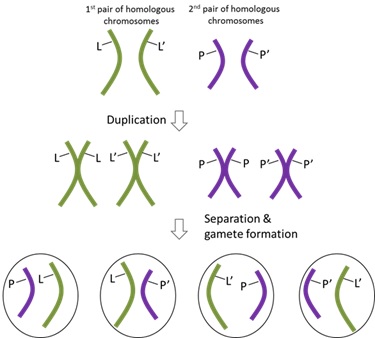
84 The mechanisms that lead to independent assortment are explained in Figure 12 below, which shows the production of gametes from one parent. If there are two alleles (with the mutated allele indicated by the presence of a ‘) of two genes (denoted as L and P) wherein the genes are on different chromosomes, there are four possible genotypes of gametes which can be produced (PL, P’L, PL’ and P’ L’). In this scenario, the two alleles of the genes are described as being unlinked because they are on different chromosomes, and the likelihood of inheriting a wild type P allele with a wild type L allele is the same as inheriting it with a mutant L’ (i.e. if a gamete inherits allele L, there is a 50% chance of inheriting a wild type allele P). This is because each allele is inherited independently of the other. In this scenario the two alleles are described as being unlinked because they are on different chromosomes.
Figure 12 – Unlinked alleles
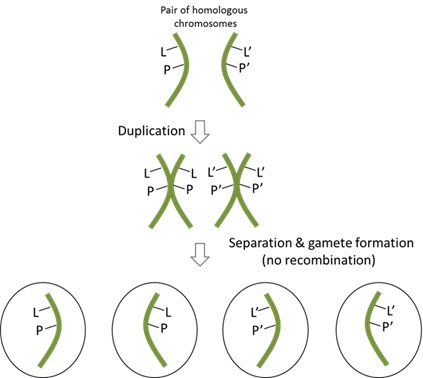
85 A different scenario is illustrated in Figure 13 below. In that scenario, the alleles of the genes are located on the same chromosome and there is no crossing over (or recombination) between them. Consequently, only two combinations of genotypes are produced (LP or L’P’). The wild-type alleles (L and P) are inherited together, and the mutant alleles (L’ and P’) are inherited together. In this scenario, the two alleles are described as being tightly linked.
Figure 13 – Linked alleles
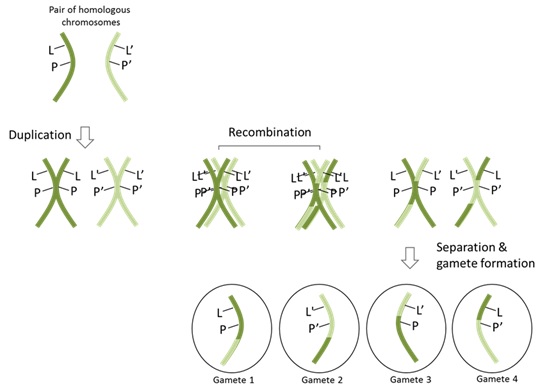
86 However, the scenario illustrated above of no recombination does not typically apply to alleles of genes on the same chromosome. Chromosomes undergo recombination during meiosis with their homologous pair. This process introduces the possibility that an allele at one locus on a chromosome will be inherited separately to an allele at a second locus on the same chromosome. If a recombination event happens between the two loci on the chromosome, and the homologous chromosome possesses a different allele, then the two alleles will be separated and a different haplotype will be transmitted to the progeny. This is illustrated in Figure 14 below. As can be seen, despite each chromosome having a particular combination of alleles, that combination of alleles will not always be inherited together. As can be seen in gametes 3 and 4, if a recombination event occurs between the loci of the L allele and the P allele, then the phase relationship between these two alleles is broken and the alleles can be inherited separately. In gamete 3, the L’ allele will be inherited with the P allele, while gamete 4 is the opposite. However, as can be seen in gametes 1 and 2, if a recombination event does not occur between the two loci, then the parental alleles will stay together on the same chromosome in the new gamete.
Figure 14 – Recombination
87 Consequently, alleles of genes that are linked (i.e. they are in close proximity on the same chromosome) will be inherited together with a higher probability than if they were not linked (i.e. there is a greater than 50% chance that alleles on the same chromosome will be inherited together).
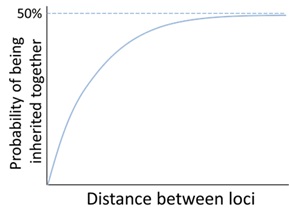
88 The probability of alleles at two loci on the same chromosome being inherited together is, in part, inversely related to their distance from each other, such that as the distance between the two loci increases, the chance of them being inherited together decreases, until the chance is the same as if they were on separate chromosomes (i.e. 50%). This is shown in Figure 15 below.
Figure 15 – The influence of distance on the probability of two loci being inherited together
89 Furthermore, as recombination happens during meiosis, the chance of alleles at two loci being separated increases for each successive generation. In other words, linkage disequilibrium (I will elaborate on this in a moment) between two loci decays each generation due to recombination. This is illustrated in Figure 16 below, which shows the decay of linkage disequilibrium (LD) across multiple generations for loci having a differing linkage (i.e. high, medium and low linkage). As can be seen, the higher the recombination fraction, the more quickly the linkage disequilibrium decays across multiple generations (c = recombination fraction, being the ratio of recombined gametes vs total gametes). The closer c is to 0.5, the less linkage there is between the two loci, that is, the more recombination happens between the loci.
Figure 16 – The decay of LD dependent on recombination between two loci
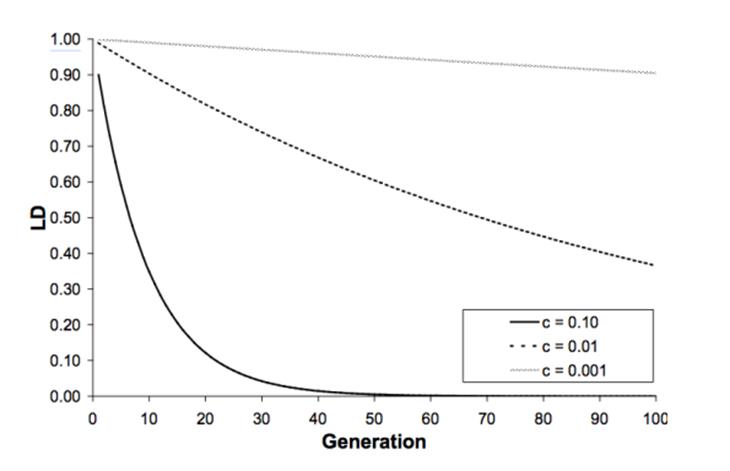
90 Before proceeding further I should note that there is a debate between the parties as to whether the information contained in Figure 16 and the immediately preceding statement concerning the c value of closer to 0.5 was part of common general knowledge as at the priority date.
Genetic linkage or distance
91 Genetic linkage is measured in a unit known as a centimorgan (cM). Two alleles which are one cM apart will have approximately a 99% chance of being inherited together.
92 But while a centimorgan is often used to infer a distance between loci, it is not a true measure of physical distance. The physical distance is measured by the number of nucleotides that separate two loci. The rate of chromosomal recombination per base varies throughout the genome. Certain regions of chromosomes have a higher propensity to recombine than other regions. As such, the physical distance between two loci that are one cM apart, will be lower if the loci are in an area with a high rate of recombination as compared with the distance between two loci in an area with a low rate of recombination.
Linkage disequilibrium
93 In Figures 12, 13 and 14 set out above, what is pictured is the genotype of a single animal and the gametes produced. The animal carries one chromosome with alleles L and P and a homologous chromosome with alleles L’ and P’. However, other animals in the population may have other genotypes. If across the whole population, a gamete that carries L also carries P or P’ according to the frequencies at which those alleles occur within the population, then the L and P loci are said to be in linkage equilibrium. Conversely, if a gamete carrying L is more likely to carry P, as compared to its frequency within the population, then the L and P loci are said to be in linkage disequilibrium (LD).
94 Another way to describe LD is to say that across the whole population, the genotypes at the L locus are not independent of the genotypes at the P locus. Consequently, if there is LD and you know an animal’s genotype at the L locus, it may help you to predict the genotype of that animal at the P locus.
95 The degree of LD can be measured by several statistics. One measure is the correlation between the alleles at the two loci (r). In relation to the above example, if r = 0 there is no LD (i.e. there is linkage equilibrium) and L is inherited with P at the frequency at which P occurs in the population. Contrastingly, if r = 1, then L’ is always carried by the same gametes as P’ and similarly the L and P alleles occur together.
96 Several factors can cause LD. One important factor is chance. If the effective size of the population is small, then only a small number of gametes form the next generation and a correlation between loci may arise in the sample simply by chance. And in each generation there is another chance sampling of gametes and r may increase or decrease.
97 Now although LD tends to build up in a small population by chance, recombination breaks down LD. Even if LD is complete (r = 1) recombination between L and P loci may generate the alternative gametes (i.e. LP’ and L’P) and therefore reduce LD. The speed with which recombination breaks down LD is shown in Figure 16 set out above. If the recombination rate is relatively small (i.e. c = 0.001 or 0.01), LD breaks down slowly. However, if the recombination rate is large (i.e. c = 0.10), LD breaks down at a relatively faster rate.
98 Generally speaking, the LD observed in populations is decreased if the effective population size is large. Contrastingly, the LD observed in populations is increased if the recombination rate between the loci is small. Further, other factors such as the physical distance between each locus, allele interaction effects and selection also affect LD.
(e) DNA genetic markers
99 A DNA genetic marker is considered to be any DNA sequence that varies from one individual to another. DNA is inherited in chromosomes. Genetic markers that are close to one another on a chromosome are more likely to be inherited together than genetic markers that are spaced further apart on a chromosome or are located on separate chromosomes. In this sense, markers act as landmarks indicating the inheritance of a specific region of DNA. When a relationship is established between a marker and a region that is known to influence a trait (i.e. a QTL as I will explain in a moment), then the marker can be used as a landmark to help trace the inheritance of this QTL.
100 While any particular marker genotype can be present in several individuals, a combination of genotypes at several markers may be unique to a specific individual, or family, and therefore can uniquely identify the genotype or phenotype of that individual/family. There are many different types of markers used in genotyping animals. However, all are predicated on one of three changes in DNA: (i) insertions/deletions of DNA (indels); (ii) point mutations (single nucleotide polymorphisms) or (iii) duplications (repeats) of DNA (both repeats of small areas of DNA or large chunks of DNA).
QTL
101 A quantitative trait locus (QTL), as I have referred to in the glossary, is a stretch of DNA that correlates with the genetic value and hence phenotype of an animal for a specific quantitative trait. The QTL is typically in LD with, or contains, gene(s) that influence the trait.
102 Genetic markers which are themselves in LD with a QTL influencing a trait can be used to identify animals which are more likely to possess the desired allele at the QTL and therefore more likely to possess and/or produce offspring with a desirable value for the trait. Such an approach does not require the genes influencing the trait to be identified, nor does the identified marker (mutation) need to be a causative mutation(s). Rather, this approach relies on identifying markers that are in LD with the causative mutations.
Microsatellites
103 A microsatellite marker, as I have referred to in the glossary, is a region of repetitive DNA where certain DNA motifs (short sequences of DNA, typically 2 to 5 bp long) are repeated (generally 3 to 50 times).
104 As a microsatellite can possess alleles that vary from a few repeats to 50 or more repeats of short sequences, these markers are known as multi-allelic markers, that is, markers with more than two possible options. As such, these markers can provide more information than a bi-allelic marker as there are many more possible variants, that is, one individual may have 5 and 7 repeats defining the two alleles present in its genotype, and another may have 6 and 11 repeats defining the two alleles present in its genotype.
Single nucleoside polymorphisms (SNPs)
105 As I have already explained in the glossary, SNPs are single nucleotide changes in the DNA sequence at a specific locus in the genome. There are millions of SNPs in any given genome. SNPs are considered to be bi-allelic in that there is an ancestral allele and a derived allele.
(f) Animal genomics and the genetic improvement of livestock
106 Genomics is the study of the genome of individuals to understand the structure of the DNA that encodes the genetic blueprint of the individual. Animal breeders and researchers are interested in how variation in the genome is associated with variation in the traits expressed by an animal.
107 The way an animal looks and performs is described as its phenotype. This includes not only physical characteristics such as colour or whether it has horns, but also how fast it grows, the composition of its milk or carcass (i.e. the quality of its meat), and how well it copes with stress and disease challenges. Phenotype is determined both by the genotype of the animal (its genome sequence) and its environment. The interaction between genotype and environment complicates how researchers can identify and select the best animals to breed the next generation to be better than the previous one.
108 By measuring the phenotype of an individual, and its relatives including its siblings and offspring, it is possible to obtain information on the breeding value of an animal or how well its offspring will perform compared to other individuals in the population. Although modern animal breeding uses advanced statistical methodologies such as Best Linear Unbiased Prediction (or BLUP) and very large amounts of computing power, it is still based on tenets such as crossing the best with the best. In this regard, being able to identify what makes them the “best of the best” genotypically, that is, selection based on genetics rather than selection of a specific genotype, and selecting for those traits has large commercial value for those in the livestock industry who want to breed for desirable traits. Further, the genotype (or genetic information) can provide information on the potential performance of animals in order to optimise their management and improve economic returns.
109 Different techniques have been available to study the contribution of different genes and their individual alleles to quantitative traits. Allelic variation can contribute to, or be linked to, a change in a trait. By knowing such information in the context of animal sorting and breeding, one can endeavour to identify animals having desirable genes (or more strictly alleles) to then form a view as to whether an animal has a desirable trait, e.g. meat tenderness, milk production, etc.
110 There were various different approaches identified in the evidence of Professor Michael Goddard called by MLA and Professor Graham Plastow called by Branhaven. It is useful to summarise them at this point. For the most part, they can be taken to be approaches used as at the priority date. But there was considerable debate with respect to approaches 5 and 6 in terms of what was known about them, how they were described and how they were applied (if at all) as at the priority date. For present purposes, I will postpone that debate concerning approaches 5 and 6 and return to it later.
Approach 1 – phenotype selection/selective breeding
111 The first approach may be called the phenotypic selection approach or selective breeding. In 2002, a standard model for this approach was to estimate the genetic parameters for the trait of interest and to use these to calculate estimated breeding values (EBVs) for animals that were candidates for selection as breeding animals.
112 For example, if the trait of interest was growth rate, one would estimate the heritability of that trait and then calculate EBVs for all animals of interest. This would involve selecting a group of animals and their relatives and measuring the growth rates of these animals. This would provide data on the phenotypes of the animals. One would also have the pedigree relationships of those animals. One could take this data on phenotypes and pedigrees and estimate the heritability of the trait, usually by the statistical method called restricted maximum likelihood (REML). Using this heritability estimate and by employing Best Linear Unbiased Prediction one would be able to obtain an EBV for the trait of interest for each animal. One could then select those animals with the most desirable EBVs to become parents for the next generation.
113 This technique did not involve any DNA analysis. Whilst it generally worked well, there were a number of limitations to the approach. For example, certain traits could not be easily measured across an animal population. A classic example is selecting for milk yield in dairy cattle. One could not select bulls based on their own milk yield. Instead, selection had to be made on the basis of information obtained from female relatives such as daughters. That worked, but it took a relatively long time.
114 In addition, this approach did not work well in respect of traits that had a very low heritability. For such traits, environmental factors may have had a large effect on phenotype. Further, the approach also did not work well for numerous traits that could only be measured in old age, such as longevity or traits which could only be measured on dead animals, such as traits relating to meat quality.
Approach 2 – candidate gene approach
115 The candidate gene approach to conducting genetic association studies focuses on determining associations between genetic variation within pre-specified genes of interest. Candidate genes are selected for study based on prior knowledge of the gene’s biological function and its relationship to the trait of interest, typically based on knowledge of analogous genes in other species.
116 To take the candidate gene approach, it is first necessary to know of the existence of one or more specific genes that affect the trait of interest. It is then necessary to identify a polymorphism in that gene. The polymorphism could be a SNP. Other types of polymorphisms are a deletion or insertion of one or more nucleotides or some other difference between the sequence of one copy of the gene and another.
117 One would then carry out experiments to ascertain whether animals with different genotypes at this polymorphism differed significantly in phenotype for the trait of interest. This is termed an association study. Polymorphisms which showed a correlation with the trait of interest would be combined. A multiple regression analysis would need to be carried out where the dependent variable was the trait of interest and the independent variables were the genotypes at the polymorphisms deemed to be associated with the trait. This would give you an EBV for each animal based on the genotypes at these selected polymorphisms.
118 But key limitations of this approach included the following. First, this approach was limited by its reliance on existing knowledge about known or theoretical biology of different traits. For example, a gene known to be associated with bone production was later discovered to also be associated with the occurrence of twinning in sheep. Second, because most traits are influenced by many different genes, each contributing a small amount to the overall trait, identifying the independent contribution of a single genetic variant in the pathway may be difficult unless the study population was relatively large. Third, as this process tends to be on a gene by gene basis, it could be very slow. Fourth, because the search for variants may not be limited to a specific region of investigation, a study’s ability to detect an association depends upon a number of different factors in addition to the function of the gene. For example, Professor Plastow worked on a project where they had identified a gene from the correct gene family, but not the correct gene. The one they were analysing mapped in a completely different region of the genome to the location of the gene involved in variation of the trait.
119 A classic example of where the candidate gene approach has been applied to identify superior animals for breeding has been in respect of a trait that is controlled by a single gene such as an abnormality caused by a recessive mutation. In this instance, there are two alleles with one being dominant and the other being recessive. If both recessive alleles are inherited, the animal may express an abnormality in the trait or the animal may not be viable. If both a dominant and recessive allele is inherited, then the abnormality is not expressed but the animal is a carrier for the condition and may pass it on to its progeny. But although the candidate gene approach often worked with single gene abnormalities, it seldom worked with complex or quantitative traits. In respect of most of the traits of economic importance, the candidate gene approach was considered by some quantitative geneticists in animal breeding at the priority date not to be usually successful. This is for a number of reasons. First, even though a gene may have a role in the physiology underlying a trait, a genetic variation in the gene may not affect the trait. Second, even if there is genetic variation in the candidate gene, it usually has a small effect on such traits at most. This is because most traits of economic importance are usually complex traits controlled by many genes and environmental factors. Third, many genes that contain polymorphisms affecting a trait were not obvious candidates at the time.
120 More generally, prior to the priority date, it was found that many candidate associations in respect of complex traits were not statistically significant, and for some of those that were significant the effect of the polymorphism was over-estimated. This in turn caused some reported associations not to be replicated in future experiments and it caused some inaccurate selection of breeding stock when used. Furthermore, the associations that were found in some cases explained little of the genetic variance for the trait and so might not significantly increase the accuracy of selection even if their effect was estimated accurately.
Approach 3 – Within family QTL linkage approach
121 A QTL analysis usually involved crossing two or more groups of animals that differed genetically with regard to the trait of interest. For example, dairy cattle might be bred to beef cattle with the aim of mapping QTL for milk production and muscle traits. Using molecular markers to distinguish between the parental lines, the offspring having the desired traits were assessed against those that did not. Markers that were genetically linked to a QTL influencing the trait of interest would segregate more frequently with trait values, whereas unlinked markers would not show significant association with phenotype. Looking within one family in the breeding experiment (a male, female, their offspring and descendants), if the desired trait was always more prevalent with a certain interval or haplotype of markers and always lower with another haplotype, then it could be concluded that in all likelihood the gene for the trait was in the first mentioned region of the chromosome. But this approach was limited in a number of ways. First, the number of animals used tended to only be a few hundred and the effects needed to be relatively large to be detected. Second, there was a problem in the specificity that could be achieved. Referring to the analogy set out at the start of my reasons, the QTL could tell you which state you were in, but not which city or the street address. You could not specify which genes or individual polymorphisms were causal without combining it with another technique to enable fine mapping such as the candidate gene approach. In this case the information was on the location of genes, often using comparative information between species. For example, region X in bovines might be equivalent to region Y in humans, and so information on region Y as to the human genes known there might enable you to make a judgment call on the bovine genes likely to be in region X. Third, because the QTL regions were relatively large, they could contain several QTL making the analysis complicated. In addition, they were often “marked” by microsatellites which were difficult to use for routine application. In both cases the linkage disequilibrium between the markers and the underlying genes (and the causative alleles or variants) were such that the association might be different in different families. This meant that in one family the gene appeared in one “state”, and another in a different “state”. This would also change over time due to recombination or shuffling of the different segments during the production of gametes.
Approach 4 – marker assisted selection
122 The fourth approach is termed marker assisted selection (or MAS). Such an approach makes use of random genetic markers that are on the same chromosome as an unknown polymorphism causing variation in the trait of interest. The marker is said to be “linked” to the polymorphism. Any type of marker such as microsatellites or SNPs could be used in marker assisted selection.
123 Such an approach could be carried out by undertaking an analysis of a large number of individuals within a family. For example, if there were several hundred offspring of a single bull (a common occurrence in farming), one would look for an association between a genetic marker (by way of DNA analysis of the animals) and a particular trait within those offspring (e.g. milk yield). A marker could be a SNP. If the SNP was an A to T polymorphism and individuals that received the A allele from their father produced more milk than individuals who inherited the T allele, this would establish linkage between this SNP and a polymorphism affecting milk yield. But it does not entail that the SNP is necessarily a biological cause of any change in that trait in the sense of a mutation in a gene responsible for that trait, but only that it is linked on the chromosome to a polymorphism that does cause a change in that trait.
124 To implement MAS in a breeding program, one would need to genotype other animals in this same family and select those that carried the allele associated with increased milk yield. But in a new breeding family, one would not know whether either of the A or T allele for the SNP would be a marker for improved milk yield. The linkage phase had to be determined for each family. The SNP and the QTL may not necessarily be in linkage disequilibrium.
125 Set out below is an illustration of this concept:
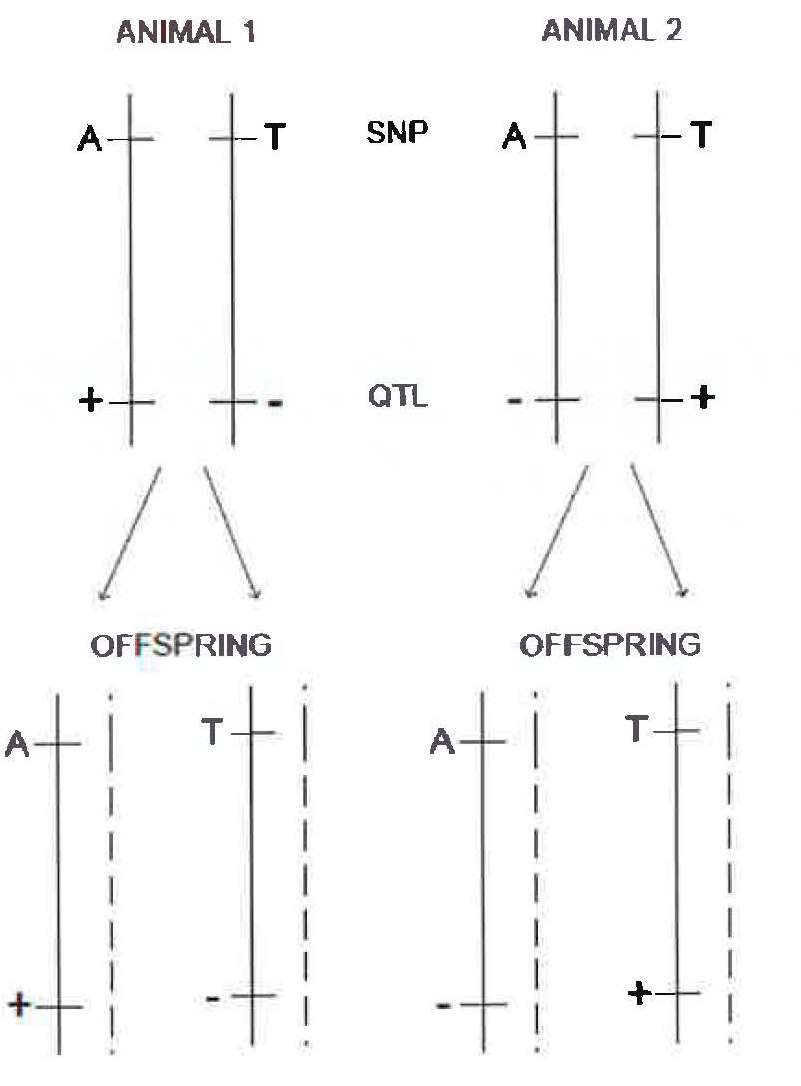
126 In the offspring of animal 1, the SNP allele A tends to be inherited with the “+” allele at the QTL as they are on the same chromosome. But in the offspring of animal 2, it is the T allele that tends to be inherited with the + allele. The SNP and QTL are linked. But across the whole population there is no linkage disequilibrium, e.g. no association between alleles at the SNP and alleles at the QTL.
127 MAS is based on the use of relatively few markers that are relatively sparsely spread across the genome. It is usually assumed that given the low number of markers, most will not be close enough to a gene affecting a trait to be relevantly in linkage disequilibrium.
Approach 5 – Genomic selection
128 The fifth approach is called genomic selection and is in some respects similar to MAS, except in terms of scale. Genomic selection makes use of random genetic markers, and it is based on having a larger number of genetic markers that are spread densely enough across the genome such that a gene for a trait of interest or a QTL will very likely be in linkage disequilibrium with at least one of the genetic markers, no matter where the gene or QTL is located on the genome. Further, the effect of all markers (and hence all the genes or QTLs) on the trait is estimated and provides an EBV. Contrastingly, the MAS approach attempts only to find a small number of markers linked to genes causing variation in a trait.
129 An advantage of the genomic selection approach is that once you can establish that there is linkage disequilibrium between one or more genetic markers and a polymorphic gene/QTL, you can then carry out the analysis across an entire population.
130 The first step in genomic selection is to identify a group of animals, designated the discovery population, and to carry out an association study on that population. This involves first measuring for at least one (but possibly many) trait of interest. The population of animals is then genotyped not for a single or small number of markers, but for many markers spread across the genome. An estimation is then made of the apparent effect of each marker on the trait. It is the apparent effect because many of the markers do not generally have a direct effect on the trait. But they are in linkage disequilibrium with a polymorphism that does have an effect on the trait.
131 One type of marker suitable for use in genomic selection is a SNP. For example, if one SNP has C and T alleles, then an animal will have one of the combinations of: CC, TT or CT. The genotype of an animal at this particular SNP can be described using the numbers 0, 1 or 2 (the number of T alleles) so that the CC genotype is designated 0, the CT genotype is designated I and the TT genotype is designated 2. The difference between these three genotypes in the trait is then estimated in the discovery population. Usually, the difference between CC and CT is assumed to be the same as the difference between CT and TT. Then the regression of the trait on the number of T alleles (0 or 1 or 2) is estimated. The regression coefficient estimates the difference between CC and CT genotypes. If the effect of each additional T allele is estimated to be +5, then the contribution of this marker to the EBV of the animal is obtained by multiplying this value (5) by the number of T alleles (0, 1 or 2). This process is undertaken for each marker using a multiple regression equation to obtain a predictive equation which uses marker genotypes as the input. The outputs are an estimate of the breeding value of each animal for each trait. The predictive equation so derived can then be tested or validated by applying it to a new group of animals with genotypes and phenotypes for the trait. This is the validation phase. It is carried out for the purpose of validating that the predictive equation is accurate.
132 The predictive equation can then be applied to a group of animals which have not been measured for phenotype, but whose genotype has been ascertained. This will generate EBVs for these animals that can then be used to select the best animals as parents for the next generation for a particular trait or a number of traits.
133 In performing the genomic selection approach, all the markers are treated impartially. It is not known which markers may be highly predictive on their own of a trait and which are less predictive. Once one obtains the data, one is able to estimate the apparent effect of each marker. If a marker is in high linkage disequilibrium with a causal mutation with a large effect on the trait, then the marker will have a large apparent effect on the trait. If the size of the discovery population is large enough, this apparent effect might be statistically significantly different from zero even when a stringent significance test is used. But one does not usually restrict the predictive equation only to markers that pass a significance test. Other markers may have a small effect and individually these markers might not pass a significance test. But the combined effect of many markers of small effects may be assumed to be important. Hence all markers are taken into account.
Approach 6 – Genome wide association studies (GWAS)
134 Genome wide association studies or GWAS investigate the entire genome. The approach is non-candidate-driven in contrast to gene-specific candidate-driven studies. The key limitations of this approach include:
GWAS identify SNPs and other variants in DNA which are associated with a trait, but, like the QTL approach, cannot on their own specify which genes are causal.
Such studies need a relatively large number of SNPs which are distributed reasonably evenly across the genome.
135 A GWAS is typically used for research purposes to find markers associated with traits. It identifies a set of potentially useful markers. A subset of the markers may subsequently be used for MAS.
The difference between approaches 5 and 6
136 The experts were questioned about what the known differences between the genomic selection (GS) approach and GWAS were as at the priority date.
MLA’s experts
137 MLA’s experts (Professor Michael Goddard, Professor Peter Visscher and Professor Benjamin Hayes) provided the following information.
138 Professor Goddard explained that both the GS approach and a GWAS would start by estimating the association between a panel of markers and the trait. Both could involve a test of which markers were associated with the trait. However, such a test was neither necessary nor sufficient if the aim was to identify cattle with superior genetic potential (which he gave as the ultimate purpose of the GS approach). If the results of a GWAS were used in further steps to identify animals with superior genetic potential, it became an exercise in the GS approach. The GS approach estimated the effects of a panel of markers on the trait (such as could be derived from a GWAS) and then combined these estimates with the genotypes of individual cattle to predict their genetic potential.
139 Professor Visscher considered the main difference between the approaches to be one of purpose. The purpose of a GWAS was to detect associations between a trait and one or more genetic markers. A GWAS did not need to be followed by a prediction/selection step to achieve this. By comparison, the purpose of the GS approach was to estimate genetic potential using GWAS data, for the use in breeding programs.
140 Professor Hayes described GS as an approach for predicting genetic value or phenotypes of animals directly from DNA markers, whilst a GWAS identified a set of useful markers that might then later be used in marker assisted selection. In terms of the methods employed, a GWAS required association testing of markers one by one, whereas the GS approach used all markers simultaneously. The requirement for association testing in a GWAS would mean that only a limited number of markers would be used, accounting for a limited proportion of the genetic variance. The number depended upon the significance threshold set for association testing and a number of other factors. When the markers and their effects derived from a GWAS were used in marker assisted selection, there was a problem known as the Beavis effect, where the effect of the most significant markers are over-estimated. This problem was more pronounced with small sample sizes and eroded the accuracy of marker assisted selection. The GS approach did not suffer from this problem, as all markers were fitted in the model simultaneously, as opposed to selecting the best ones.
Branhaven’s experts
141 Branhaven’s experts (Professor Graham Plastow, Professor Jeremy Taylor and Dr Tad Sonstegard) provided the following information. And if it is necessary to say so, I found Professor Plastow’s explanation the most helpful of the experts on this aspect.
142 Professor Plastow explained that the GS approach was based on using a panel of sufficiently dense markers that some of them will be in LD with causative mutations. A GWAS by comparison was typically used for research purposes and to find markers associated with new traits. Typically these GWAS results were used to identify what markers and linked genes might explain variation in the traits. The analysis might use the same tools as GS or other additional tools to identify markers that are associated with the trait of interest. The resulting markers could then be used by including them in “marker assisted selection” or “marker assisted management” to identify cattle that could be sorted to different endpoints. These steps would not typically be taken with the GS approach where all of the markers were used for the subsequent steps (e.g. calculating breeding values). Professor Plastow’s understanding was that the GS approach nearly always used the same SNP panel for every stage, whereas in a GWAS the subsequent steps nearly always used a smaller number of SNPs.
143 Professor Taylor explained that the GS approach, in general, did not require statistical tests for individual markers to assess their strength of trait association. But such tests were an important part of a GWAS, which implicitly ranked the SNPs based on their strengths of association with the trait, which would have consequences for the design of assays for downstream genotyping. The GS approach attempted to predict the effect of all variants responsible for trait variation, whereas GWAS only attempted to predict merit based on the largest effect variants. Professor Taylor also noted that GWAS could be performed using selective genotyping whereas the GS approach could not.
General
144 But there was a degree of overlap between the two approaches. Both approaches had the same starting point of panels of genome wide markers, employed similar experimental designs and broadly fell within the category of “marker assisted selection” methods. Indeed, Professor Goddard characterised the GS approach as the application of GWAS data to the estimation of breeding values, stating:
So whether you regard it as a GWAS or whether you regard it as a genomic selection experiment, it actually looks the same. You genotype the animals for a genome-wide panel of markers; you measure them for the traits; and then that’s where the GWAS stops, really. It does the association and it stops, but if you’re going to use that information then to estimate breeding values on animals, it looks just like genomic selection.
145 In general terms of what was known as at the priority date, it would seem that the definitions and boundaries of approaches 5 and 6 were not that well defined; indeed there was considerable linguistic imprecision in how the experts described these approaches before me. Moreover, approaches 5 and 6 were not applied as at the priority date in relation to bovines.
The 253 Application
146 The specification is headed “Compositions, Methods and Systems for Inferring Bovine Traits”.
147 The specification describes the invention in general terms as providing methods, compositions and systems for managing, selecting, breeding and cloning cattle utilising information regarding genetic diversity among cattle, particularly single nucleotide polymorphisms (SNPs) and the effect or potential effect of nucleotide occurrences of SNPs on important traits.
148 The 253 Application is expressed to be a divisional application based, as I have said, on the parent Australian patent application no 2003303599 filed on 31 December 2003, the entire content of which is said to be incorporated by reference into the 253 Application. The 253 Application also claims the benefit of priority of US patent application No 60/437,482 filed on 31 December 2002 (the US priority document), the entire content of which is also said to be incorporated by reference into the 253 Application.
149 The claims defining the invention are set out in the following terms (pages 67 to 69):
1. A method for identifying a trait of a bovine subject from a nucleic acid sample of the bovine subject, comprising identifying in the nucleic acid sample an occurrence of at least three single nucleotide polymorphisms (SNPs) wherein the at least three SNPs are associated with the trait, and wherein the at least three SNPs occur in more than one gene [:]
[a] and wherein at least one of the SNPs corresponds to position 300 of any one of SEQ ID NOS: 19473 to 21982, or
[b] the SNP is about 500,000 or less nucleotides from position 300 of any one of SEQ ID NOS: 19473 to 21982.
2. The method of claim 1, wherein the at least one SNP is selected from at least one of the SNPs and nucleotide occurrences indicated in Table 1A.
3. The method of claim 1 or 2, wherein occurrences of at least five SNPs are identified.
4. The method of claim 1 or 2, wherein occurrences of at least seven SNPs are identified.
5. The method of claim 1 or 2, wherein occurrences of at least ten SNPs are identified.
6. A method for sorting one or more bovine subjects, comprising:
(a) identifying a trait for a first bovine subject from a nucleic acid sample of the first bovine subject, by a method comprising identifying an occurrence of at least three single nucleotide polymorphisms (SNPs) wherein the at least 3 SNPs are associated with the trait, and the at least three SNPs occur in more than one gene, wherein at least one SNP corresponds to position 300 of at least one of SEQ ID NOS: 19473 to 21982, or the SNP is about 500,000 or less nucleotides from position 300 of any one of SEQ ID NOS: 19473 to 21982; and
(b) sorting the first bovine subject based on the identified trait, and optionally repeating steps (a) and (b) for additional subjects.
7. A method for cloning a bovine subject with a desired trait, comprising:
(a) identifying an occurrence of at least three single nucleotide polymorphisms (SNPs) wherein the at least 3 SNPs are associated with the trait, and the at least three SNPs occur in more than one gene, wherein at least one SNP corresponds to position 300 of one of SEQ ID NOS: 19473 to 21982, or the SNP is about 500,000 or less nucleotides from position 300 of any one of SEQ ID NOS: 19473 to 21982;
(b) isolating a progenitor cell from the bovine subject; and
(c) generating a cloned bovine from the progenitor cell.
8. A method of inferring a trait in a bovine, comprising hybridizing a nucleic acid sample from a bovine subject to a system wherein the hybridizing comprises: a substrate; and specific binding pair members corresponding to a series of at least three SNPs, wherein the at least three SNPs are associated with a trait in the bovine subject, and the at least three SNPs occur in more than one gene, wherein at least one of the SNPs corresponds to position 300 of any one of SEQ ID NOS: 19473 to 21982, or the SNP is about 500,000 or less nucleotides from position 300 of any one of SEQ ID NOS: 19473 to 21982, where selective hybridization of the nucleic acid sample to the system indicates the presence of a SNP associated with a trait.
9. The method of any one of the preceding claims, wherein the trait is marbling, tenderness, quality grade, muscle content, fat thickness, feed efficiency, red meat yield, average daily weight gain, disease resistance, disease susceptibility, feed intake, protein content, bone content, maintenance energy requirement, mature size, amino acid profile, fatty acid profile, milk production, a milk quality susceptibility to the buller syndrome, stress susceptibility and response, temperament, digestive capacity, production of calpain, caplastatin and myostatin, pattern of fat deposition, ribeye area, fertility, ovulation rate, conception rate, fertility, or susceptibility to infection with and shedding of pathogens.
10. The method of any one of the preceding claims, wherein the trait is fat thickness, retail yield, tenderness, marbling, or average daily gain.
11. A bovine subject resulting from the method of claim 7.
12. A system for determining nucleotide occurrences of nucleotide polymorphisms (SNPs) in a bovine population comprising:
a hybridisation substrate that includes specific binding pair members corresponding to a series of at least 3 SNPs, wherein
the series of SNPs are associated with a trait from a bovine;
the SNPs occur in more than a single gene; and
at least one SNP corresponds to position 300 of any one of SEQ ID NOS: 19473 to 21982; or
the SNP is about 500,000 or less nucleotides from position 300 of any one of SEQ ID NO: 19473 to 21982.
13. An isolated polynucleotide identified according to the method of claim 8.
14. An isolated polynucleotide when used in any one of methods 1 to 10 comprising:
(a) a fragment of at least 20 contiguous nucleotides of a bovine genome, or
(b) a complement of (a);
wherein the isolated polynucleotide of (a), or (b), comprises a nucleotide occurrence of a single nucleotide polymorphism (SNP) associated with a trait, wherein the SNP is about 500,000 or less nucleotides from position 300 of any one of SEQ ID NOS: 19473 to 21982, and wherein the isolated polynucleotide is less than or equal to about 500,000 nucleotides.
15. The method of any one of claims 1 or 6 to 8, substantially as hereinbefore described.
150 At this point it should be noted that the parties and the witnesses before me have, with respect to claim 1, referred to the integer “and wherein at least one of the SNPs corresponds to position 300 of any one of SEQ ID NOS: 19473 to 21982” as limb (a). The other integer, which is disjunctive relative to limb (a), “the SNP is about 500,000 or less nucleotides from position 300 of any one of SEQ ID NOS: 19473 to 21982”, has been referred to as limb (b). I will use similar short hand references from time to time in the following sections of my reasons. For convenience, in the text of claim 1 set out above I have inserted “[a]” and “[b]” notations together with “[:]” before the start of those integers.
151 Before proceeding further I would note the following:
(a) First, MLA’s principal attack is that the alleged invention in each claim is not a manner of new manufacture (ground of appeal 13). It is said that the alleged invention is not the proper subject matter of a patent because the claims are to known methods that use gene sequences (SNPs) that are naturally occurring and the claims include the use of gene sequences that are not defined by reference to a sequence identified in the 253 Application nor to any sequence at all. Further, it is said that the claims are to mere desiderata because they are not so defined and, further, the claims include gene sequences that are yet to be discovered, identified or created. Further, it is said that the grant of a patent with respect to such methods is or would be “generally inconvenient” in that the indeterminate scope of the claims would prevent or hinder research and development of methods to identify or infer traits in bovine subjects using SNPs, contrary to the public interest. The other dimension to MLA’s attack on manner of manufacture is that it says that it is apparent on the face of the specification that the alleged invention does not satisfy the threshold test for patentability. It is said that the specification makes clear that all the inventors did was to discover naturally occurring bovine SNPs and create a high density map of the bovine genome based on SNP markers that were evenly distributed within the putative bovine genome using the assembled human genome as a scaffold (all by known methods), discover naturally occurring associations between the specified SNPs and bovine traits, and identify SNPs that were in linkage disequilibrium with the specified SNPs according to known methods.
(b) Second, MLA says that each claim is not a patentable invention because it lacks utility (ground of appeal 10). It is said that the promise of the invention was to provide a method of distinguishing between randomly-chosen animals, in particular bovines, for a particular trait or the promise said to be embodied in [0101] of the specification, which I will set out in a moment. But it is said, inter-alia, that because none of the specified SNPs have been validated it is more likely than not that something falling within each of the claims would not be able to provide the basis for an inference or identification of the trait purportedly associated with the specified SNPs in a bovine.
(c) Third, MLA makes a novelty attack (grounds of appeal 1 to 4). The attack depends upon a proper identification of the priority date. As I have said, the 253 Application derives from the parent application filed on 31 December 2003, which in turn claims priority from the US priority document filed on 31 December 2002. In my reasons, where I refer to the “earliest priority date” I am referring to 31 December 2002, and where I refer to the “deferred priority date” I am referring to 31 December 2003, unless I state otherwise; further, where I simply refer to “priority date” I am embracing either option unless I state otherwise. Now the 253 Application was filed on 1 June 2010, but the claims were amended by a statement of proposed amendments submitted on 17 October 2013 (the October 2013 amendments). Accordingly MLA says that the priority date is 17 October 2013, a suggestion I easily dismiss, alternatively 31 December 2003. If the priority date is 17 October 2013, then it is said that claims 1 to 6, 8 to 10 and 12 to 15 have been anticipated. If the priority date is 31 December 2003, then it is said that claims 1 to 6, 9, 14 and 15 have been anticipated. If the priority date is 31 December 2002 (Branhaven’s position) then it is said that claims 1 to 6, 9, 14 and 15 have been anticipated.
(d) Fourth, MLA says that the invention as defined by each claim lacks an inventive step (grounds of appeal 5 to 7). I do not need to descend into the detail for the moment, except to make the point that there are different relevant prior art bases relied upon by MLA depending on the correct priority date.
(e) Fifth, MLA asserts a lack of sufficiency (ground of appeal 8). There are three dimensions to the attack: (i) It is said that the claims are so unclear that a skilled person cannot work the invention; (ii) Further, it is said that the 253 Application fails to provide sufficient information for a person skilled in the art to perform the claimed invention without undue experimentation or prolonged study of matters presenting initial difficulty due to the failure to validate the specified SNPs or the non-specified SNPs; (iii) Further, it is said that on the assumption that each claim can be divided into two claims (“alternatively, inventions”) with the second concerning what I have described earlier as integer (b) (claim 1), then the specification lacks a sufficient description of how to make something falling within the ambit of the claims concerning limb (b). It is said that there is no sufficient guidance to determine whether a SNP or group of SNPs was a relevantly non-specified SNP without undue experimentation or prolonged study of matters presenting initial difficulty.
(f) Sixth, MLA asserts that each claim lacks clarity (ground of appeal 12). In addition to asserting a lack of clarity based upon the last mentioned point, it is said that, inter-alia, the following terms or phrases are unclear:
(i) “a method for identifying a trait of a bovine subject”;
(ii) “the at least three SNPs are associated with the trait”;
(iii) “wherein the at least three SNPs occur in more than one gene”; and
(iv) “the SNP is about 500,000 or less nucleotides from position 300 of any one of SEQ ID NOS 19473 to 21982”.
(g) Seventh, MLA asserts that the complete specification does not end with a claim or claims defining the invention (ground of appeal 11). For this purpose MLA advances arguments relied upon relating to lack of clarity and, in part, lack of sufficiency.
(h) Eighth, MLA also asserts a lack of fair basis in relation to each claim (ground of appeal 9). It is said, inter-alia, that to the extent that each claim encompasses specified SNPs (I have previously described this as limb (a) (claim 1)) that are associated with a trait other than the five traits for which associations have been identified in the specification, the claims travel beyond the matter described in the specification. Further, it is said that to the extent that each claim encompasses non-specified SNPs (I have previously described this as limb (b) (claim 1)), the claims travel beyond the matters described in the specification because the specification only describes 2,510 specified SNPs associated with bovine traits. By reason of limb (b), MLA says that each claim extends to any SNP that is “about 500,000 or less nucleotides from position 300 of any one of” the specified SNPs. Moreover, it says that each claim only requires that one of the “at least three SNPs associated with the trait” be a specified SNP or a non-specified SNP. As a consequence, so it is said, the other two SNPs can be more than about 500,000 nucleotides from the specified SNPs. In other words they can be anywhere in the bovine genome. A further dimension to the lack of fair basis challenge is that it is said that the specification provides no real and reasonably clear disclosure of the combination of integers comprising: (i) at least three SNPs; (ii) in more than one gene; and (iii) wherein at least one of the SNPs is selected from the SNPs defined by the claims.
152 Before the delegate, I should note that MLA’s opposition only succeeded in relation to a lack of clarity and a manner of manufacture point concerning claim 13. The delegate rejected MLA’s opposition concerning lack of inventive step, lack of sufficiency, lack of fair basis, lack of clarity (save for claim 13) and manner of manufacture (save for claim 13) (see the delegate’s reasons at [134] to [136]). MLA did not press its opposition before the delegate on lack of novelty or lack of utility. I would also make some other observations. The case put before me was considerably more expanded than that put before the delegate. Not only were new grounds of lack of novelty and lack of utility run before me, but the other grounds were elaborately reworked (see the third amended notice of appeal of 17 pages). Further, MLA did not call before the delegate or tender statements from the three quantitative geneticists that it called before me, namely, Professor Michael Goddard, Professor Benjamin Hayes and Professor Peter Visscher. In relation to Branhaven, it tendered a statement before the delegate from Professor Graham Plastow (a molecular geneticist) who it called before me. But it also called before me Dr Tad Sonstegard (a molecular geneticist) and Professor Jeremy Taylor (a quantitative geneticist), but no statements from these experts were put before the delegate. Of course, the appeal before me was not required to be (and was not) confined to the legal grounds and forensic landscape before the delegate.
153 It is convenient to now discuss the specification in more detail.
The field of the invention and background
154 Both the field of the invention and background information are described at [0002] to [0022] (pages 1A to 9). It is appropriate to set out [0002] to [0006], [0009] to [0014] and [0022]:
FIELD OF THE INVENTION
[0002] The invention relates generally to gene association analyses and more specifically to polymorphisms and associated traits of bovine species.
BACKGROUND INFORMATION
[0003] Under the current standards established by the United States Department of Agriculture (USDA), beef from bulls, steers, and heifers is classified into eight different quality grades. Beginning with the highest and continuing to the lowest, the eight quality grades are prime, choice, select, standard, commercial, utility, cutter and canner. The characteristics which are used to classify beef include age, color, texture, firmness, and marbling, a term which is used to describe the relative amount of intramuscular fat of the beef. Well-marbled beef from bulls, steers, and heifers, i.e., beef that contains substantial amounts of intramuscular fat relative to muscle, tends to be classified as prime or choice; whereas, beef that is not marbled tends to be classified as select. Beef of a higher quality grade is typically sold at higher prices than a lower grade beef. For example, beef that is classified as “prime” or “choice”, typically, is sold at higher prices than beef that is classified into the lower quality grades.
[0004] Classification of beef into different quality grades occurs at the packing facility and involves visual inspection of the ribeye on a beef carcass that has been cut between the 12th and 13th rib prior to grading. However, the visual appraisal of a beef carcass cannot occur until the animal is harvested. Ultrasound can be used to give an indication of marbling prior to slaughter, but accuracy is low if ultrasound is done at a time significantly prior to harvest.
[0005] Currently there are no cost effective methods for identifying live cattle that give accurate prediction of the genetic potential to produce beef that is well-marbled. Such information could be used by feedlot operators to identify animals for purchase prior to finishing, to identify animals under contract for one or more premium programs administered by a packer, by feedlot managers to make management decisions regarding individual animals within a lot (including nutrition programs and sale dates), by cow-calf producers in marketing their animals to various feedlots or in making decisions regarding which animals will be sold on various carcass evaluation grids. Such information could also be used to identify cattle that are good candidates for breeding. Thus, it is desirable to have a method which can be used to assess the beef marbling potential of live cattle, particularly young cattle well in advance of the arrival of the animal at the packing house.
[0006] Another characteristic of beef that is desired by consumers is tenderness of the cooked product. Currently there are no procedures for identifying live animals whose beef, if cooked properly, would be tender. Currently, there are two types of procedures which are used by researchers to assess the tenderness of meat samples after they have been aged and subsequently cooked. The first involves a subjective analysis by a panel of trained testers. The second type is characterized by methods used to cut or shear meat samples that have been removed from an animal and aged. One such method is the Warner-Bratzler shear force procedure which involves an instrumental measurement of the force required to shear core samples of whole muscle after cooking. Neither of these procedures can be used to any practical effect in a fabrication setting as the need to age product prior to testing would lead to maintenance of inventory of fabricated product that would be cost prohibitive. Consequently, the methods are used at research facilities but not at packing plants. Accordingly, it is desirable to have new methods which can be used to identify carcasses and live cattle that have the potential to provide beef that, if cooked properly, will be tender.
…
[0009] Modern day breeding programs in animal agriculture originated from fundamental observations made upon the first domestication of animals. Early humans observed differences in a broad range of characteristics between the offspring produced by mating different parents and they took advantage of this observation by only mating individuals that demonstrated the most desirable characteristics. By following this strategy for several generations our ancestors were able to create populations of animals that exhibited only desirable traits that best fit their needs. This strategy, called selective mating or selective breeding, is based on identifying the best progeny from one generation and making them the parents for the next generation. Selective breeding results in the development of individuals that are superior for one or more traits and is the backbone for modern day genetic improvement programs in animal agriculture.
[0010] Through the utilization of selective breeding strategies geneticists have been able to define the fundamental genetic parameters that influence the expression of traits. Breeding experiments revealed that some traits, like coat color, were expressed in a qualitative manner and could be easily passed onto the next generation while other traits, like growth rate or adult size, were expressed in a quantitative fashion and only small progress could be made at each generation. Subsequent research in the field of molecular genetics has now revealed that qualitative trait effects are caused by the action of a single gene while quantitative traits are caused by the action and interaction of many different genes.
[0011] In addition to contributions of genetics, it has been determined that genetic source alone did not account for all of the differences observed among groups of closely related individuals and that environment and management also played a role in determining the expression of specific traits …
[0012] The primary objective of any genetic improvement program is to ascertain the genetic potential of individuals for a broad range of economically important traits at a very early age. While the classical breeding approach has produced steady genetic improvement in livestock species it is limited by the fact that accurate prediction of an individual’s genetic potential can only be achieved when the animal reaches adulthood (fertility and production traits) or is harvested (meat quality traits). This is particularly problematic for meat animals since harvested animals obviously cannot enter the breeding pool. Furthermore, it is difficult to utilize the classical breeding approach for traits that are difficult (disease resistance) or costly (meat tenderness) to measure.
[0013] To overcome the previous problems with the classical breeding approach animal breeders and geneticists turned to the new fields of molecular genetics and genomics. These disciplines offered the promise that the underlying genes responsible for genetic variation of important traits could be identified. Targeted research programs were initiated to ascertain the location and functional differences of specific genes that contribute to genetic variation for defined traits. The primary goal of molecular breeding programs in livestock species is to develop genetic assays for economically important traits that can be tested on individual animals at an early age, can be used for traits that are difficult to measure, that provide an accurate estimate of an animals genetic potential for expression of the trait, and account for a large proportion of the total genetic variation observed for the trait in commercial populations.
[0014] To date, three different experimental approaches have been utilized to identify genes that effect economically important traits in livestock species: Candidate Gene Approach, Differential Gene Expression Approach, and Within Family Quantitative Trait Loci (QTL) Linkage Approach. Limited success has been achieved for each of these methods in identifying genes that contribute to genetic variation for defined traits. However, each method also has limitations, as the primary objectives of the molecular breeding approach described above have not been achieved. Accordingly, a need exists for methods that assist in a determination of the genetic potential of individuals for a broad range of economically important traits at a very early age. A description of each of the experimental approaches attempted thus far, and the limitations for each is outlined below:
…
[0022] In summary, three different experimental approaches have been used with limited success to identify genes, chromosomal regions or DNA markers that account for a large proportion of the genetic variation observed in economically important traits in livestock species. The results achieved from research programs utilizing these methods have not been widely utilized to date because they do not account for enough of the total genetic variation to allow accurate prediction of an animal’s performance for a specific trait. Furthermore, even when successful these approaches are only capable of identifying additive genetic components while ignoring non-additive genetic components such as dominance (i.e. dominating trait of an allele of one gene over an allele of another gene) and epistasis (i.e. interaction between genes at different loci) which are critical to the development of diagnostics that can be utilized by animal breeders to accurately predict genetic potential for economically important traits in livestock species.
155 It is appropriate to make a number of observations at this point.
156 First, under the heading “field of the invention”, it is stated that the invention “relates generally to gene association analyses and more specifically to polymorphisms and associated traits of bovine species”. In my view, the fields of quantitative genetics and molecular genetics are both embraced, with the asserted invention also to be considered and understood in that context. I will explain these two fields later.
157 Second, under the heading “background information”, the desirability of methods that can be used to identify cattle with the potential for traits such as marbling, tenderness, yield and growth rate is discussed.
158 Third, it is said that classical breeding approaches involve the selective mating or breeding of cattle. One such approach that was used before the priority date was progeny selection. Precise physical detection and analysis of DNA is not involved in progeny selection. Rather it involves making a prediction about an animal’s genetic merit based on its progenies’ performance. Information about the performance of a large number of cattle and information about their family history is necessary.
159 Fourth, there is a distinction to be made between qualitative traits and quantitative traits. Qualitative traits, such as coat colour, are identifiable as falling into one of a number of categories. Further, they may be caused by the action of a single gene. Quantitative traits, such as growth rate, show variation on a continuous scale between individuals. They can also be caused by the interaction of multiple genes. The limits of classical breeding approaches in predicting the potential for such traits are discussed.
160 Further, it is recognised that genetics alone cannot account for all the differences observed amongst groups of closely related individuals. Environment and management may also play a role in determining the expression of specific traits. Breeders using parameters that govern differences in the expression of specific traits between individuals enable them to evaluate the genetic makeup of different individuals within a population. This enables them to make steady progress in improving the expression of traits that have economic significance to the commercial production of livestock species.
161 Fifth, in their attempts to ascertain the genetic potential of cattle, there is a discussion of the shift in attention by animal breeders and geneticists to “the new fields of molecular genetics and genomics”. Three different approaches (and their limitations), which have been used to identify genes that affect economically important traits in livestock, are discussed.
162 The first approach discussed in the background section is the candidate gene approach. This involves targeting a specific gene based on the hypothesis that it may have an effect on a particular trait. This may be based on information known about genes that contribute to the trait in other species. The hypothesis is then tested. The DNA sequence of the target gene is looked at and it is assessed whether differences in sequence between individuals (i.e. polymorphisms) are associated with variations in the trait, or whether the location and functional differences of specific genes contribute to genetic variation for defined traits. This approach has been said to be used successfully in identifying genes and sequence variants that have an effect on a particular trait. But a number of practical and technical limitations are identified. The approach is largely based on information from published resources and effectively looks at a single gene at a time. This makes it cheaper to carry out than other approaches. The candidate gene approach could be combined with other approaches, such as QTL mapping.
163 The second approach discussed in the background section is the differential gene expression approach. The levels of expression of particular genes in certain tissues or under certain environmental conditions are looked at and inferences drawn as to which genes control a specific trait. Targeted genes are chosen based on knowledge of biochemical pathways or functions of genes in other organisms. This approach has been used to elucidate biochemical pathways and to understand cellular functions. But it is of limited utility in predicting the genetic potential of animals for specific traits.
164 The third approach discussed in the background section is within family QTL mapping. As previously described by me, QTL mapping is a technique that uses polymorphisms as genetic markers to map regions of the genome that are associated with quantitative traits. The “within family QTL mapping approach” involves small families of related individuals that have been bred-up or assembled. DNA samples are taken from all individuals in the population. Phenotypic measurements for the targeted traits are taken on the progeny. A set of polymorphic DNA markers is genotyped against the entire research population. Data from the population of animals is then analysed to determine whether variation in a particular marker or a linked set of markers is associated with variation in the trait. This approach, although being described as widely used, is subject to a number of limitations, for example, a lack of marker density, i.e. an insufficient number of genetic markers available to identify the precise location of the QTL on the chromosome.
165 Each of the three approaches is described as having limited success. The experts before me agreed that each of these approaches was used before the priority date, but there was some disagreement between the experts as to the degree of success of each of these approaches at that time.
166 Sixth, the background section stated that “a need exists for methods that assist in a determination of the genetic potential of individuals for a broad range of economically important traits at a very early age” (at [0014]). In other words it was perceived (or on one view “known”) that there was a need for more accurate methods to predict genetic potential. Indeed it is noted that the three approaches discussed in the background section (and summarised above) do not “allow accurate prediction of an animal’s performance for a specific trait” (at [0022] and see also [0005]). Contrastingly, later in the specification it is stated that “[t]his invention identifies animals that have superior traits, predicted very accurately …” (at [0101]); I will postpone for the moment discussion of the “very accurately” reference and its significance.
167 I will return to the question of the promise of the invention later in my reasons (see [0022] and [0101] of the specification) as it is relevant to, inter-alia, MLA’s assertion of lack of utility.
168 In summary, the background section summarises the limitations of the approaches that were available before the priority date. It emphasised the need for new approaches to the genetic improvement of cattle.
Summary of the invention
169 Although the summary of the invention and general statements dealing with some of the embodiments are set out from [0023] to [0030] (pages 9 to 11), it is convenient to set out only [0023] to [0025], [0027] and [0029] to [0030]:
[0023] The present invention provides methods, systems, and compositions that allow the identification and selection of cattle with superior genetic potential for desirable characteristics. Accordingly, the present invention provides methods, compositions, and systems for managing, selecting and mating, breeding, and cloning cattle. These methods for identification and monitoring of key characteristics of individual animals and management of individual animals maximize their individual potential performance and edible meat value. The methods of the invention provide systems to collect, record and store such data by individual animal identification so that it is usable to improve future animals bred by the producer and managed by the feedlot. The methods, compositions, and systems provided herein utilize information regarding genetic diversity among cattle, particularly single nucleotide polymorphisms (SNPs), and the effect of nucleotide occurrences of SNPs on important traits.
[0024] The present invention further provides methods for selecting a given animal for shipment at the optimum time, considering the animal’s genetic potential, performance and market factors, the ability to grow the animal to its optimum individual potential of physical and economic performance, and the ability to record and preserve each animal’s performance history in the feedlot and carcass data from the packing plant for use in cultivating and managing current and future animals for meat production. These methods allow management of the current diversity of cattle to improve the beef product quality and uniformity, thus improving revenue generated from beef sales.
[0025] This invention allows the identification of animals that have superior traits that can be used to identify parents of the next generation through selection. These methods can be imposed at the nucleus or elite breeding level where the improved traits would, through time, flow to the entire population of animals, or could be implemented at the multiplier or foundation parent level to sort parents into most genetically desirable. The optimum male and female parent can then be identified to maximize the genetic components of dominance and epistasis, thus maximizing heterosis and hybrid vigor in the market animals.
…
[0027] In another embodiment, the invention provides methods to draw an inference of a trait of a bovine subject by determining the nucleotide occurrence of at least one bovine SNP that is determined using methods disclosed herein, to be associated with the trait. For example, the inference can be drawn by determining the nucleotide occurrence of at least one SNP identified in Tables 1A and 1B (i.e. a SNP corresponding to position 300 of SEQ ID NOS:19473 to 21982). The inference can be drawn regarding, for example, fat thickness, retail yield, marbling, tenderness, or average daily gain.
…
[0029] In another embodiment, the present invention provides a method for identifying a bovine target sequence, such as a gene, associated with a trait, by identifying an open reading frame present in a target region of the bovine genome, wherein the target region is located on the bovine genome less than or equal to about 500,000 nucleotides of a single nucleotide polymorphism (SNP) corresponding to position 300 of any one of SEQ ID NOS:19473 to 21982, and analyzing the open reading frame to determine whether it affects the trait, thereby identifying a bovine gene associated with the trait. In one aspect, the target region is located within about 5000 nucleotides of a single nucleotide polymorphism (SNP) corresponding to position 300 of any one of SEQ ID NOS:19473 to 21982.
[0030] In another embodiment, the present invention provides a method for identifying a bovine single nucleotide polymorphism (SNP) associated with a trait, that includes identifying a test SNP in a target region of a bovine genome, wherein the target region is less than or equal to about 500,000 nucleotides of a SNP position corresponding to position 300 of one of SEQ ID NOS:19473 to 21982, and identifying an association of the test SNP to the trait, thereby identifying the test SNP as associated with the trait In certain aspects, the target region includes at least 20 contiguous nucleotides of SEQ ID NOS:24493 to 64886. In another aspect, for example, the target region includes at least 20 contiguous nucleotides of SEQ ID NOS:19473 to 21982. The present invention also provides isolated polynucleotides that include the identified SNPs.
170 As described, the invention uses “information regarding genetic diversity among cattle, particularly SNPs, and the effect of nucleotide occurrences of SNPs on important traits” (at [0023]). I would note at this point that Branhaven’s case is that, before the priority date, very few SNPs had been identified in cattle and the technology that was available for genotyping them was either expensive or of low throughput. Indeed Branhaven has contended that SNPs were not a commonly used genetic marker at the priority date. It has contended that a major transition towards the use of SNP markers only occurred later in 2008, when a research tool called the BovineSNP50 SNP chip was commercialised. I would note also at this point that it was accepted by all parties that before the priority date, the bovine genome had not been sequenced and that version 1 of the bovine genome was not published until about mid-2005.
171 Further, as to the form of the invention described in [0027], which I have set out above, it is to be noted that [0027] refers to 2,510 specified SNPs and five identified traits, namely, fat thickness, retail yield, marbling, tenderness and daily gain, but in non-limiting terms. In comparison, claim 1 requires the identification of at least three SNPs (one of which must be a limb (a) or limb (b) SNP) associated with the trait of interest (not limited to the five identified traits) in the use of the claimed method, although the specification elsewhere contemplates the use of greater numbers of SNPs and explains that this might be done to infer a higher likelihood that the trait will exist. I will return to this later when discussing the question of whether the “at least 3 SNPs” requirement is arbitrary and also the question of fair basis concerning a “trait” in claim 1 (i.e. beyond the five traits identified).
172 I would also note that a further embodiment is described at [0030], which refers to the identification of other SNPs within a target region of about 500,000 nucleotides of one of the 2,510 SNPs identified by the sequence listings and Tables 1A and 1B. I will return later to the question of whether this reflects the latter part of claim 1 referred to as limb (b) (see also [0035] of the specification).
Detailed description of the invention
173 A detailed description of the invention is then set out from [0031] to [0211] (pages 12 to 66a). Key provisions thereof include [0031] to [0035], [0036], [0038], [0051] to [0060], [0071] to [0077], [0079] to [0082], [0088] to [0091], [0101], [0106], [0108], [0112] to [0114], [0116], [0117], [0121], [0122], [0125] to [0128], [0130] to [0135], [0138], [0140], [0141], [0143], [0147], [0149], [0150], [0152] and [0155].
174 It is appropriate to set out some of these paragraphs:
[0031] The specification hereby incorporates by reference in their entirety, the files contained on the two compact discs filed herewith. Two copies of each of the two compact discs are filed herewith. The first compact disc includes a file called “mmi1100wo Table 1A.doc,” created December 31, 2003, which is 11299 kilobytes in size, and a file called “mmi1100wo Table 1B.doc,” created December 31, 2003, which is 11266 kilobytes in size. The Second disc includes a sequence listing which is included in a file called “MMI1100WO SEQUENCE LISTING.txt,” created December 31, 2003, which is 88096 kilobytes in size.
…
[0034] Presently, feedlots contain pens which typically have a capacity of about 200 animals, and market to packers, pens of cattle that are fed to an average endpoint. The endpoint is calculated as a number of days on feed estimated from biological type, sex, weight, and frame score. Animals are initially sorted to a pen based on the estimated number of days on feed and incoming group. However, sorting is done by a series of subjective and suboptimal parameters, as discussed herein. The cattle are fed to an endpoint in order to maximize the percentage of animals from which Grade USDA Choice beef can be obtained at slaughter without developing cattle that are too fat, and thus get discounted for insufficient red meat yield. The present invention provides a method for maximizing a physical characteristic of a bovine subject, including optimizing the percentage of bovine subjects that produce Grade USDA Choice and Prime beef in the most efficient manner.
[0035] In one embodiment, the present invention provides an isolated polynucleotide that includes a fragment of at least 20 contiguous nucleotides of the bovine genome, or a complement thereof, wherein the isolated polynucleotide includes a nucleotide occurrence of a single nucleotide polymorphism (SNP) associated with a trait, wherein the SNP is in disequilibrium with a SNP corresponding to position 300 of any one of SEQ ID NOS:19473 to 21982. In certain aspects, the polynucleotide is located about 500,000 or less nucleotides from position 300 of SEQ ID NOS:19473 to 21982 on the bovine genome. As disclosed in the Examples herein, the linkage disequilibrium for cattle is about 500,000 nucleotides. Therefore, it is expected that other SNPs can be identified that are associated with the same traits based on the fact that these other SNPs are located less than or equal to about 500,000 nucleotides of the identified associated SNP on the bovine genome. In certain aspects, the polynucleotide is from an Angus, Charolais, Limousin, Hereford, Brahman, Simmental or Gelbvieh bovine subject.
[0036] The attached sequence listing provides sequences of contigs (SEQ ID NOS:24493 to 64886) generated from the bovine genome. It will be understood that contigs can be aligned such that SNPs that are on separate contigs, but are located within 500,000 nucleotides on the bovine genome, can be identified. For example, alignment of contigs and determination of distance between contigs provided herein, can be accomplished by using the sequence information of the human genome as a scaffold. Tables 1A and 1B (filed herewith on the compact disc), lists contigs that are “nearby” (i.e. within 500,000 nucleotides on the bovine genome) an associated SNP.
…
[0038] In another aspect, the present invention provides an isolated polynucleotide that includes a polynucleotide that is at least 20 nucleotides in length and is at least 90% identical to a fragment of at least 20 contiguous nucleotides of a bovine genome; or a complement thereof, wherein the fragment of at least 20 contiguous nucleotides of the bovine genome comprises a nucleotide occurrence of a single nucleotide polymorphism (SNP) associated with a trait, wherein the SNP is about 500,000 or less nucleotides from position 300 of any one of SEQ ID NOS:19473 to 21982. In certain aspects, for example, the polynucleotide is at least 90% identical to a fragment of at least 10, 15, 20, 25, 50, or 100 contiguous nucleotides of SEQ ID NOS:19473 to 21982. In certain aspects, the polynucleotide comprises position 300 of SEQ ID NOS: 19473 to 21982.
…
[0051] In another embodiment, the invention provides a method for drawing an inference regarding a trait of a bovine subject by determining the nucleotide occurrence of at least one bovine SNP that is associated with the trait. A SNP is associated with a trait when at least one nucleotide occurrence of the SNP occurs more frequently in subjects with a certain characteristic of the trait in a statistically significant manner, for example with greater than 80%, 85%, 90%, 95%, or 99% confidence. Therefore, in certain aspects, the methods include identifying whether the nucleotide occurrence is a bovine SNP allele identified herein as associated with a trait. A bovine “SNP allele” is a nucleotide occurrence of a SNP within a population of bovine animals. The inference, in certain aspects, is drawn by determining the nucleotide occurrence of one or more SNPs corresponding to position 300 of SEQ ID NOS:19473 to 21982. These SNPs are referred to herein as “associated SNPs.” The inference can be drawn regarding a variety of traits as discussed herein, such as, for example, fat thickness, retail yield, marbling, tenderness, or average daily gain. In certain aspects, the bovine subject is an Angus, Charolais, Limousin, Hereford, Brahman, Simmental or Gelbvieh bovine subject.
[0052] As illustrated in the Example provided herein, a high density SNP map of the bovine genome was constructed and analyzed for the presence of SNPs that are associated with a trait at a confidence level of 0.01 or greater. The identified SNPs are referred to herein as “SNPs that are associated with a trait” or “associated SNPs.” The predictive value of the associated SNPs allow a determination of the genetic potential of a bovine animal to express multiple economically important traits, termed the molecular breeding and selection value. This information is utilized to enhance the efficiency and accuracy of breeding, sorting and cloning of animals.
[0053] The analysis disclosed in the Examples herein, utilized methods of the present invention, to generate a high-density genetic map of the bovine genome based on single nucleotide polymorphic (SNP) markers. The high-density genetic map was created through a whole genome sequence of the bovine genome using the shotgun sequencing approach as described by Venter, J.C, et al., (Science 291:1304-1351 (2001)). Shotgun sequencing was performed with four different bovine individuals that represent different breed types. Upon whole genome assembly of the sequenced fragments all sequence variants were identified and cataloged. Sequence variants that differ by a single nucleotide became candidate SNP markers for the high-density map. The relative position of each candidate SNP within the bovine genome was determined by using the assembled human genome as scaffolding. Candidate SNPs were chosen based on their locations so that the map is evenly distributed across the bovine genome. The genetic SNP map is evenly distributed where the average genetic distance between any two adjacent markers is 0.5 cM.
[0054] Furthermore, phenotypic data from 3791 bovine animals was collected from a three by three factorial feeding and carcass data collection experiment, comparing three biological types (English, Continental and Brahman crosses) within three different days on feed (early, optimum and late). Animals were randomly assigned to treatment groups based on biological type. All cattle entered the experiment within 90 kg of body weight. These groups were blocked across starting and harvest date. Blood samples were collected on each individual animal at the start of the feeding period and assigned an electronic ID that was matched to the collection sample. At the completion of the feeding and harvest period data were compiled and analyzed for relevant statistical parameters. Statistically significant associations between specific SNPs and targeted traits were identified by methods disclosed herein for utilizing a high-density genetic SNP map in the performance of whole genome association studies in bovine animals. Using methods and results provided herein, the effect of the associated SNP on the target trait through allele frequency differences in the SNP was determined. Furthermore, as disclosed herein, SNPs that are adjacent to or in close proximity to some of the associated SNPs were identified that are associated with the same traits as an associated SNP.
[0055] As discussed in detail in the attached Examples, DNA samples were pooled from bovine subjects that represent high and low phenotypic extremes for the expression of a target trait in a population of bovine animals (e.g. high fat). The traits selected for analysis were marbling, tenderness, fat thickness, yield, and daily gain. A total of 2510 SNPs were identified that are associated with these traits (Tables 1A and 1B).
[0056] Tables 1A and 1B, both of which are filed herewith on a compact disc, disclose the SNPs identified by the analysis, and provide the SNP names for the SNPs corresponding to position 300 of SEQ ID NOS:19473 to 21982. The sequences disclosed in SEQ ID NOS:SNP1 to SNP4000 are regions from which amplicons were generated. Table 1B also indicates the location of the amplicon-generating regions within a larger bovine genomic sequence contig (SEQ ID NOS:24493 to 64886) (See column 2 of Table 1B, labeled “In Sequence,” which lists a contig name (e.g., “19866880525139”) and positions (e.g. “923-1522”) within the contig of an amplicon which includes the SNP at position 300. A sequence identifier for the amplicon (SEQ ID NOS:19473-21982) is listed in Table 1A. Furthermore, Tables 1A and 1B identify the nucleotide occurrences that have been detected for each of these SNPs, and identifies traits that have been identified to be associated with these SNPs using methods disclosed herein. All of the SNPs listed in Tables 1A and 1B were associated with the respective trait(s) with a confidence level of 0.01, or higher confidence. Finally, Table 1A provides the sequence of an extension primer that was used to determine the nucleotide occurrence of the SNPs (SEQ ID NOS:21983 to 24492).
[0057] Each SNP in Tables 1A and 1B is characterized by the trait(s) found to be in association: marbling, tenderness, fat thickness, daily gain and retail yield. For each of the five traits, “High” refers to animals reaching the 90th percentile of that phenotypic measurement based on numeric ranking for the trait. “Low” refers to animals in the 10th percentile or less of that phenotypic measurement based on the numeric ranking of the trait.
[0058] In certain aspects of the invention directed at methods for inferring traits such as the traits listed in Tables 1A and 1B, nucleotide occurrences are determined for one or more associated SNPs. Therefore, in one aspect, for example, the method is used to infer fat thickness, by determining a nucleotide occurrence of at least one SNP corresponding to the SNPs indicated in Tables 1A and 1B as associated with fat thickness. For this aspect, as a non-limiting example, a nucleotide occurrence of the SNP at position 300 of SEQ ID NO:19473 can be identified and compared to the nucleotide occurrences listed in Tables 1A and 1B 1 for SEQ ID NO:19473. A thymidine residue at position 300 of SEQ ID NO:19473 infers a higher likelihood that the bovine subject will produce meat that has high tenderness. In addition, as a non-limiting example, a nucleotide occurrence at position 300 of SEQ ID NO:19474 can be determined and used alone or in combination with the nucleotide occurrence at position 300 of SEQ ID NO:19473, to infer tenderness. For example, if position 300 of both SEQ ID NO:19473 and SEQ ID NO:19474 are thymidine residues, there is an even greater likelihood that the bovine subject will produce meat that has high tenderness, than for either nucleotide occurrence alone.
[0059] In another aspect, the method is used to infer retail yield, by determining a nucleotide occurrence of at least one SNP corresponding to the SNPs indicated in Table 1A as associated with retail yield. In another aspect, the method is used to infer marbling by determining a nucleotide occurrence of at least one SNP corresponding to the SNPs indicated in Table 1A as associated with marbling. In another aspect, the method is used to infer daily gain, by determining a nucleotide occurrence of at least one SNP corresponding to the SNPs indicated in Table 1A as associated with daily gain.
[0060] For any trait, a “relatively high” characteristic, indicates greater than average, and a “relatively low” characteristic indicates less than average. For example “relatively high marbling”, indicates more abundant marbling than average marbling for a bovine population. Conversely, “relatively low marbling”, indicates less abundant marbling than average marbling for a bovine population. Furthermore, in certain aspects, methods of the present invention infer that a bovine subject has a significant likelihood of having a value for a trait that is within, for example, the 5th, 10th, 20th, 25th, 30th, 40th, 50th, 60th, 70th, 75th, 80th, 90th, or 95th percentile of bovine subjects for a given trait. For example, a method presented herein can provide an inference that a bovine subject has a significant likelihood of having a marbling value that is within the 10th percentile of marbling for a bovine population …
…
[0072] The method, in certain examples, includes identification of the causative mutation influencing the trait directly or the determination of 1 or more SNPs that are in linkage disequilibrium with the associated trait.
[0073] The method can include a determination of the nucleotide occurrence of at least 2 SNPs. At least 2 SNPs can form all or a portion of a haplotype, wherein the method identifies a haplotype allele that is in linkage disequilibrium and thus associated with the trait. Furthermore, the method can include identifying a diploid pair of haplotype alleles.
…
[0075] As used herein, the term “at least one”, when used in reference to a gene, SNP, haplotype, or the like, means 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, etc., up to and including all of the haplotype alleles, genes, and/or SNPs of the bovine genome. Reference to “at least a second” gene, SNP, or the like, means two or more, i.e., 2, 3, 4, 5, 6, 7, 8, 9, 10, etc., bovine genes, SNPs, or the like.
[0076] Polymorphisms are allelic variants that occur in a population that can be a single nucleotide difference present at a locus, or can be an insertion or deletion of one, a few or many consecutive nucleotides. As such, a single nucleotide polymorphism (SNP) is characterized by the presence in a population of one or two, three or four nucleotides (i.e., adenosine, cytosine, guanosine or thymidine), typically less than all four nucleotides, at a particular locus in a genome such as the human genome. It will be recognized that, while the methods of the invention are exemplified primarily by the detection of SNPs, the disclosed methods or others known in the art similarly can be used to identify other types of bovine polymorphisms, which typically involve more than one nucleotide.
…
[0079] As used herein, the term “infer” or “inferring”, when used in reference to a trait, means drawing a conclusion about a trait using a process of analyzing individually or in combination, nucleotide occurrence(s) of one or more SNP(s), which can be part of one or more haplotypes, in a nucleic acid sample of the subject, and comparing the individual or combination of nucleotide occurrence(s) of the SNP(s) to known relationships of nucleotide occurrence(s) of the SNP(s) and the trait. As disclosed herein, the nucleotide occurrence(s) can be identified directly by examining nucleic acid molecules, or indirectly by examining a polypeptide encoded by a particular gene where the polymorphism is associated with an amino acid change in the encoded polypeptide.
[0080] Relationships between nucleotide occurrences of one or more SNPs or haplotypes and a trait can be identified using known statistical methods. A statistical analysis result which shows an association of one or more SNPs or haplotypes with a trait with at least 80%, 85%, 90%, 95%, or 99%, or 95% confidence, or alternatively a probability of insignificance less than 0.05, can be used to identify SNPs and haplotypes. These statistical tools may test for significance related to a null hypothesis that an on-test SNP allele or haplotype allele is not significantly different between groups with different traits. If the significance of this difference is low, it suggests the allele is not related to a trait.
[0081] As another example, associations between nucleotide occurrences of one or more SNPs or haplotypes and a trait (i.e. selection of significant markers) can be identified using a two part analysis in the first part, DNA from animals at the extremes of a trait are pooled, and the allele frequency of one or more SNPs or haplotypes for each tail of the distribution is estimated. Alleles of SNPs and/or haplotypes that are apparently associated with extremes of a trait are identified and are used to construct a candidate SNP and/or haplotype set. Statistical cut-offs are set relatively low to assure that significant SNPs and/or haplotypes are not overlooked during the first part of the method.
[0082] During the second stage, individual animals are genotyped for the candidate SNP and/or haplotype set. The second stage is set up to account for as much of the genetic variation as possible in a specific trait without introducing substantial error. This is a balancing act of the prediction process. Some animals are predicted with high accuracy and others with low accuracy.
…
[0088] The present invention can also be used to provide information to breeders to make breeding, mating, and or cloning decisions. This invention can also be combined with traditional genetic evaluation methods to improve selection, mating, or cloning strategies.
[0089] The subject of the present invention can be any bovine subject, for example a bull, a cow, a calf, a steer, or a heifer or any bovine embryo or tissue, and includes all breeds of bovines. For methods of the invention directed at sorting bovine subjects, managing bovine subjects, improving profits related to selling beef from a bovine subject, the animal can be a young bovine subject ranging in ages from conception to the time the animal is harvested and beef and other commercial products obtained. The method of the present invention can be performed after the animal is purchased and first enters the feedlot.
[0090] A “trait” is a characteristic of an organism that manifests itself in a phenotype. Many traits are the result of the expression of a single gene, but some are polygenic (i.e., result from simultaneous expression of more than one gene). A “phenotype” is an outward appearance or other visible characteristic of an organism. Many different non-bovine livestock traits can be inferred by methods of the present invention. Traits analyzed in methods of the present invention include, but are not limited to, marbling, tenderness, quality grade, quality yield, muscle content, fat thickness, feed efficiency, red meat yield, average daily weight gain, disease resistance, disease susceptibility, feed intake, protein content, bone content, maintenance energy requirement, mature size, amino acid profile, fatty acid profile, milk production, hide quality, susceptibility to the buller syndrome, stress susceptibility and response, temperament, digestive capacity, production of calpain, calpastatin and myostatin, pattern of fat deposition, ribeye area, fertility, ovulation rate, conception rate, fertility, heat tolerance, environmental adaptability, robustness, susceptibility to infection with and shedding of pathogens such as E. Coli, Salmonella sp. and other human pathogens.
…
[0101] This invention identifies animals that have superior traits, predicted very accurately, that can be used to identify parents of the next generation through selection. These methods can be imposed at the nucleus or elite breeding level where the improved traits would, through time, flow to the entire population of animals, or could be implemented at the multiplier or foundation parent level to sort parents into most genetically desirable. This invention provides a method for determining the optimum male and female parent to maximize the genetic components of dominance and epistasis thus maximizing heterosis and hybrid vigor in the market animals.
…
[0106] A series of markers or a series of SNPs as used herein, can include a series of at least 2, 3, 4, 5, 6, 7, 8, 9, 10, 15, 20, 25, 30, 35, 40, 45, 50, 75, 100, 150, 200, 250, 500, 1000, 2000, 2500, 5000, or 6000 markers, for example.
…
[0108] The nucleotide occurrence of at least 2 SNPs can be determined. The at least 2 SNPs can form a haplotype, wherein the method identifies a haplotype allele that is associated with the trait. The method can include identifying a diploid pair of haplotype alleles for one or more haplotypes.
…
[0112] Accordingly, methods of the invention can involve determining the nucleotide occurrence of at least 2, 3, 4, 5, 10, 20, 30, 40, 50, etc. SNPs. The SNPs can form all or part of a haplotype, wherein the method can identify a haplotype allele that is associated with the trait. Furthermore, the method can include identifying a diploid pair of haplotype alleles.
[0113] In another embodiment, the present invention provides a method for identifying a bovine genetic marker that influences at least one trait by analyzing bovine genetic markers of a genome-wide genetic marker map for association with the trait. The genetic marker can be a single nucleotide polymorphism (SNP), or can be at least two SNPs that influence the trait. Because the method can identify at least two SNPs, and in some embodiments, many SNPs, the method can identify not only additive genetic components, but non-additive genetic components such as dominance (i.e. dominating trait of an allele of one gene over an allele of a another gene) and epistasis (i.e. interaction between genes at different loci). Furthermore, the method can uncover pleiotropic effects of SNP alleles (i.e. SNP alleles or haplotypes effects on many different traits), because many traits can be analyzed for their association with many SNPs using methods disclosed herein.
[0114] In one aspect, expression products of genes near the at least two identified genetic markers are analyzed, to determine whether the expression products interact. In certain aspects, at least 2 SNPs are identified for inferring the genetic potential of a bovine animal for one, two, or more traits. At least 2 of the single nucleotide polymorphisms are located on different chromosomes. Furthermore, at least 2 of the single nucleotide polymorphisms can be separated by at least 10,000 base pairs on the bovine genome. In certain examples, at least 2 of the single nucleotide polymorphisms occur in different genes.
…
[0116] The present invention provides methods to allow the simultaneous discovery of any and all SNP markers that associate with one or more traits in one or more regions throughout the entire genome. Furthermore, the present invention provides methods for utilization of the predictive diagnostic to determine the genetic potential of an animal to express any targeted trait(s). The genetic potential of a bovine animal to express multiple economically important traits, termed the molecular breeding and selection value, is utilized to enhance the efficiency and accuracy of breeding, sorting and cloning of animals.
[0117] The present invention provides methods for developing a high-density genetic map of the bovine genome based on single nucleotide polymorphic (SNP) markers. The high-density genetic map is created through a whole genome sequence of the bovine genome using the shotgun sequencing approach. Shotgun sequencing is performed with several different bovine individuals that represent different breed types. Upon whole genome assembly of the sequenced fragments all sequence variants are identified and cataloged. Sequence variants that differ by a single nucleotide become candidate SNP markers for the high-density map. The relative position of each candidate SNP within the bovine genome is determined by using the assembled human genome as scaffolding. Candidate SNPs are chosen based on their locations so that the map is evenly distributed across the bovine genome. The invention includes methods for creating an evenly distributed genetic SNP map where the average genetic distance between any two adjacent markers is 0.5 cM (i.e. 500,000 nucleotides).
…
[0121] The methods infer the discovery of one or more, and in some cases, all SNPs that show association to a target trait and therefore, account for a large proportion of the genetic variation observed in the expression of the trait in a population of bovine animals. The methods allow identification of SNPs that account for additive as well as non-additive genetic variation, such as dominance and epistasis, observed in the expression of the trait.
[0122] The methods infer the discovery of any and all SNPs that show association to one or more target traits. Furthermore, whereby certain traits have either positive or negative correlations to each other, the methods allow identification of all SNPs that enhance or uncouple the correlation. For example, the presence of external fat on beef carcasses is highly correlated with marbling or intra-muscular fat. External fat is an undesirable trait that causes discounts in beef carcasses, whereas marbling is a desirable trait that results in premiums. The present invention provides methods for the identification of all SNPs that may uncouple the correlation between external fat and marbling, for example.
…
[0125] Accordingly, nucleotide occurrences can be determined for essentially all, or all of the SNPs of a high-density, whole genome SNP map. This approach has the advantage over traditional approaches in that since it encompasses the whole genome, it identifies potential interactions of gene products expressed from genes located anywhere on the genome without requiring preexisting knowledge regarding a possible interaction between the gene products. An example of a high-density, whole genome SNP map is a map of at least about 1 SNP per 10,000 kb, at least 1 SNP per 500 kb or about 10 SNPs per 500 kb, or at least about 25 SNPs or more per 500 kb. Definitions of densities of markers may change across the genome and are determined by the degree of linkage disequilibrium from marker to marker.
[0126] In another embodiment of the invention, a method is provided for identifying SNPs that are associated with a trait by using the associated SNPs disclosed herein. The method is based on the fact that other markers in close proximity to the associated SNP marker will also associate with the trait because markers in linkage disequilibrium with the associated SNP marker will also be in linkage disequilibrium with the gene(s) influencing the trait. SNPs in linkage disequilibrium can be used in lieu of determining a SNP or mutation to predict the presence or absence of a phenotypic trait or contributor to a phenotypic trait. Accordingly, in certain embodiments, the present invention provides a method for identifying a SNP associated with a trait, that includes identifying a test SNP that is in disequilibrium with a SNP corresponding to position 300 of SEQ ID NOS:19473 to 21982.
[0127] As illustrated in the Examples section, it has been determined that disequilibrium exists across the region of 500,000 bp from the associated SNP in each direction. Other markers within this 500,000 bp region will also be in disequilibrium with the associated SNP and with the trait of interest, and can be used to infer associations with the trait of interest. Genomic segments containing the markers can be adjacent to the associated SNP marker or contained within a separate island of sequence distant from the associated SNP.
[0128] Genetic markers within 500,000 bp of the associated SNPs disclosed herein in Tables 1A and 1B (position 300 of SEQ ID NOS:19473 to 21982), can be discovered by a number of different methods known in the art. In one aspect of the invention, bovine sequence that is within 500,000 bp of the associated SNP can be used to identify new DNA markers. This sequence can be created from whole-genome shotgun sequencing, BAC-sequencing, or sequence generated from comparative maps. The bovine sequence can be used to develop bovine specific sequencing primers. These primers can be used to sequence at least 2 individual bovine animals and the alignments from these sequences can be used to identify SNP markers and microsatellite markers.
…
[0130] These new markers can be genotyped in pools of animals or individual animals representing the high and low ends of the phenotypic distribution for the trait to determine association between the new marker(s) and the trait. Markers with a significantly different allele frequency in the high and low groups are also in disequilibrium with the trait.
[0131] Accordingly, in another embodiment, the present invention provides a method for identifying a bovine single nucleotide polymorphism (SNP) associated with a trait that includes identifying a test SNP in a target region of a bovine genome, wherein the target region is less than or equal to about 500,000 nucleotides from a SNP position corresponding to position 300 of one of SEQ ID NOS:19473 to 21982, and identifying an association of the test SNP to the trait. In certain aspects, the target region consists of at least 20 contiguous nucleotides of SEQ ID NOS:24493 to 64886. In other aspects, the target region consists of at least 20 contiguous nucleotides of SEQ ID NOS:19473 to 21982.
[0132] In certain aspects, the test SNP is located less than or equal to about 500,000, 400,000, 300,000, 250,000, 200,000, 100,000, 50,000, 25,000, 10,000, 5,000, 1,000, or 100 nucleotides from a position corresponding to position 300 of at least one of SEQ ID NOS:19473 to 21982. The test SNP is expected to be associated with the same trait as a SNP that corresponds to position 300 of SEQ ID NOS:19473 to 21982 that is located less than or equal to about 500,000 nucleotides from the test SNP, as discussed further herein.
[0133] The trait can be any bovine trait as discussed herein. For example, the trait can be marbling, tenderness, quality grade, muscle content, fat thickness, feed efficiency, red meat yield, average daily weight gain, disease resistance, disease susceptibility, feed intake, protein content, bone content, maintenance energy requirement, mature size, amino acid profile, fatty acid profile, milk production, susceptibility to the buller syndrome, stress susceptibility and response, temperament, digestive capacity, production of calpain, caplastatin and myostatin, pattern of fat deposition, ribeye area, fertility, ovulation rate, conception rate, fertility, susceptibility to infection with or shedding of pathogens. In certain specific examples, the trait is fat thickness, retail yield, tenderness, marbling, or average daily gain.
[0134] In another embodiment, the present invention provides a method for identifying a bovine gene associated with a trait, by identifying an open reading frame present in a target region of the bovine genome, wherein the target region is located on the bovine genome less than or equal to about 500,000 nucleotides of a single nucleotide polymorphism (SNP) corresponding to position 300 of any one of SEQ ID NOS:19473 to 21982, and analyzing the open reading frame to determine whether it affects the trait.
[0135] In certain aspects, the target region is located less than or equal to about 500,000, 400,000, 300,000, 250,000, 200,000, 100,000, 50,000, 25,000, 10,000, 5,000, 1,000, 100, or 50 nucleotides from a single nucleotide polymorphism (SNP) corresponding to position 300 of any one of SEQ ID NOS:19473 to 21982.
…
[0138] If a polynucleotide is identified as at least 90% identical to the SNP-containing polynucleotide, the bovine polynucleotide is a target polynucleotide for the trait.
…
[0140] The invention, in another aspect includes methods for creating a high density bovine SNP map. The SNP markers and their surrounding sequence are compared to model organisms, for example human and mouse genomes, where the complete genomic sequence is known and syntenic regions identified. The model organism map may serve as a template for ensuring complete coverage of the animal genome. The finished map has markers spaced in such a way to maximize the amount of linkage disequilibrium in a specific genetic region.
[0141] This map is used to mark all regions of the chromosomes, in a single experiment, utilizing thousands of experimental animals in an association study, to correlate genomic regions with complex and simple traits. These associations can be further analyzed to unravel complex interactions among genomic regions that contribute to the targeted trait or other traits, epistatic genetic interactions and pleiotropy. The invention of regional high density maps can also be used to identify targeted regions of chromosomes that influence traits.
…
[0152] Medium to high-throughput systems for analyzing SNPs, known in the art such as the SNPStream® UHT Genotyping System (Beckman/Coulter, Fullerton, CA) (Boyce-Jacino and Goelet Patents), the Mass ArrayTM system (Sequenom, San Diego, CA) (Storm, N. et al. (2002) Methods in Molecular Biology 212: 241-262.), the BeadArrayTM SNP genotyping system available from Illumina (San Diego, CA) (Oliphant, A., et al. (June 2002) (supplement to Biotechniques), and TaqManTM (Applied Biosystems, Foster City, CA) can be used with the present invention. However, the present invention provides a medium to high-throughput system that is designed to detect nucleotide occurrences of bovine SNPs, or a series of bovine SNPs that can make up a series of haplotypes. Therefore, as indicated above the system includes a solid support or other method to which a series of oligonucleotides can be associated that are used to determine a nucleotide occurrence of a SNP for a series of bovine SNPs that are associated with a trait. The system can further include a detection mechanism for detecting binding of the series of oligonucleotides to the series of SNPs. Such detection mechanisms are known in the art.
…
[0155] Numerous methods are known in the art for determining the nucleotide occurrence for a particular SNP in a sample. Such methods can utilize one or more oligonucleotide probes or primers, including, for example, an amplification primer pair that selectively hybridizes to a target polynucleotide, which corresponds to one or more bovine SNP positions …
175 It is also necessary to set out in more elaborate terms than is usual the illustrative examples described at [0190] to [0208] (pages 59 to 66):
[0190] The following examples are intended to illustrate but not limit the invention.
EXAMPLE 1
Generation of a high-density bovine genetic snp map
[0191] This example illustrates the generation of a high density bovine genetic SNP map created through a whole genome sequencing of the bovine genome using the shotgun sequencing approach. This approach was selected to provide hundreds of thousands of SNP markers, as described by Venter, J.C, et al., (Science 291:1304-1351 (2001), in order to perform a whole-genome association study with adequate density of markers to ensure discovery of markers in disequilibrium with mutations influencing the targeted traits.
[0192] Shotgun sequencing was performed with four different bovine individuals that represented different breed types. The shotgun sequencing was performed according to the methods of Venter, J.C, et al., (Science 291:1304-1351 (2001)). By this method, random fragments of bovine sequence were generated and size selected to 2.5 and 10 kb. These fragments of bovine DNA were inserted into a sequencing vector to create high quality plasmid libraries suitable for high throughput sequencing.
[0193] Shotgun sequencing was performed with four different bovine subjects that represented several different breed types: Angus, Limousin, Brahman and Simmental. Upon whole genome assembly of the sequenced fragments, contigs were formed from consensus sequence, and sequence variants were identified and cataloged. 786,777 sequence variants that differed by a single nucleotide became candidate SNP markers for the high-density SNP map. The relative position of each candidate SNP within the bovine genome was determined using the assembled human genome as scaffolding creating a candidate map of 242,181 human-mapped markers. Upon positioning of the SNPs within the genome, individual markers were tested to determine informativeness within the cattle population using 210 animals representing diverse breeds (Angus, Charolais, Limousin, Hereford, Brahman, Simmental and Gelbvieh) and Mendialian segregation (21 trios of parents and progeny). Selected markers were polymorphic in the majority of the breeds tested. Any markers within a region that failed the test were discarded and replaced with another marker in the region. These markers were also validated against the test population. This process was repeated until a relatively evenly distributed genetic SNP map was obtained, where the average genetic distance between any two adjacent markers is 0.5 cM.
Example 2
Identification of Bovine snps associated with tenderness, fat, marbling, yield, and/or daily gain
[0194] This example illustrates the identification of SNPs from the high-density bovine SNP map identified in Example I, that are associated with the traits meat tenderness, fat thickness, marbling, yield, and/or daily gain.
[0195] DNA samples from bovine subjects were obtained by collecting 50 ml of whole blood from the 4,791 bovine subjects. 25 ml of whole blood was used for DNA extraction using standard methods and concentrations of DNA were calculated using standard fluorimetric methods. Animals representing less than or equal to the 10th percentile of low numeric phenotypic animals (44 individuals) and the 90th percentile and greater of high phenotypic animals (44 individuals) were identified for each trait. The low numeric values were identified as “Low” and the high numeric values were identified as “High”. DNA samples were pooled from bovine individuals that represent high and low phenotypic extremes for the expression of a target trait in a population of bovine animals with each of the 44 animals contributing equally to the pool of DNA. A separate “High” and “Low” pool was created for each biological type (English, Continental, and Brahman crosses) by treatment group (Early, Optimum, Late) for each of the five traits resulting in 90 total pools. In addition to the 90 pools listed above, another group was formed based on animals that were 5 standard deviations above the mean for numeric tenderness values. Eleven animals were included in this group of individuals and the pool was compared to the other tenderness groups resulting in a total of 91 pools. Each pool was tested against each of the 6189 mapped and validated SNP markers. The SNP detection platform utilized in the experiment was the Beckman Coulter SNP-IT system, utilizing single-base extension of the SNP base. Allele frequency was estimated for each pool based on the fluorescence intensity of each of the two incorporated fluorescent labels corresponding to the SNP alleles. These estimates were adjusted for marker specific characteristics and incorporation differences. A test statistic was developed based on a Chi-square distribution of differences among allele frequencies of the high minus low pools. These test statistics were summed across the 9 breed by treatment groups within each trait resulting in Chi-square distribution. SNP markers reaching a threshold test statistic of 46.96294 for the trait of tenderness and 21.66599 (p<.01) for the remaining four traits of retail yield, daily gain, fat thickness and marbling were identified as associated SNPs and are listed in Tables 1A and 1B.
[0196] The high-density SNP map was used to identify SNPs that are associated with a series of bovine traits. The traits included marbling, tenderness, fat thickness, yield, and daily gain. Tables 1A and 1B (filed herewith on a compact disc) provide the identity of SNPs that associated with one or more of the traits analyzed. Twenty five hundred and ten associated SNPs were identified for all five traits.
[0197] Table 1A provides the following information, from left to right columns: SNP name; a sequence identifier of the sequence listing filed herewith, for an amplicon, wherein the SNP position is position 300 of the amplicon; position of the SNP within the amplicon (i.e. position 300); The nucleotide sequence and SEQ ID NO: for an extension primer capable of priming polynucleotide synthesis across the SNP position; trait(s) that are associated with the SNP; Characteristics of the trait that are associated with specific nucleotide occurrences at the SNP; Nucleotide occurrences that have been detected at the SNP position; And the sequence identifier of contig sequences that are located within 500,000 nucleotides from the SNP on the bovine genome. Table 1B provides the following information from left to right columns: SNP name; A sequence name of a contig that includes the SNP position, as well as the position numbers within the contig for an amplicon that includes the SNP; Position of the SNP within the amplicon (i.e. position 300); The nucleotide sequence for an extension primer capable of priming polynucleotide synthesis across the SNP position; trait(s) that are associated with the SNP; Characteristics of the trait that are associated with specific nucleotide occurrences at the SNP; Nucleotide occurrences that have been detected at the SNP position; And the sequence identifier of contig sequences that are located within 500,000 nucleotides from the SNP on the bovine genome.
Example 3
Determination of the distance of disequilibrium in cattle
[0198] This example utilizes a few of the associated SNPs disclosed in Example 2, to identify additional SNPs that are associated with the same traits, using the physical proximity on the genome of the SNPs. Furthermore, the results are used to calculate a distance of disequilibrium in cattle. In this example, “shear force” is used to refer to tenderness, “vision retail yield” is used to refer to retail yield, and “average daily gain” is used to refer to daily gain.
[0199] In the past 10 years numerous methods have been developed to identify alleles associated with phenotypic effects, traits or diseases. Linkage disequilibrium and measures of linkage disequilibrium have been of particular interest for studies of complex traits or diseases … LD occurs where blocks or regions of neighboring markers are co-inherited from a common ancestor. The degree of LD varies considerably throughout the genome and is a function of time, recombination events, mutation rate and population structure. The extent of LD can vary from a few thousand base pairs to several centimorgans. This has been most extensively documented in human studies … Similar results have been observed in other species including cattle (F.Farnir, W.Coppieters, J-J. Arranz, et. al., “Extensive Genome-wide Linkage Disequilibrium in Cattle” Genome Research 10:220-227 (2000)). These studies and others have also shown that a SNP or multiple SNPs associated with a phenotype can be used as predictive of gene(s) causing differences in trait phenotypes within a region of high LD although they may or may not be the precise causative gene (as further examples, see also: A.M. Glazier, JH Nadeau and TJ Aitman, “Finding Genes that Underlie Complex Traits” Science 298: 2345-2348 (2002) … While it has been established that markers can be identified that associate with a specific trait, and, therefore, become diagnostic for the trait, the distance that disequilibrium reaches has not been determined in cattle with a dense marker map. Therefore, an experiment to determine the disequilibrium distance in cattle was performed using the high-density SNP map disclosed in Example 1.
[0200] The high-density SNP map disclosed in Example 1 was used to identify SNPs that are in physical proximity to a few of the associated SNPs disclosed in Example 2. Nucleotide occurrences of the SNPs were determined using the method disclosed in Example 2. A determination of whether on-test SNPs was associated with a trait was performed as disclosed in Example 2.
[0201] As discussed above, the study was performed to verify the assumption that markers that are in close physical proximity on the bovine genome will associate with the same trait(s) because markers in linkage disequilibrium with the associated SNP marker will also be in linkage disequilibrium with the mutation(s) influencing the trait.
[0202] As indicated in Table 2, SNP3 (MMBT22302) is significantly associated with the trait of average daily gain (“ADG” in Table 2). Several SNPs were identified using the high-density SNP map of Example I that are located at various distances from SNP3 on the bovine genome (Table 2). For example, SNP2 is 466,047 nucleotides from SNP3. Furthermore, SNP5 was identified which is 408,732 nucleotides from SNP3. SNP6 was identified which is 1.0 million nucleotides from SNP3. Finally, SNP4 was identified, which is 308,742 nucleotides.
[0203] As illustrated in Table 2, SNPs that were located within 500,000 nucleotides of SNP3 also were associated with average daily gain, whereas those that were located greater than 500,000 nucleotides from SNP3 were not associated with average daily gain. For example, linkage disequilibrium reaches 466,047 bases to SNP2, but not to SNPI at 1.5 Mb; linkage disequilibrium reaches to 408,732 bases to SNP5, but not to SNP6 at 1.0 Mb. SNP4, which is 308,742 nucleotides from SNP3, was discovered by sequencing the contig of DNA that maps to this region in 4 different breeds of cattle. It is also in disequilibrium with average daily gain.
[0204] Table 2. Disequilibrium analysis in relation to SNP distance from MMBT22302.
SNP | At Position 300 in SEQ ID NO | Marker Association P<.01 Trait | Human Chromosome Location | bp location | Difference from MMBT22302 | |
1 | MMBT22310 | not in patent | Not Sign | HC16 | 45425130 | 1,507,460 |
2 | MMBT13976 | 20291 | ADG | HC16 | 46466543 | 466,047 |
3 | MMBT22302 | 19666 | ADG | HC16 | 46932590 | |
4 | MMBT09532 | 21944 | ADG | HC16 | 47241332 | 308,742 |
5 | MMBT09533 | 19999 | ADG | HC16 | 47341322 | 408,732 |
6 | MMBT09535 | 21078 | Not Sign | HC16 | 47958246 | 1,025,656 |
[0205] To further analyze linkage disequilibrium, a similar analysis was performed using another SNP identified as an associated SNP in Example 2. SNP9 (MMBT03905) is significantly associated with vision retail yield (VRY). SNPs 7-8 and 10-12 were identified that are various distances from SNP9 (Table 2). Again, SNPs that were located less than or equal to about 500,000 nucleotides from the associated SNP, were also associated with the trait, whereas those that were present greater than 500,000 nucleotides from a known associated SNP, were not associated. For example, SNPs 8 and 11 were identified as also being highly significantly associated with VRY and are located less than 500,000 bp from SNP9 (Table 3). On the other hand, SNPs 7 and 12, which are greater than 500,000 bp from SNP9, were not associated with the trait. Furthermore, through additional sequencing, SNP10 was discovered and also found to be in linkage disequilibrium with VRY.
Table 3. Disequilibrium analysis in relation to SNP distance from MMBT3905.
SNP | At Position 300 in SEQ ID NO | Marker Association P<.01 Trait | Human Chromosome Location | bp location | Difference from MMBT03905 | |
7 | MMBT12437 | not in patent | Not Sign | HC04 | 177035705 | 518426 |
8 | MMBT03904 | 20327 | VRY | HC04 | 177201331 | 352800 |
9 | MMBT03905 | 19816 | VRY | HC04 | 177554131 | |
10 | MMBT03906 | 20240 | VRY | HC04 | 177900170 | 346039 |
11 | MMBT05906 | 20045 | VRY | HC04 | 178047550 | 493419 |
12 | MMBT03907 | not in patent | Not Sign | HC04 | 178113631 | 559500 |
[0206] As indicated in Tables 1A and 1B, SNP16 (MMBT02782) is highly significantly associated with shear force (SHF, Table 4). SNPs 14, 15, 17 and 18 were identified which are located within 500,000 nucleotides of SNP16 (Table 4). Once again, all of these SNPs, which are within 500,000 nucleotides of an associated SNP, were found to be associated with the same trait. That is, SNPs 14, 15, 17, and 18 were all found to be associated with SHF (Table 4). On the other hand, SNPs 1 and 7, which are located beyond 1.0 million nucleotides from SNP16, were not associated with SHF.
[0207] Table 4. Disequilibrium analysis in relation to SNP distance from MMBT02782.
SNP | At Position 300 in SEQ ID NO | Marker Association P<.01 Trait | Human Chromosome Location | bp location | Difference from MMBT02782 | |
13 | MMBT02777 | 19767 | Not Sign | HC04 | 46401363 | 1594271 |
14 | MMBT02781 | 20791 | SHF | HC04 | 47777758 | 217876 |
15 | MMBT19460 | 20790 | SHF | HC04 | 47778002 | 217632 |
16 | MMBT02782 | 20901 | SHF | HC04 | 47995634 | |
17 | MMBT03688 | 20765 | SHF | HC04 | 48379141 | 383507 |
18 | MMBT02784 | 20764 | SHF | HC04 | 48492482 | 496848 |
19 | MMBT02786 | not in patent | Not Sign | HC04 | 49190953 | 1195319 |
[0208] The results of this Example indicate that disequilibrium in cattle exists across the region of 500,000 nucleotides from an associated SNP, in each direction. Therefore, it is expected that when an associated SNP is identified, other markers within this 500,000bp region will also be in disequilibrium with the associated SNP and with the trait of interest, and can be used to infer associations with the trait of interest.
The steps of the claimed invention
176 The specification appears to disclose steps in the following sequence:
(a) Step 1 appears to involve the construction or generation of a high density bovine genetic SNP map.
(b) Step 2 appears to involve the construction or generation of an association study to identify bovine SNPs in the said map associated with various traits.
(c) Step 3 appears to involve the construction or generation of a study to determine a distance over which a degree of linkage disequilibrium (LD) is able to be established.
177 Before turning to detailed questions of construction, it is appropriate at this point to say something about each of these steps in turn and MLA’s arguments, including its characterisation of these steps. A brief discussion at this point will facilitate ease of comprehension in what later follows concerning inter-alia construction, some aspects of the manner of manufacture argument (at least whether that is disclosed on the face of the specification) and the question of inventive step.
(a) Step 1 – Construction of a high density bovine genetic SNP map
178 The specification describes that a high-density SNP map of the bovine genome was constructed and analysed for the presence of SNPs that were associated with a trait at a confidence level (p) value of 0.01 or greater (at [0052]). The specification says that “the methods of the present invention” were used to generate a high density genetic map of the bovine genome based on SNPs (at [0053]). The specification at [0141] also refers to the “invention of regional high density maps”.
179 MLA contends that the methods used to create the map and the creation of the map itself did not involve any invention and the map was known to be desirable. MLA contends that there was nothing new in constructing a high density genetic map using SNPs. It has also said that (as the specification indicates at [0076]), there is no magic in the use of SNPs because the methods of the invention could be used with other types of DNA markers. MLA says that SNPs are particularly suitable for whole of genome marker studies, because of their abundance throughout the genome.
180 Further, MLA says that the map was created using the same techniques that had been used to sequence the human genome, namely, shotgun sequencing as described in Venter JC et al, “The Sequence of the Human Genome” (2001) 291 Science 1304 (Venter). It is said that this is acknowledged at [0053]: see also [0128] and [0191]. MLA says that Venter was probably the “most important publication” in 2001 and was read and noted not only by scientists but also the general public. It was one of the first publications of a draft sequence of the human genome. Further, it points to the fact that a day before Venter was published, another paper that disclosed the sequence of the human genome was published by Lander ES et al, in the journal Nature, which was the result of the research by the International Human Genome Sequencing Consortium: Lander ES et al, “Initial Sequencing and Analysis of the Human Genome” (2001) 409 Nature 860 (Lander).
181 MLA also says that the only difference between the approach in Venter and the one used by the inventors for the 253 Application (for convenience, I will use the term “inventors” rather than “apparent inventors”, even though MLA asserts a lack of inventive step) is that the inventors’ “map”, which was not provided in or with the specification, was constructed using only parts of the bovine genome and mapping those parts against the human genome as a scaffold. The inventors did not obtain and assemble the entire bovine genome sequence without a reference scaffold. The relative positions of SNPs were determined using comparative mapping of the bovine genome with the human genome (at [0053]). MLA says that comparative mapping of genomes against another species was a well-known technique where the genes of a particular species were unknown.
182 Further, MLA contends that in the 253 Application, the SNPs were genotyped in the test animals using known methods (e.g. [0152]).
183 Now I accept that Example 1 describes in detail the generation of a high-density bovine genetic SNP map. DNA from four different animals of four different breeds was used ([0192] and [0193]). Shotgun sequencing produced sequenced fragments which were then assembled to form “contigs” from fragments which had some consensus. Variants in sequence were identified and catalogued. 786,777 sequence variants that differed by a single nucleotide became candidate SNP markers for the proposed high-density SNP map. The relative position of each SNP was determined using the human genome as a scaffold ([0193]). This led to a culling of the identified SNPs to 242,181 human-mapped markers.
184 I also accept that individual markers were then selected and tested to determine informativeness in 210 cattle from diverse breeds and selected markers were found that were polymorphic in the majority of breeds. Further, I accept that the process appears to have been that if a selected SNP within a region was not sufficiently informative (polymorphic), it was discarded and replaced with another marker “in the region”. The SNPs were validated against the test population. MLA says in reliance upon Professor Jeremy Taylor’s evidence, which I accept, that the term “validated” here does not mean that the SNPs were validated as associated with a trait in a broader population, but merely validated as being true SNPs. This process was repeated to obtain a relatively evenly distributed map of SNPs, with an aim to obtaining an average genetic distance between markers of 0.5 centimorgan (cM). It appears from [0195] that there were 6,189 such mapped markers.
185 Further, as MLA correctly contended, the map itself was not provided. The inventors only give (random) contigs (SEQ ID Nos 24493 to 64886) which “can be aligned … using the sequence information of the human genome as a scaffold” (at [0036]). While the inventors list contigs that are “‘nearby’ (i.e. within 500,000 nucleotides on the bovine genome)” a SNP that has been shown to be associated with a trait, they do not say where the contigs or associated SNPs are relative to any map. MLA says that the contigs provided in Table 1A are very small stretches of DNA. I accept that a review of Table 1A and the Sequence Listing shows that on average the contigs are about 1,484 nucleotides long. MLA says, in reliance upon Professor Benjamin Hayes’ evidence, that relying on them to find SNPs in 2002 and associate them with a trait of interest would have been difficult. Now even if I accept that that may have been so, that does not of itself establish any ground of opposition. Now in the absence of a SNP map, Professor Graham Plastow for Branhaven said that he would go to databases that existed at 2002 and look for information in the region of interest. But, so MLA contends, there is no evidence as to the extent and usefulness of any such database that existed at that time, other than Professor Peter Visscher’s indication that he would go to the publically available expressed sequence tag (EST databases), which he could interrogate to find SNPs. But even if I were to accept MLA’s point, that does not take it far. I will discuss the relevant grounds of opposition later.
186 Further, MLA points out that at [0125], the specification says that an advantage of the map, which Branhaven did not provide, is that it encompasses the whole genome. But as MLA rightly contends, no claim relates to a method of creating a high-density SNP map of a bovine genome.
187 Now MLA had to accept Professor Taylor’s explanation that the exercise undertaken by the inventors in mapping the shot-gun sequences from four bovine subjects to the human genome was “non-trivial”. But MLA contends that his explanation did not suggest that what was done was beyond the calling of a skilled worker. It was simply a lot of hard work. Further, MLA contends that the person skilled in the art need not have done what the inventors did to reach the alleged invention. An alternative would have been to sequence and assemble the whole bovine genome which, as Dr Tad Sonstegard explained, would have taken Celera about 26 days. Indeed, it was Celera that did the sequencing disclosed in the 253 Application. Further, MLA says that after being invited by counsel for Branhaven to explain what technical difficulties would arise in this exercise, Dr Sonstegard could identify little other than the need to set up a DNA sequencing facility and pipelines for processing the sequencing information, much like what was done in the human genome project, but without the need to map the whole genome and instead assemble it against other animal/human genomes, which were only matters of time and cost. MLA says that the fact that Branhaven spent the time and money to do this on a very large scale does not entitle them to a patent that uses the “map” that it generated in a known way to predict the genetic potential of animals. In a loose sense I accept that the inventors used the human genome work as a “blueprint” for the work on the cattle genome. I will return to these interesting questions later when I discuss in more detail the question of inventive step. But before proceeding further, I should say something further on the human genome project.
188 As I have said, the inventors of the 253 Application created their high density genetic map of the bovine genome using the shotgun sequencing approach described by Venter (at [0053]). MLA has asserted that it was obvious to do so on that basis. As it pithily submitted, “work in humans leads the way”. It even goes so far as to assert that the notional person skilled in the relevant art, instead of adopting the approach of comparative mapping to the human genome that was employed by the inventors of the 253 Application, could have sequenced and assembled the entire bovine genome as was done in the human genome project. But as at the priority date of the 253 Application, the evidence before me clearly established that there were significant technical challenges facing breeders and those working in the field of bovine molecular genetics.
189 Now there is little doubt that the sequencing of the human genome was a noteworthy scientific achievement. In 1998, a private venture group announced its intention to build a unique genome sequencing facility to determine over a three year period the sequence of the human genome. The group was headed by J Craig Venter. The sequencing was performed by a whole-genome random shotgun method with subsequent assembly of the sequenced segments (Venter, 1305 and 1306). The human genome assembly was published in February 2001, but at that time the “detailed manual curation and interpretation of the genome” was “just beginning” (Venter, 1306).
190 The following concluding observations were made in Venter (at 1345):
Experience in applying the whole-genome shotgun sequencing approach to a diverse group of organisms with a wide range of genome sizes and repeat content allows us to assess its strengths and weaknesses ... With more complex genomes like those of … or human, map information, in the form of well-ordered markers, has been critical for long-range ordering of scaffolds. For joining scaffolds into chromosomes, the quality of the map (in terms of order of the markers) is more important than the number of markers per se. Although this mapping could have been performed concurrently with sequencing, the prior existence of mapping data was beneficial.
191 The above observations make it clear that it was important to the success of the work that existing map information in the form of well-ordered markers was available.
192 Further, I would also note that the sequencing of the bovine genome itself did not commence until March 2003. Even then, version 1 of the bovine genome was not published until about mid-2005. Further, work on even a “working draft” of the bovine genome had not yet begun, let alone being made available as any well-ordered marker map as at 31 December 2002 (the earliest priority date) or as at 31 December 2003 (the deferred priority date).
193 The history of the bovine genome sequencing project and its impact is usefully summarised in Womack JE, “The Impact of Sequencing the Bovine Genome” (2006) 46 Aust J Exp Agr 151, which observed that the sequencing of the bovine genome was the culmination of more than a decade of international collaboration to bring together resources to chart the genome. Even as at 2006 much work remained in annotation of the genome and the discovery of DNA polymorphisms within and between breeds.
194 Finally on this aspect, Branhaven’s experts also distinguished the sequencing and mapping approach described in the Venter paper from the comparative mapping approach described in the 253 Application, a topic I will return to later.
(b) Step 2 – Identification of bovine SNPs associated with traits
195 MLA says that there was nothing new in any of the analysis involving step 2.
196 The specification describes how the relevant phenotypic data from cattle was collected (see [0054]). Using this data, SNPs in the high-density SNP map that had a statistically significant association with targeted traits were identified, in “whole genome association studies” ([0054]). At [0080], the specification says that known statistical methods can be used to identify relationships between SNPs and a trait.
197 The specification identifies the traits tested at [0055]: marbling, tenderness, fat thickness, yield, and daily gain. 2,510 SNPs were identified as purportedly being associated with these traits. The SNPs and the traits with which they were associated (at a p-value better than 0.01) are identified in Tables 1A and 1B (which MLA says were not disclosed in the priority document) ([0055]). Those SNPs are at position 300 of SEQ ID NOS: 19473 to 21982 (filed on compact discs at the time of filing the PCT application that became the 253 Application) ([0056]).
198 Example 2 describes in greater detail the association study that identified which of the bovine SNPs in the map created in Example 1 were associated with one of five traits of interest. DNA samples were taken from 4,791 cattle with different phenotypes for each trait and pooled for each biological type of animal and each trait. Each pool was tested against each of the 6,189 SNPs mapped and validated (in the sense described by Professor Taylor). A Chi square statistical test was used to analyse associations between SNPs and each trait ([0195] to [0197]). None of the associations were validated outside the discovery population. MLA says that it would still be necessary to determine an association between these SNPs and the five specific traits in a different population in order to validate the initially observed association, thereby confirming that the SNP/trait association would be useful outside of the discovery population. MLA says that the manner in which this information was collected and analysed casts doubt on the validity of any associations asserted in Tables 1A and 1B.
199 I will return to these questions later, particularly in the context of lack of utility.
(c) Step 3 – Determination of the distance of linkage disequilibrium (LD) in cattle
200 Now the specification at [0030] introduces the idea that the single SNP can be one that is within about +/- 500,000 or less nucleotides of a SNP position corresponding to position 300 on one of Sequence ID Nos 19473 to 21982, where those SNPs at those positions are described in the specification as “associated SNPs”.
201 At [0035], the specification states:
the linkage disequilibrium for cattle is about 500,000 nucleotides. Therefore, it is expected that other SNPs can be identified that are associated with the same traits based on the fact that these other SNPs are located less than or equal to about 500,000 nucleotides of the identified associated SNPs on the bovine genome.
202 This is based on Example 3 as demonstrating that the extent of LD for cattle is about +/- 500,000 nucleotides ([0208]). In Example 3, some SNPs adjacent to the identified SNPs were found to be associated with the same trait as a specified SNP ([0054], [0201] to [0206]).
203 But MLA contends that the way in which the extent of LD has been determined is flawed. MLA contends, based upon the evidence of Professor Hayes, that multiple SNPs may be associated with the same trait but not in association with each other if there are multiple QTLs. Indeed, MLA says that this is very likely for quantitative traits, which have numerous genes affecting the expression of the phenotype.
204 MLA points to [0208] of the specification that states:
The results of this Example [3] indicate that disequilibrium in cattle exists across the region of 500,000 nucleotides from an associated SNP, in each direction. Therefore, it is expected that when an associated SNP is identified, other markers within this 500,000 bp region will also be in disequilibrium with the associated SNP and with the trait of interest, and can be used to infer associations with the trait of interest.
205 In other words, so MLA contends, the specification is saying that SNPs within the about +/- 500,000 nucleotide region will be in LD with the specified SNP and with the trait of interest, and therefore that region is being used as a proxy for LD.
206 MLA says that this is confirmed at [0126], where the specification states that “SNPs in linkage disequilibrium can be used in lieu of determining a SNP or mutation to predict the presence or absence of a phenotypic trait or contributor to a phenotypic trait”. It is said that similar statements are made at [0035] and [0052].
207 Indeed, MLA says that there are numerous general statements made in the specification as to the invention extending to SNPs that are within (about) 500,000 nucleotides of the specified SNPs, without reference to LD at all: see e.g. [0029], [0030], [0036], [0038], [0131], [0132], [0134] and [0135]. MLA says that to the extent these paragraphs also require that the non-specified SNP must be in association with the trait, such association does not mean that the non-specified SNP will be in LD with a specified SNP. MLA says that it may be in LD with another QTL for the trait.
208 Further, at [0127], the specification states that:
it has been determined that disequilibrium exists across the region of 500,000 bp from the associated SNP in each direction. Other markers within this 500,000 bp region will also be in disequilibrium with the associated SNP and with the trait of interest, and can be used to infer associations with the trait of interest. Genomic segments containing the markers can be adjacent to the associated SNP marker or contained within a separate island of sequence distant from the associated SNP.
209 MLA says that in the context of the other teachings in the specification, the “Other markers” in the second sentence is describing that all other markers within that region are taken to be in LD with the specified SNP.
210 Further, MLA says that the specification explains ([0128] et seq) that the non-specified SNPs can be identified by whole-genome shotgun sequencing (the method of the inventors), BAC-sequencing or sequence generated from comparative maps. Further, the non-specified SNPs can be genotyped in pools of animals and associated with traits as the inventors did. In other words, so MLA contends, to identify the non-specified SNPs and associate them with a trait, the skilled person must repeat everything the inventors did.
211 Further, MLA rejects Branhaven’s suggestion that LD is significant because it was necessary for the inventors to determine LD before identifying an association between the specified SNPs and traits. MLA says that it is plain from the face of the specification that Examples 1 and 2 were conducted before Example 3. Example 3 began with associated SNPs identified as a result of Examples 1 and 2.
212 I will return to the question of LD shortly as it has relevance, inter-alia, to a good point raised by MLA concerning limb (b) of claim 1.
Construction of the claims
(a) Principles of construction
213 The principles governing the construction of patent specifications, including claims, are well established. A claim is construed from the perspective of a person skilled in the relevant art as to how such a person would have understood the patentee to be using the words of the claim in the context of the specification as a whole. Further, a claim is to be construed in the light of the common general knowledge including the art before the priority date.
214 A measure of common sense should be used. And ordinary words should be given their ordinary meaning unless a person skilled in the art would give them a technical meaning or the specification ascribes a special meaning.
215 In terms of how the body of the specification may be used in construing a claim, the claim should be construed in the context of the specification as a whole even if there is no apparent ambiguity in the claim. Nevertheless, it is not legitimate to narrow or expand the boundaries of the monopoly as fixed by the words of a claim by adding to these words glosses drawn from other parts of the specification. More particularly, if a claim is clear and unambiguous, to say that it is to be read in the context of the specification as a whole does not justify it being varied or made obscure by statements found in other parts of the specification.
216 Now the specification may stipulate the problem in the art before the priority date and the objects of the invention that are designed to address or ameliorate this. Accordingly, the specified objects may be useful in construing a claim in context. Nevertheless, the specified objects are not controlling in terms of construing a claim; glosses cannot be drawn from the objects.
217 A claim should be given a purposive construction. Words should be read in their proper context and a too technical or narrow construction should be avoided. Further, the integers of a claim should not be considered individually and in isolation. Further, a construction according to which the invention will work is to be preferred to one in which it may not. But to give a claim a purposive construction “does not involve extending or going beyond the definition of the technical matter for which the patentee seeks protection in the claims” (Sachtler GmbH and Co KG (formerly Sachtler AG) v RE Miller Pty Ltd (2005) 221 ALR 373; [2005] FCA 788 at [42] per Bennett J). To apply a purposive construction does not justify extending the patentee’s monopoly to the ideas disclosed in the specification. I also adopt what was said in Artcraft Urban Group Pty Ltd v Streetworx Pty Ltd (2016) 245 FCR 485 at [72] to [78] per Greenwood J (agreed to by Rares J at [142], [145] and [146]). Further, I would also refer to Lord Hoffmann’s observations in Kirin-Amgen Inc v Hoechst Marion Roussel Ltd (2004) 64 IPR 444; [2004] UKHL 46 at [27] to [34] concerning a purposive approach to construction.
218 As I have said, a claim is to be construed from the perspective of how a person skilled in the art would have understood the patentee to be using the words, informed by the notional skilled addressee’s relevant general knowledge and what has been disclosed in the specification. But to consider such a perspective does not entail that the Court necessarily requires expert evidence to assist on construction. If it is clear that the claims are to be read according to their ordinary meaning with no special meaning given to any word or phrase, if the science or technical issues are easily comprehensible and if, more generally, the Court does not require expert assistance in understanding the context of the claims, then expert evidence on construction may not only be unnecessary, but unhelpful and distracting. The nature and complexity of the patent in suit and the issues raised will determine the utility or necessity for expert evidence on construction. In the present case, I have to some extent been assisted on questions of construction by the expert evidence adduced by the parties, but the significance and weight of such evidence should not be over-stated. After all, the proper construction of a claim is ultimately a question of law for me, albeit that I must adopt the perspective that I have just described.
219 In terms of the skilled addressee, one is using a hypothetical construct. The following principles are applicable:
(a) First, to identify the characteristics of the skilled addressee, the field to which the invention relates must be identified.
(b) Second, the skilled addressee is taken to be a person of ordinary skill (as opposed to a leading expert) in that field and equipped with the relevant common general knowledge including the art before the priority date.
(c) Third, the qualifications and experience of the skilled addressee will depend on the particular case, having regard to the nature of the invention and the relevant industry. Formal qualifications are not essential. Practical skill and experience in the field may suffice. A patent specification is addressed to those having a practical interest in the subject matter of the invention; such persons are those with practical knowledge and experience of the kind of work in which the invention is intended to be used.
(d) Fourth, the hypothetical person skilled in the art may possess an amalgam of attributes drawn from a team of persons whose combined skills, even if disparate, would normally be employed in interpreting and carrying into effect instructions such as those contained in the specification.
(e) Fifth, as the skilled addressee comes to a reading of the specification with the common general knowledge of persons skilled in the relevant art, they read it knowing that its purpose is to describe and demarcate an invention. But the person skilled in the art is not particularly imaginative or inventive.
(f) Sixth, the skilled addressee does not come to reading the specification seeking failure.
220 As I have said, the legal construct may not be a single person but may be a team of persons whose combined skills would normally be employed in that art in interpreting and carrying into effect instructions such as those contained in the relevant instrument.
(b) Person skilled in the art
221 As I have said, a patent application should be construed through the eyes of the hypothetical person (or team of persons) who is likely to have a practical interest in the subject matter of the invention and may often work in the art with which the invention is connected (KD Kanopy Australasia Pty Ltd v Insta Image Pty Ltd (2007) 71 IPR 615; [2007] FCA 481 at [16] per Kiefel J (as she then was) and GlaxoSmithKline Consumer Healthcare Investments (Ireland) (No 2) Ltd v Apotex Pty Ltd (2016) 119 IPR 1; [2016] FCA 608 at [277(c)]). In the present context in relation to construing the 253 Application, the person skilled in the art includes both a molecular geneticist with experience in livestock genetics, particularly cattle, and a quantitative geneticist with some experience in animal breeding programs. A molecular geneticist would be concerned with the structure and function of genetic material, with physically isolating and manipulating genetic material for the purposes of improving the genetic potential of livestock, with identifying and mapping microsatellites, SNPs and the like, and with undertaking gene association analyses. But given the nature of the invention claimed in the 253 Application, the molecular geneticist in the present context would work with a quantitative geneticist. A quantitative geneticist is someone who has expertise in the use of statistical models and methodology to study genetic data. In my view a quantitative geneticist would also be part of the notional team. Let me elaborate.
222 In the present case, the field of the invention includes the study of associations that exist between naturally occurring polymorphisms in genes and known and desired traits of animals. This involves the discovery of naturally occurring genetic markers, the design of an association study to test for associations between traits and the genetic markers, the collection of data from the study and the use of known statistical methods to analyse that data, the validation of the associations between genetic markers and traits and, finally, the genotyping of an animal for validated genetic markers to predict the animal’s value for the trait, including for the purpose of breeding. As I have said, the persons skilled in the relevant art for this purpose would be molecular geneticists and quantitative geneticists with experience in animal breeding programs. As I have indicated, the molecular geneticist would conduct the physical aspects of the study, such as discovering genetic markers and genotyping animals for those markers and obtaining the data for use in the association study, and the quantitative geneticist would be involved in the design of the association and validation studies and perform analyses of the data in order to determine associations and understand their accuracy and utility for sorting animals for their potential value for traits of interest, including for the purpose of breeding.
223 Now Branhaven accepts that the notional team would comprise a quantitative geneticist and molecular geneticist but it contends that the skilled addressee is primarily a molecular geneticist with experience in livestock genetics, particularly cattle. But in my view the specification and claims are also concerned with associations between genetic markers and traits, a quantitative geneticist’s work. In the present context being considered, in my view the role of the quantitative geneticist is not secondary to that of the molecular geneticist. Generally speaking I would rank them equally in that sense.
224 Conformably with that conclusion, I would observe in respect of the evidence called before me that where a topic has related to molecular genetics, I have preferred the expert opinion of a molecular geneticist in preference to that of a quantitative geneticist. Likewise, where a particular topic has related more to quantitative genetics, I have preferred the views of a quantitative geneticist over that of a molecular geneticist. I make this observation as it is relevant as to why on some topics I have preferred one set of experts called by one party over the other set of experts called by the other party. Let me elaborate on the expert evidence called by the parties in terms of the individual witnesses, their qualifications and their reliability. I would say at the outset though that I found all expert witnesses to be highly talented and expert in their particular fields.
(c) MLA’s expert witnesses
225 MLA adduced evidence from three quantitative geneticists and a patent attorney.
Professor Michael Goddard
226 Professor Michael Goddard is a quantitative geneticist with over 30 years’ experience in area of the genetic improvement of livestock, in particular cattle. He received a Bachelor of Veterinary Science in 1972 from the University of Melbourne and completed a PhD in animal genetics at the University of Melbourne in 1977. From 1977 to 1983, Professor Goddard was a lecturer in biometrics at James Cook University. From 1983 to 1993, he worked at the Victorian State Department of Agriculture, carrying out genetic improvement research of dairy cattle, beef cattle and sheep, leading that research from 1988 to 1993. Since 1988, he has been chairman of the Genetics Committee of the Australia Dairy Herd Improvement Scheme. From 1988 to 2014, he was a member of the National Beef Cattle Genetics Advisory Committee of Meat Livestock Australia. Since 1993, Professor Goddard has held several positions concurrently. In 1994, he was employed as Director of Animal Genetics and Breeding Unit at the University of New England, leading and carrying out research in the genetic improvement of beef and dairy cattle and pigs. From 1993 to 1998 and from 2000 to 2012, he was the program manager of three Cooperative Research Centres in the area of genetics and beef cattle. From 2008 to 2012, he was chief scientist of the Cooperative Research Centres for Beef Genetic Technologies, responsible for the quality of the research program. In 1998, Professor Goddard was appointed a professorial fellow at the University of Melbourne, heading research projects in the area of the genetic improvement of traits in livestock, including cattle. Since 1998, he has held a joint position with the University of Melbourne and the Victorian State Department of Agriculture in which role he teaches animal genetics and had led and carried out research in genetic improvement of dairy cattle, beef cattle, sheep and pigs.
227 Professor Goddard has published widely in the field of livestock genetics, in particular the use of DNA information in the genetic improvement of livestock. In 2015, he was elected a Fellow of the Royal Society.
228 Generally, his evidence on contentious issues concerning quantitative genetics was reliable. However, without descending into the detail, I do not consider that the AV/Goddard patent application (Australian patent application No 2007335195 that I describe later) and its contents sits particularly well with Professor Goddard’s evidence concerning the “chilling effect” of the 253 Application, his evidence concerning limb (b) of claim 1, particularly given that on p 4 it is stated “[m]arkers in linkage disequilibrium are generally within about 1 cM to 5 cM of a locus of interest” (cf limb (b) of claim 1 which uses 0.5 cM), and his evidence concerning the use or potential use as at the priority date of genome-wide selection using dense marker maps (cf the problems he discusses in 2007 in the AV/Goddard patent application at pp 4 and 5).
Professor Benjamin Hayes
229 Professor Benjamin Hayes is a quantitative geneticist, in particular a complex traits geneticist. I found his evidence useful and reliable in the field of quantitative genetics. In 1995, he received a Bachelor in Agricultural Science from The University of Queensland and completed a PhD in the area of livestock breeding at Central Queensland University in 2000. He has more than 15 years’ professional academic experience in the area of the genetic improvement of livestock and crops. He has held several professional positions in the field of genetics. From 2001 to 2003, he was a quantitative geneticist at the Victorian Department of Natural Resources and Environment, and then was senior researcher in genetics and breeding at Akvarforsk Genetics, Norway until 2005. Since 2005 he has held a position as a research leader and principal scientist, Computational Biology and Biosciences Research at the Victorian Department of Primary industries.
230 Since 2016, he has been a professor at the Queensland Alliance for Agriculture and Food Innovation at the University of Queensland, having been associate professor in the faculty of Life Sciences at Latrobe University from 2009 to 2016.
231 His research has included genetic improvement of livestock, crop, pasture and aquaculture species, with a focus on integration of molecular information into breeding programs for genetic enrichment of crops and livestock. He has published widely in journals including Nature, Science and Nature Genetics.
Professor Peter Visscher
232 Professor Peter Visscher is a quantitative geneticist, in particular a complex trait geneticist. He also gave evidence on quantitative genetics that I found to be of assistance. He received a Bachelor of Science from the Dronten Higher Agricultural College (Netherlands) in 1986 and a Master of Science degree in animal genetics from the University of Edinburgh in 1988, and completed a PhD in quantitative animal genetics from the University of Edinburgh in 1991. From 1992 to 1993, he was a research fellow at the Victorian Institute of Animal Science, and then worked as a senior research fellow from 1994 to 1995 at the Roslin Institute (Scotland), an animal science institute. From 1995 to 2005, he lectured at the University of Edinburgh. From 2005 to 2011 he was the senior principal research fellow at the Queensland Institute of Research Medicine, before being appointed professor and chair of quantitative genetics at the University of Queensland in 2011. His research has included quantitative and population genetics, genetic epidemiology, genomics, human genetics and bioinformatics. He has published widely in journals including Nature, Science and Nature Genetics. In 2010, he was elected a Fellow of the Australian Academy of Science.
Dr Scott Whitmore
233 Dr Scott Whitmore is a patent attorney at Phillips Ormonde Fitzpatrick, where he is a partner of the Chemistry and Life Sciences group. Although his affidavit was tendered, he was not cross-examined. I need say nothing further at this point.
General
234 Now each of MLA’s experts were quantitative geneticists who did not have significant direct experience in undertaking laboratory work in the application of molecular genetic techniques, including the molecular genetic techniques disclosed in the 253 Application. Further, from my assessment of the evidence, MLA’s experts appeared to accept that the identification and genotyping of markers, the sequencing of DNA and the construction of a genetic marker map for the bovine genome were endeavours that lay more within the field of a molecular geneticist rather than a quantitative geneticist.
235 I raise the matter at this point because MLA’s case on inventive step had as an important foundation that the steps that a molecular geneticist would have had to take to construct and characterise an informative SNP map for the entire bovine genome were a matter of routine. In that context, MLA’s witnesses’ lack of direct hands on experience (as distinct from administrative supervision of others) in the application of molecular genetic techniques is not unimportant. That lack of relevant experience is reflected in their evidence on this issue. For example, Professor Goddard, although conceding that he was not a molecular geneticist, asserted in relation to the method described in the 253 Application that there were no technical hurdles that needed to be overcome in order to identify a sufficient number of SNPs. Similarly, Professor Visscher asserted that the techniques to find and identify SNPs were well known by reference to various unnamed publications. Further, such a lack of relevant experience is also reflected in the apparent inconsistencies pointed to by Branhaven in the evidence of Professors Goddard and Visscher concerning the technical difficulties involved in performing molecular genetic techniques. I do not need to elaborate further for the present context. On the question of molecular genetics, I have generally preferred the evidence of Branhaven’s witnesses concerning SNP mapping and identifying SNPs.
(d) Branhaven’s expert witnesses
236 Branhaven adduced evidence from two molecular geneticists and a quantitative geneticist, each of whom provided reliable and helpful evidence in their fields.
Dr Tad Sonstegard
237 Dr Tad Sonstegard is a molecular geneticist with over 20 years’ experience in livestock genetics. He received a Bachelor of Science (agricultural biochemistry) from Iowa State University in 1987 and completed a PhD in cellular, developmental and molecular biology and genetics at the University of Minnesota in 1995. From 1997 to 2015, Dr Sonstegard worked as a research geneticist in the Agricultural Research Service at the United States Department of Agriculture (USDA), where his research focused on improving cattle breeding using genetic technologies. In the period of 1997 to 2003, he worked on developing new genetic markers and using them to map regions of the genome associated with quantitative traits.
238 Dr Sonstegard was also the project leader in the project that led to the development of a fibre optic beadchip that specifically assays SNP markers at over 53,000 locations in the bovine genome. The methodology was central to the development of a commercial genotyping chip known as the BovineSNP50 in 2009, which is still the core material on a number of standard genotyping tools used in most bovine genomic selection studies. For that work, he was awarded the USDA, ARS Technology Transfer Award, the Federal Labs Consortium Award for Excellence in Technology Transfer and the Secretary of Agriculture’s Award for Superior Service in 2009.
239 Dr Sonstegard’s experience both before and after the priority date made him well qualified to give before me what I considered to be compelling evidence in relation to the nature of the work required in practice in SNP discovery and characterisation.
Professor Graham Plastow
240 Professor Graham Plastow is a professor in the Department of Agricultural, Food and Nutritional Science at the University of Alberta, whose current research is focused on the genomics of animal health in beef and swine and feed efficiency in beef and dairy cattle. He received a Bachelor of Science (biological sciences) and a PhD in molecular genetics from the University of Leicester in 1977 and 1981 respectively.
241 From 1983 to 1996, he worked in various roles at Dalgety plc, a British agri-food company, ultimately managing the central research and development of its agri-food business. In particular he undertook genomics research to improve pig breeding, and became responsible for the genomics program at Dalgety subsidiary, Pig Improvement Company (PIC).
242 From 1996 he was biotechnology manager for PIC, and when PIC became part of Sygen International in 2001 he became that company’s chief technology officer until 2005, responsible for its research and development programs.
243 Since 2011, he has been the chief executive officer of Livestock Gentec, a Canadian centre for livestock genomic research based at the University of Alberta. He has more than 30 years’ practical experience in the management and implementation of multidisciplinary research projects in the field of molecular genetics (specifically in the agri-food sector), and more than 20 years’ experience working with many of the world’s largest pig, cattle and poultry breeding companies.
244 His evidence on molecular genetics on contentious questions was particularly helpful to me.
Professor Jeremy Taylor
245 Professor Jeremy Taylor is a quantitative geneticist and the Curators’ Distinguished Professor of Genetics and Animal Sciences and the Wurdack Chair in Animal Genomics at the University of Missouri. He received a Bachelor of Science (mathematical statistics) from the University of Adelaide in 1979 and completed a PhD in quantitative genetics at the University of New England in 1982.
246 From 1982 to 1984, he was a postdoctoral research associate in the Department of Animal Science at Cornell University. From 1984 to 1986, he lectured in biometrics and animal production at James Cook University. From 1986, he was associate professor in the Faculty of Genetics and Department of Animal Science at Texas A&M University, and was professor there from 1993 to 1999.
247 In 1999, he founded a livestock genomics company, Genomic FX. Its goal was to commercialise genetic diagnostics that could be used to determine livestock’s predisposition to levels of expression of economically important traits. From 2001 to 2002, he was Director of Genomics and Associate Director of Bioinformatics with the Research Triangle Institute, before taking up a position at the University of Missouri.
248 He has authored and co-authored a large number of articles and papers on cattle genomics and breeding strategies. He has been a member of the Editorial Board for Animal Genetics and acted as a reviewer for a number of relevant publications. He has also been a member of advisory committees for a number of research consortia focused on the breeding and genetic improvement of livestock, including the BovineSNP50 iBMC Consortium from 2005 to 2009.
249 I should also note that Professor Taylor has also trained and worked in a molecular biology laboratory and currently runs such a laboratory. Accordingly his evidence in both the fields of quantitative genetics and molecular genetics was reliable.
(e) Dr Kerr’s declaration
250 Before proceeding further, it is convenient at this point to dispose of another evidentiary question.
251 MLA sought leave to tender before me a statutory declaration of Dr Richard Kerr (one of the inventors for the 253 Application) dated 3 November 2014 that had in fact been relied upon by Branhaven before the delegate below. MLA contended that leave ought be given pursuant to r 34.31(1) of the Federal Court Rules 2011 (Cth) to admit the declaration, alternatively, that section 190(3) of the Evidence Act 1995 (Cth) should apply, on the basis of the following matters. First, the matters to which the evidence relates could not genuinely be in dispute, given that it was evidence put on by Branhaven in the hearing below by a named inventor. Second, Branhaven has been on notice of MLA’s proposal to rely on the declaration since 9 May 2017. Third, the declaration was made “conscientiously believing the statements contained in this declaration to be true and correct”. Fourth, the object of providing a “quick and efficient inexpensive” procedure (that is, the appeal before me) is achieved by allowing MLA to rely on the declaration on its face without the need to call Dr Kerr. Fifth, Branhaven has not adduced any evidence of prejudice and nor could there be since it has been on notice since 9 May 2017 of MLA’s intention to rely on the declaration, and Dr Kerr apparently resides in Hobart and was willing and able to assist Branhaven during the opposition.
252 The matters upon which MLA seeks to rely arising from the declaration are as follows. First, the only inventor that Branhaven tendered evidence from in the case below was of a quantitative geneticist. Second, in his declaration, Dr Kerr does not assert any invention in the creation of the bovine “map” produced by the patent applicants. Third, in his declaration, Dr Kerr does not suggest that any new technique was involved in the association study. Fourth, Dr Kerr does not assert that the invention requires linkage disequilibrium to be proven between a specified and non-specified SNP, only that “we thought that any other SNP within 0.5 cM (or within 500,000 bp) of a SNP showing association is expected to be in LD with the same causal mutation”. Fifth, Dr Kerr says that the minimum of three SNPs was chosen simply as being a sensible minimum (less than which would capture non-quantitative traits) and he does not mention the requirement that they occur in at least two genes.
253 I would reject the tender of the declaration as it is not admissible evidence.
254 At the outset I would note that the basis on which MLA has sought to tender the declaration has changed. It originally relied on s 81 of the Evidence Act but quite rightly no longer presses this; Dr Kerr is not relevantly a person who could be taken to have made an admission on behalf of Branhaven and it was misconceived to suggest that merely because the declaration was tendered by Branhaven before the delegate that the declaration or part thereof could by such a circumstance alone be transmogrified into an admission. MLA now relies on r 34.31(1) of the Federal Court Rules or s 190(3) of the Evidence Act in support of the tender of the declaration. But as to this, I would make the following observations.
255 First, as to r 34.31(1), the predecessor to that rule, being O 58 r 8(1) of the previous Federal Court Rules 1979 (Cth), did not of itself render evidence admissible which would otherwise have been inadmissible: see Commissioner of Patents v Sherman (2008) 172 FCR 394 at [34]. Further, the difference in wording between O 58 r 8(1) and r 34.31(1) (the phrase “and saving all just exceptions” does not appear in the present r 34.31(1)) does not change this. Like the former rule, r 34.31(1) provides that the evidence is admissible “with the leave of the Court”. That wording is still consistent with the operation of the Evidence Act. In any event, a minor change to the wording of a rule of Court could not of itself override the Evidence Act. The substance of the reasoning in Sherman is unaffected by the change. That minor change in wording is simply a consequence of the different formal approach to the drafting of the current rules. The extrinsic material indicates that the introduction of the Federal Court Rules in 2011 was not intended to bring about any substantial alteration to the previous rules’ operation: Voxson Pty Ltd v Telstra Corporation Ltd (No 7) (2017) 343 ALR 681; [2017] FCA 267 at [11] and [20] per Perram J. The express requirement in r 34.31(1) that leave of the Court be obtained before the tender of material recognises that the tender requires consideration by reference to substantive and procedural law (including the Evidence Act) that might be applicable in the circumstances. Similarly, the statutory requirements imposed by the Evidence Act in relation to evidence are not to be subjugated to the requirement for a relatively quick and inexpensive procedure (i.e. the appeal process). Only admissible evidence tendered to the Court in the de novo hearing can be considered subject to the operation, inter-alia, of s 190(3) of the Evidence Act.
256 Second, as to s 190(3) of the Evidence Act, the matters on which MLA seeks to rely from the declaration are either genuinely in dispute (s 190(3)(a)) or irrelevant. The following matters are genuinely in dispute. First, whether there was inventiveness in the creation of the bovine SNP map. Second, whether the association study by the inventors involved any “new technique”. Third, whether “limb (b)” of claim 1 requires LD with a specified SNP. Fourth, whether the “at least three SNPs” in claim 1 is an arbitrary parameter. Now s 190(3)(a) asks whether “the matter to which the evidence relates” is genuinely in dispute, which directs attention to the substance of the evidence. It is apparent that MLA wishes to rely on the declaration as supporting its substantive submissions on the matters listed. In any event, whatever probative value there might be in the declaration is far outweighed by the prejudice Branhaven would suffer if it were admitted now. It has been given no reasonable opportunity to test the matters that MLA says arise from the declaration by way of cross-examination of Dr Kerr or through participation by Dr Kerr in the concurrent evidence session which addressed those matters.
257 Now MLA asserts that Branhaven could suffer no prejudice because it was on notice from 9 May 2017 of MLA’s intention to rely on the declaration, but this was less than 4 business days before the start of the hearing. It was also after the time for the filing of evidence that had been ordered. Further, MLA’s submission that Dr Kerr “was willing and able to assist” before the delegate does not advance the matter.
258 Moreover, the usual rule is that it is for the party who seeks to rely on evidence of a witness (i.e. MLA) who must make that witness available for cross-examination by the other party.
259 Further, MLA does not appear to rely on s 190(3)(b) of the Evidence Act, but the provision would not assist it in any event. The application of the hearsay rule will not cause or involve any unnecessary expense or delay as it will simply mean that the declaration is not admitted into evidence.
260 Finally, in the context of considering s 190(3) I would make some other points. Evidence untested in cross-examination from an inventor on the above issues could have only secondary significance at best on questions such as construction (which is from the perspective of the hypothetical person skilled in the art with the relevant common general knowledge) and inventive step (see my decision in BlueScope Steel Limited v Dongkuk Steel Mill Co., Ltd [2017] FCA 1537 at [31] to [38]). I have more direct and probative evidence from six expert witnesses called before me. Moreover, to allow reliance upon the declaration now would likely open up whether other material before the delegate should be tendered before me. Some reasonable limits and finality must be imposed in relation to this appeal. These are further points as to why I would not exercise any discretion I have under s 190(3) in favour of MLA. Further and for completeness, MLA did not seek to invoke ss 64, 67 and 68 of the Evidence Act.
261 I reject MLA’s tender of the declaration of Dr Kerr and refuse the leave sought.
(f) Terms requiring construction
262 The 253 Application contains 15 claims defining the invention. Each claim needs to be considered separately and independently for the purpose of assessing the various grounds of opposition. Moreover, each claim must be construed as a whole. Further, it is trite to observe that the integers of a claim in combination determine its scope and it is the interaction between those integers to produce a new result that is the essence of the claimed invention.
263 The principal claim for present purposes is claim 1, which is in the following terms (inserting “[:]”, “[a]” and “[b]” as explained by me earlier for clarity):
A method for identifying a trait of a bovine subject from a nucleic acid sample of the bovine subject, comprising identifying in the nucleic acid sample an occurrence of at least three single nucleotide polymorphisms (SNPs) wherein the at least three SNPs are associated with the trait, and wherein the at least three SNPs occur in more than one gene[:]
[a] and wherein at least one of the SNPs corresponds to position 300 of any one of SEQ ID NOS: 19473 to 21982, or
[b] the SNP is about 500,000 or less nucleotides from position 300 of any one of SEQ ID NOS: 19473 to 21982.
264 The principal issues that arise relating to the construction of claim 1 and its ambit are or concern the following:
(a) The meaning of “a method for identifying a trait” and also the scope of “for” in that phrase.
(b) The meaning of “at least three SNPs are associated with the trait”.
(c) The meaning of “at least three SNPs occur in more than one gene”.
(d) The requirement of at least three SNPs in more than one gene.
(e) The requirement that at least one SNP be a specified SNP or a non-specified SNP.
(f) The operation of limb (b) and whether it is necessary that the SNP referred to in limb (b) be in linkage disequilibrium (LD) with the specified SNP (i.e. the SNP referred to in limb (a)).
265 Now the various aspects of claim 1 that I have identified cannot be considered in isolation. The claim is directed, first, to a method for identifying, in the sense of inferring as I later explain, a trait in a bovine subject. That purpose informs the nature and scope of other aspects of the claim, including the requirement that the at least three SNPs identified from the nucleic acid sample of the subject be associated with the trait. That requirement of association is to be understood having regard to the purpose for which the SNPs are to be used. So, the association must be suitable to enable the method to be carried out by drawing an inference of the kind referred to in the claim. Further, limb (b) of claim 1 should also not be considered in isolation. The claim requires that each of the at least three SNPs used in the claimed method be associated with the trait of interest. If the SNPs are not so associated, then there will be no use of the claimed method. But having said that in all its generality, in my view it will be necessary to amend the claim so that limb (b) makes it plain that relevant LD is stipulated. Otherwise the claim lacks clarity and does not properly define the invention. I will elaborate on this later.
266 More generally on this question of association, I would also note that it is the association that I have just identified, which is a limiting and defining feature of the claim, that gives rise to the ability to infer or predict the potential for the trait in the subject. Now Branhaven says that the degree of association involved is left to the skilled person’s implementation of the claimed method in a particular case, having regard to the application to which the method is to be put (and the accuracy etc, desired) and the guidance given in the body of the specification. Branhaven contends that the words “associated with” are a classic example of a workable standard suitable to the intended use. I would note at this point that I agree with its submission in one sense but not in another. In my view, the claim will need to be amended to stipulate the degree of statistical significance required in order to establish the required “association”. And usually and appropriately that would be stipulated at a p-value of equal to or less than 0.05, but in this case desirably at equal to or less than 0.01; I will hear counsel further on such matters if Branhaven applies to amend claim 1. If statistical significance is not stipulated, there is in my view no adequate measure for or objectivity enshrined in the concept “association” or its cognate verbial forms. Subjectivity and imprecision would rule the day. In my view, absent a proper stipulation for statistical significance, the claim lacks clarity and the invention has not been properly defined.
267 Now the claim is further limited by the requirement that the at least three SNPs occur in more than one gene, and that at least one of the SNPs be among the 2,510 SNPs identified in limb (a) or fall within limb (b) referring to the +/- 500,000 nucleotide region around the limb (a) SNPs. There is an issue as to whether limb (b) is merely concerned with the distance of the SNP referred to from a specified SNP, or whether it entails that the respective SNPs be in LD with each other. In my view, for the claim to make any sense to the skilled addressee and to conform to the relevant association required by the prefatory words, LD is required for limb (b). As I have already indicated, LD should be required and stipulated in limb (b) to avoid problems dealing with a lack of clarity and a failure to define the invention.
268 Claims 2 to 10 and 15 of the 253 Application are similarly method claims. The other claims are to bovine subjects, systems and isolated polynucleotides resulting from or relating to the methods of the invention.
269 Claim 2 limits the identification of a trait of claim 1 to the use of at least one specified SNP (i.e. one of the specific SNPs identified in Table 1A referred to at [0027] of the 253 Application). Claims 3 to 5 are dependent on claims 1 or 2, and require the use of at least five, seven or ten SNPs respectively. I would note that MLA contends that these further limitations are arbitrary, as it similarly contends in relation to the three SNPs and more than one gene integers of claim 1. I should say at this point that I have rejected all of MLA’s assertions concerning arbitrariness and parameteritis as I explain later.
270 Claim 6 concerns a method for sorting bovine animals using the identification as described in claim 1. Claim 7 concerns a method for cloning a bovine animal shown to have a desired trait by using the identification of a trait described in claim 1. Claim 11 is to the animal cloned according to the method of claim 7. Claim 8 is a method for the same identification of a trait as claim 1, where occurrence of the SNPs is identified using a hybridisation technique. Claim 12 is to a system for determining SNPs that uses the same feature as claim 8. Claims 9 and 10 embrace the method of any one of the preceding claims, but specify certain traits. Claim 10 is limited to the five traits tested. Claim 13 is to isolated DNA identified according to claim 8. Claim 13 was found by the delegate to be unclear. Claim 13 was also found by the delegate to not constitute patentable subject matter. As I elaborate on later, I have formed views similar to the delegate on claim 13. Claim 14 is to isolated DNA of at least 20 nucleotides in length when used in any of claims 1 to 10. This claim also gives rise to potential problems in terms of patentable subject matter that I discuss but dismiss later. Claim 15 is a so-called omnibus claim.
(i) The meaning of “identifying”
271 In my view the word “identifying” in the phrase “a method for identifying a trait” refers to drawing an inference about the potential for the trait to exist in a bovine subject using the claimed method. This meaning is clear from the context of the claim. It also reflects a workable and common sense understanding of the claim from the perspective of the notional skilled person.
272 Moreover, I do not consider that the use of the word “identifying” instead of “inferring” in the claim gives rise to any difficulty of interpretation. The skilled person would well understand that the claim is not directed to a method for identifying as a matter of fact or certainty that the trait actually exists in the relevant subject. The use of genetic markers is only ever capable of offering a prediction about the existence of the trait.
273 Now the experts had differing views as to the meaning of this term.
274 Branhaven’s experts, whose evidence I preferred on this aspect, considered that “identifying” was meant to refer to a method for inferring a trait, that is, predicting the genetic potential of an animal to possess a trait. Moreover, given that other experts such as Professor Goddard and Professor Visscher could not make proper sense of the term “identifying” in its context, this all supports my view that the skilled addressee would construe “identifying” as meaning “inferring”. Now admittedly there is a difference in language as compared with the use of the word “inferring” in claim 8. MLA contends that it would be unusual to give the same meaning to two different terms used in the specification and claims. And for that reason alone, MLA says that the term is unclear. But in my view it is well apparent that, in context, the terms “identifying” and “inferring” were used as synonyms and the skilled addressee would so understand.
275 Further, MLA says that whether the word “identifying” or “inferring” is used, strictly speaking it is not appropriate language to use in relation to quantitative traits (e.g. marbling, or growth rate). This is said to add to a lack of clarity. But I consider that such a submission has an element of linguistic preciousness about it that in my view a skilled addressee is unlikely to indulge in. If “identifying” is to be taken as meaning “inferring”, which I have concluded, I see little practical difficulty with it being used in the context of considering quantitative traits.
276 For completeness I note that before the delegate, MLA submitted that as at the priority date, it was not possible to identify a trait. It was only possible to predict or infer whether a SNP was associated with a trait. Contrastingly, Branhaven contended that the phrase “identifying a trait” was synonymous with “inferring a trait”. The delegate noted that the title of the 253 Application referred to “inferring bovine traits”, and that the specification contained definitions for “infer” and “inferring” when used in reference to a trait. But there was no definition for “identifying a trait”. Further, the delegate accepted the evidence of Professor Plastow, who stated that the skilled person reading the 253 Application would regard the phrase “identifying a trait” to be the same as “inferring a trait”. Based on the disclosure in the specification and the evidence of Professor Plastow, the delegate considered that “identifying a trait” had the same meaning as “inferring a trait”. I have taken a similar view on the evidence before me and my own consideration of the specification including the claims.
277 There is a further question that it is convenient to address at this point relating to the meaning of the word “for” in the phrase “method for identifying a trait”.
278 MLA has contended that this feature only means that the method must be suitable for or capable of use for inferring a trait. This means that if the prior art does not specifically teach that use, it does not matter, assuming that the method is otherwise suitable for that purpose.
279 MLA has referred to Pharmacia & Upjohn AB (opposition by CSL Limited) [2000] APO 58, where the delegate considered a similar phrase in relation to a process of filtration: “for removing at least one virus selected from the group comprising of hepatitis A, polio virus or parvo virus”. The delegate accepted that this phrase merely meant that the filtration method was capable of removing one of the listed types of virus and that viral particles may not necessarily be present in the solution to be filtered, noting “that is consistent with the nonlimiting way that the term ‘for’ has traditionally been construed by the courts in Australia”. MLA asserts that a similar view has been taken in respect of “Swiss-style” claims which provide for the use of compound X in the manufacture of a medicament for a specified and new therapeutic use. In Otsuka Pharmaceutical Co Ltd v Generic Health Pty Ltd (No 4) (2015) 113 IPR 191; [2015] FCA 634 (Otsuka), an appeal from which was dismissed (Otsuka Pharmaceutical Co Ltd v Generic Health Pty Ltd (No 2) (2016) 120 IPR 431; [2016] FCAFC 111), Yates J queried whether, for the purpose of determining infringement of such a claim, it matters that the alleged infringer does not advertise or promote the medicament specifically for the therapeutic use defined in the claim. His Honour said (at [172]):
I do not think it necessarily does. The question is whether, objectively ascertained, the medicament that results from the claimed method or process is one that has the therapeutic use defined in the claim. The question is not really about how the alleged infringer markets its product, although, plainly, its conduct in that regard may well assist in determining, objectively, whether the accused product has the claimed therapeutic use.
280 MLA says that by analogy when considering the anticipation of a Swiss-style claim, if there is disclosure of the manufacture of compound X, it does not matter that the disclosure does not refer to the therapeutic use; it is enough that it is capable of that use. Similarly, for claim 1 of the 253 Application, MLA says that it does not matter whether the anticipatory document describes using the disclosed method to infer or identify a trait or sort animals, as long as it is capable of doing so.
281 I would reject MLA’s ambitious submission.
282 In my view, the word “for” in this context means more than that something must be “suitable for” use in the method claimed. The word “for” in the expression “method for identifying a trait” places a limit on what is claimed such that the method claimed must result in being able to identify, in the sense of infer as I have discussed, a trait. It is clear in context that claim 1 stipulates a purpose constraint. It is not simply that the method must be suitable for identifying a trait. Now on MLA’s case, it does not matter if the prior art does not actually teach the claimed method; a disclosure of something that could be used for that purpose would be sufficient. But I agree with Branhaven that this is misconceived. The word “for” might be given such a construction in the context of a product claim, where the word can be understood as intended to characterise the construction of the product by reference to its suitability for the use to which it is to be put. But in the context of a method claim, such a construction is inapposite. Such a construction involves disregarding what is an important limiting and characterising feature of the claim.
283 In Otsuka, Yates J observed that the reference in a “Swiss type” claim to the use of a substance in the manufacture of a medicament “for” a specified therapeutic purpose is intended to impose the kind of limitation that MLA seeks to avoid, which is a limitation that confers novelty over a disclosure of the substance itself. As Branhaven points out, that is the reason why Swiss type claims were developed. And the point made by Yates J at [172] rather related to a different point. In my view, it is not in doubt that a claim to a method “for” treating a particular condition or disease involves a purpose limitation, such that a product that is merely suitable for that purpose will not infringe the claim, and will not anticipate if it is the subject of a prior disclosure. Similarly, the use of such a product for a different purpose will not infringe or anticipate. What is necessary to consider is a use or disclosure of the method itself. The same considerations apply to the method of claim 1 of the 253 Application.
284 I will return to this question again when I discuss novelty.
(ii) The meaning of “associated”
285 What is meant by the word “associated” in the phrase “at least three SNPs are associated with the trait”? Branhaven contends that these words require that the at least three SNPs identified from the nucleic acid sample of the bovine subject be associated with the trait of interest. It says that the context indicates that the association must be a meaningful one. It says that the body of the specification confirms that the association must be statistically significant.
286 Branhaven says that there is no difficulty in the use of the phrase “associated with” in the claim. It says that the idea of an association between a genetic marker and a trait of interest is a well-understood concept that the experts were readily able to deal with. It says that it is a concept that underpins molecular genetics and genomics generally and that the evidence indicates that it is a concept that is readily capable of being applied by the skilled person to the circumstances of a particular case.
287 Now Branhaven had to accept that no degree of association is specified in the claim. But it is said that the draftsperson is not required to use only exact expressions and that general or relative expressions are common in patent claims. It is said that a lack of precise definition is permissible so long as the claim provides a workable standard suitable to the intended use.
288 Branhaven says that the body of the specification provides guidance to the skilled person as to the nature of this requirement. At [0051], the specification states:
A SNP is associated with a trait when at least one nucleotide occurrence of the SNP occurs more frequently in subjects with a certain characteristic of the trait in a statistically significant manner, for example with greater than 80%, 85%, 90%, or 99% confidence.
289 Branhaven says that the work conducted by the inventors in generating a high density SNP map of the bovine genome employed a confidence level of 0.01 or greater [0052] for the association of the SNPs with a trait. But it says that the claim is not so limited.
290 Branhaven says that the claim requires the existence of a statistically significant or meaningful association with the trait of interest, but does not place any precise limit on the “degree” of association required. It is said that this is appropriate given the nature of the claimed method. It contends that the incorporation of a hard limit would arbitrarily and artificially restrict the claim, and that the level of accuracy required of the method and thus the degree of association that is appropriate to put it into effect will vary from case to case. It is said that this will depend on the application to which the method is to be put including matters such as the purpose of drawing the inference, the size of the target population, the nature of the trait of interest, the number of SNPs to be employed in the method, and the like. It contends that the evidence confirmed that the skilled person would readily be able to address these matters in a particular case.
291 Branhaven says that it is not the case that the claim covers any degree of association between the SNPs and the trait when determined by any means. It is said that the association must be such as to allow the claimed method to be carried out according to its terms: i.e. to enable the skilled person to draw an inference about the potential for the trait to exist in the subject. It says that the accuracy required of that prediction is a matter for the skilled person in the particular case and that this is the result of a practical and common sense construction of the claim.
292 It also contends that there is no substance to the contention that the claim is unclear because the skilled person will not know whether the particular associations between SNPs and traits fall inside or outside the scope of the claim. It says that to so contend misunderstands the subject matter of the claim. It is said that the claim does not claim particular associations between SNPs and traits. It claims a method of identifying a trait in a bovine subject, based on the identification from a nucleic acid sample of the bovine subject of at least three SNPs associated with the trait. It is said that the skilled person will readily understand when he or she is employing such a method, and identifying SNPs for that purpose. It says that the degree of association will depend on the particular case.
293 Branhaven further contends that there is no territory left outside the claim by reason of the incorporation of the requirement that the SNPs be “associated with the trait” which the skilled person would wish to exploit. It is said that this is not a case of the skilled person being unable to draw a line between infringement and non-infringement in any practical sense.
294 I must say that I have difficulty accepting Branhaven’s contentions.
295 Before I proceed further, let me be clear as to what is being precisely talked about here. I am concerned to consider “association” in terms of a correlation with a trait. I am not concerned at this point with the quantum of the percentage variation in a trait to be explained by a SNP(s). But as soon as one talks of correlation, one really needs to consider statistically significant correlations. After all, a correlation that has no statistical significance is useless as an objective measure or observation of anything. It is a correlation in the ether or a perceived correlation infected with subjective imprecision. In terms of the skilled addressee, he must be able to know or ascertain what is the correlation within claim 1 and what is the correlation without claim 1. Moreover, an invention where the correlation was not required to be statistically significant would, in my view, fail for lack of utility. The promise of the invention would be illusory and its fulfilment a fantasy.
296 Let me now make the following observations.
297 First, it is apparent that the claim as drafted includes any SNP covered by the claim that has any degree of association with any trait. In my view the claim lacks clarity because although a non-zero association might be determined by one study, it might be found not to have any association at all when tested using a different study. Moreover, the fact that the claim includes any degree of association also gives rise to lack of utility.
298 Second, even if the term “associated” requires a particular degree of association, which in my view would be to add an impermissible gloss, then the claim still lacks clarity. The person skilled in the art will not know the degree of statistical significance required to establish the requisite association. So in my opinion whether or not the claims require a particular degree of association, there is no workable standard in the claim for a person skilled in the relevant art to determine if a SNP that they are using is “associated” with a trait.
299 Now Branhaven confidently asserts that the measure of statistical significance or level of confidence is a matter that can readily be determined by a molecular geneticist. Now in one sense this is true. But in another sense it does not answer the lack of clarity question. Reasonable minds may differ as to the degree of association and statistical significance required. How is one to tell whether what one is doing falls inside or outside of claim 1? In my view claim 1 should be sufficiently clear on this point, which at the moment it is not.
300 In my view, statistical significance should be stipulated in a meaningful way and capable of objective assessment. Conformably with relevant aspects of the specification, a p value of equal to or less than 0.01 suggests itself; perhaps one could be more generous to Branhaven at a p value of equal to or less than 0.05, but I doubt it. I will leave that question open for further argument if Branhaven applies to amend claim 1. As to whether each of the 3 SNPs should separately satisfy the relevant p value or only the combination of the three, I will leave that for discussion on another occasion if necessary.
301 Finally and for completeness, I note that before the delegate, MLA had contended that there was a conflict in the specification regarding statistical thresholds for association. On the one hand, [0052] described an “associated SNP” as a SNP associated with a trait at a confidence level at 0.01 or greater. On the other hand, [0080] described SNPs associated with a trait as having statistical association of at least 80% to 99% confidence. Branhaven submitted that there was no conflict between the two paragraphs, because [0080] described a method of using a bovine SNP map to identify SNPs associated with traits. It was not seeking to define an “associated SNP”. The delegate was satisfied that [0052] related to the confidence level for identifying a SNP associated with a trait, whilst [0080] described a possible application of statistical methods to determine relationships between SNP nucleotide occurrences associated with traits. She construed the phrase “associated with a trait” to mean that the SNP was statistically determined to be significantly associated with a trait.
302 I have taken a slightly different approach. In my view that phrase should have been so expressed with some stipulation for the degree of statistical significance. But it has not been so expressed. Accordingly, the claim lacks clarity and does not properly define the invention absent some appropriate amendment to deal with my concern. I do not consider at all that it is appropriate to leave this to the skilled person. Some objective stipulation of statistical significance and the appropriate degree is necessary.
303 I will hear further from counsel on that question if Branhaven applies to amend.
(iii) The meaning of “gene”
304 What is meant by “gene” in the phrase “at least three SNPs occur in more than one gene”? Claim 1 requires that at least two SNPs must be present on one gene and at least one SNP must be present on a different gene. It also encompasses three SNPs that are present on three different genes.
305 Now the experts each attributed different meanings to “gene” at the priority date. Although the experts agreed with the self evident proposition that “gene” would include exons, they had different views as to whether introns and regulatory regions would also be included in the meaning of the word.
306 MLA contends that the specification does not provide any guidance at all on this matter. Furthermore, the specification does not explain why it is necessary for the SNP to be in a gene (or for that matter in more than one gene). MLA contends that it is not possible for the person skilled in the art to have known, in December 2002, whether a SNP was in a gene, especially in the absence of any explanation in the specification as to why there must be “at least three SNPs occurring in more than one gene”.
307 Further, MLA contends that the variation in meanings attributed to the word “gene” by the experts means that the term does not provide a workable standard by which a third party will know whether or not they infringe the claim. It contends that this is made worse by the fact that there was no complete bovine genome mapped at the priority date, let alone a database of bovine genes identified within that map. This meant, so MLA contended, that one would need to sequence and map the bovine genome to be somewhere along the way to determining whether or not all relevant genes had been accounted for and thus know whether the at least three SNPs were in more than one gene as required in the claims. And relying upon Professor Goddard’s evidence, MLA contends that in 2002 you could not be sure that a SNP did or did not appear in a gene given there was no whole bovine genome and given the limitations in the comparative mapping process.
308 Further, MLA contends that the 253 Application is insufficient because it was not possible to determine whether a SNP fell within a gene in December 2002 without undue experimentation, as the sequence of a bovine genome was not available at that time and was not published until 2009. Further, even though it was possible to do comparative mapping, not all genes in the human genome have equivalents in the bovine genome. I will discuss this lack of sufficiency assertion later in my reasons.
309 At this point I reject MLA’s construction arguments. I will deal with the other dimensions to MLA’s arguments later.
310 First, merely because different views are expressed by experts and more than one construction is open does not entail that the claim lacks clarity. Further, it is a molecular geneticist as opposed to a quantitative geneticist, who is responsible for identifying and determining the location of genetic markers in the genome. Accordingly, on this construction question I prefer Branhaven’s experts. A reasonable definition of “gene” was given by Professor Plastow, a molecular geneticist, as simply “a unit of inheritance associated with a biological function in [an] animal”. I would note that even accepting that knowledge of the components of a “gene” has expanded over time, the term has been used to describe the basic unit of heredity since 1906, the year the term “genetics” was apparently coined.
311 Second, if I need to go further, in my view it would include both exons and introns pre-splicing, the mechanism for which I have described earlier. It would also include necessary regulatory regions. It seems to me that a person skilled in the art would at least so construe “gene”, although it may be debated whether regulatory non-coding aspects of DNA would be included and the true boundary of any particular “gene”. I would note, if it matters, that some support is given for including introns (that is, a gene can be considered pre-splicing and has the potential to give rise to multiple transcripts depending upon how splicing occurs) by the implications flowing from the following passage in Venter (at 1317) and albeit that Venter itself was not part of common general knowledge:
A gene is a locus of cotranscribed exons. A single gene may give rise to multiple transcripts, and thus multiple distinct proteins with multiple functions, by means of alternative splicing and alternative transcription initiation and termination sites. Our cells are able to discern within the billions of base pairs of the genomic DNA the signals for initiating transcription and for splicing together exons separated by a few or hundreds of thousands of base pairs. The first step in characterizing the genome is to define the structure of each gene and each transcription unit. (my emphasis)
312 For completeness, let me say something about the proceedings before the delegate. I note that experts for both parties gave evidence before the delegate that the requirement that “the at least three SNPs occur in more than one gene” meant that each SNP was located in a different gene. Because the specification did not define “gene”, the delegate concluded that the term was to be given the meaning that the skilled person would give it in light of common general knowledge and context. However, MLA contended that because the boundaries defining a gene could be difficult to establish, a skilled person might misunderstand the requirement that the SNPs occur in more than one “gene”. Its expert Dr Barendse stated in his third declaration that “more than one gene” meant “more than one location of the genome”. But the delegate considered that this description conflicted with a previous declaration of Dr Barendse, which described “gene” in terms that were not contradicted by Professor Plastow and were approved by the delegate:
In my view, a gene generally refers to both introns and exons and associated regulatory regions (such as promoters) of a particular coding sequence, and DNA immediately 5’ and 3’ to the coding sequence (the 3’ and 5’ untranslated regions) are involved in the regulation of the gene. However, it is difficult to establish with certainty the precise bounds of the 5’ and 3’ untranslated regions.
313 From my own consideration of the evidence, I would conclude that the skilled addressee would construe the term “gene” in a similar fashion. And I do not consider there to be any lack of clarity in that respect.
314 Let me deal with another matter at this point. MLA contends that whatever “gene” means, no scientific or other justification is given in the specification for this limitation. Put another way, MLA contends that this feature is merely an arbitrary parameter, and cannot provide a basis for novelty and inventive step. It says that it was well known that the majority of bovine traits are quantitative traits that are influenced by many genes. Therefore, you would need at least one SNP associated with each of these influencing genes to predict genetic potential, and you might need a large number of SNPs (each associated with a different influencing gene) in order to predict genetic potential with any degree of accuracy. I will deal with this question in part in the following section.
(iv) The requirement of at least three SNPs in more than one gene
315 MLA says that there is no scientific or other justification given in the specification for the requirement that you need to have at least three SNPs, nor that they need to occur in more than one gene. It asserts that these requirements are entirely arbitrary and that the claims are infected with parameteritis. It is worth dealing with the matter at this point, although the context for raising it relates more to MLA’s attacks concerning, inter-alia, novelty and other grounds of opposition that I will discuss later.
316 At [0075] of the 253 Application, the specification states that the term “at least one” when used in reference to a SNP means any number “up to and including all of the … SNPs of the bovine genome”; see also e.g. [0027] (“at least one bovine SNP”), [0051] (“at least one bovine SNP”), [0071] (“for at least one single nucleotide polymorphism”), [0072] (“1 or more SNPs”), [0091] (“using one SNP or a population or series of SNPs”), [0095] (“of at least one single nucleotide polymorphism”), [0097] (“of at least one single nucleotide polymorphism”), [0107] (“at least one nucleotide occurrence of a single nucleotide polymorphism”), [0110] (“at least one nucleotide occurrence of a single nucleotide polymorphism”) and [0147] (“typically at least one SNP or haplotype … nucleotide occurrences for at least one SNP in the DNA samples”). The same is said to apply for “at least two” or the like; see also e.g. [0073] (“nucleotide occurrence of at least 2 SNPs. At least 2 SNPs can form all or a portion of a haplotype”), [0075] (“Reference to ‘at least a second’ gene, SNP, or the like, means two or more”), [0098] (“at least one or at least two SNPs for the bovine subject”), [0108] (“nucleotide occurrence of at least 2 SNPs can be determined. The at least 2 SNPs can form a haplotype”), [0113] (“can be at least two SNPs that influence the trait”) and [0114] (“at least 2 SNPs are identified for inferring the genetic potential”). Further, at [0106] the specification explains that a series of SNPs can include “at least 2, 3, 4, 5, … 6000 markers, for example”. No explanation or preference is given. The only mention of at least two SNPs occurring in different genes is at [0114] (“In certain examples, at least 2 of the single nucleotide polymorphisms occur in different genes”) and [0142] (“in embodiments where SNPs that affect the same trait are identified that are located in different genes”). But no explanation is given as to why this would be preferred.
317 MLA contends that Dr Sonstegard could only guess at why a minimum of three SNPs was chosen. Further, Professor Plastow stated that if he were applying the method of the 253 Application, he would use as many of the markers as he could, which MLA contends supports the conclusion that the at least three SNPs requirement is an arbitrary limitation.
318 Accordingly, MLA contends that the requirement that at least three SNPs are in more than one gene is irrelevant to the invention and that it is likely to have been added to avoid a finding of lack of novelty or inventive step. MLA refers to Otsuka Pharmaceutical Co Ltd v Generic Health Pty Ltd (No 2) (2016) 120 IPR 431; [2016] FCAFC 111, where the Full Court (Besanko, Nicholas and Beach JJ) upheld the primary judge’s conclusion that a particular integer of the claim(s) in suit was an arbitrary limitation. Further, some reliance was placed on my observations at [115] and [116] to the following effect:
But even if I had accepted the appellants’ construction, the patent would still be invalid for, inter alia, lack of novelty. By the use of the said phrase, assuming that on this hypothesis it constituted a separate integer, claims 1 and 7 would be infected with what patent lawyers would diagnose as parameteritis. This affliction involves an attempt to re-patent the prior art by limiting claims by reference to a series of parameters not mentioned in the prior art (Raychem Corp’s Patents [1998] RPC 31 at 37 per Laddie J) The parameter could be something measured on test equipment, which equipment did not exist at the time of the prior art. In the present case, the parameter is a statement in essence of a scientific theory positing a link between relevant disorders of the central nervous system and the 5-HT1A receptor subtype. But to inject this parameter adds nothing to the invention. It does not create a new process or method. It does not create a new use of an old product. In substance, one still has an old use of an old product or a more limited class of an old use of an old product. Scientific knowledge may have been enhanced by the identified association, whatever “association” or “associated” means, between the receptor subtype and the disorder. But there is no new invention by the addition of such knowledge to the claim language or using it as a limitation.
Indeed, the artificiality of such a result and the nebulous verbiage of the claim language gives me confidence in my first conclusion that the phrase “disorders of the central nervous system associated with [the] 5-HT1A receptor subtype” is not a separate and essential integer.
319 Further, in Williams Advanced Materials, Inc. v Target Technology Company LLC (2004) 63 IPR 645; [2004] FCA 1405, Bennett J said in relation to the patent in suit that (at [48]):
there is nothing in the specification that suggests that the proportions or the ranges of the metals in the alloys are in any way part of the invention, other than the mere reference to them …
She accordingly diagnosed a case of “parameteritis”.
320 MLA contends that the same applies in the present case. It contends that there is no suggestion that the requirement that there be at least three SNPs occurring in more than one gene is in any way part of the invention.
321 I reject MLA’s contentions. I do not consider that the stipulation of at least 3, 5, 7 or 10 SNPs to be arbitrary parameters. Likewise I do not consider the stipulation of at least more than one gene to be an arbitrary parameter.
322 First, there is no need for any particular scientific or technical justification to be given in the specification for what is a limiting feature of the claim. And even if a feature was included so as to narrow the claim and avoid an argument about the prior art, that does not of itself defeat a patent.
323 Second, it is difficult to see how the requirement of multiple SNPs could be said to be arbitrary. After all, part of MLA’s expert evidence was to the effect that you needed to identify multiple SNPs to do anything useful in terms of inferring association with a quantitative trait. Now true it is that the particular arithmetic stipulations in a simplistic sense are arbitrary (in the sense that, 2, 4, 6 or 8 might perhaps equally have been stipulated). But they are not relevantly arbitrary, particularly given that the claim had to have clarity and to properly define the invention.
324 Third, there is support in the specification for the limitation to “at least three SNPs” in the claim. The use of at least three SNPs in the method is encompassed by the broad disclosure of the use of SNPs in passages in the specification, including in the summary of the invention. The claim to a method involving the use of at least three SNPs is simply a narrower embodiment. Further, the specification addresses the use of greater numbers of SNPs in the method of the invention (using the example of two SNPs associated with a particular trait) and explains why this might be done, namely, to infer a higher likelihood that the trait will exist.
325 Fourth, I also do not consider the more than one gene requirement to be an arbitrary parameter. Indeed, most of the experts readily accepted that when one was dealing with quantitative traits, multiple genes were involved. The requirement limits the claim by requiring that the SNPs employed be located in two or more genes. The requirement that the SNPs be located in genes is consistent with the way in which the high density bovine SNP map was created by the inventors.
326 Finally, the arithmetical stipulation for the SNPs and the more than one gene requirement it must be recalled are properly words of limitation. And as I have said, there are various references in the specification supporting the use of and requirement for multiple SNPs and more than one gene (see for example [0058] and [0114]), although I accept that an example is not given for 3 or more SNPs in more than one gene. But the specification did not need to give such an example, and its absence does not make parameteritis symptomatic.
(v) At least one SNP must be a specified SNP or a non-specified SNP
327 Claim 1 requires that at least one SNP is a specified SNP (being a SNP identified in SEQ ID NOS: 19473 to 21982 of Table 1A, the specified SNPs) (limb (a)), or a SNP that is within +/- about 500,000 nucleotides of a specified SNP (a non-specified SNP) (limb (b)). For present purposes I will accept that “about” means +/- 10% and so one is referring to a range of +/- 550,000 nucleotides. For convenience, however, I will just refer to the region of +/- 500,000 nucleotides (or use alternative terminology such as 500 kilobases or 500 kb). The evidence shows that the region so covered by the claim is in a strict sense around two thirds of the bovine genome, although this is more a jury point of MLA when one comes to appreciate that the non-specified SNP should be in linkage disequilibrium with the specified SNP (as I will require by way of amendment if one is applied for and granted).
328 As to the other two SNPs (of the at least three SNPs), they can be anywhere in the bovine genome provided that the at least three SNPs occur in at least two genes. Of course, the limitation is that one of the 3 SNPs has to be a limb (a) or limb (b) SNP, and all 3 must be associated with the same trait.
329 Now, I accept that the claim includes many SNPs that were not known as at the priority date, given that the bovine genome had not yet been sequenced and many SNPs were yet to be discovered. MLA has contended that as a consequence the claim is very broad. I must say that for my part this is another jury point. The breadth of a claim per se is not a separate ground for invalidity and nor does breadth entail a lack of clarity or a lack of definition. A claim may be broad, yet have clearly defined boundaries. But I will consider MLA’s breadth point later when it has relevance to and in the context of specific grounds of opposition.
(vi) The operation of limb (b) and linkage disequilibrium (LD)
330 Before examining in detail some of the arguments concerning limb (b) and whether the limb (b) SNP needs to be in LD with the limb (a) SNP, it is appropriate that I say something briefly concerning the evidence.
Some evidence
331 The experts gave evidence on the topic, some of which it is necessary to set out.
332 Professor Goddard made the following statements in relation to the specification and claims of the 253 Application:
(a) It appeared that the use of the region of +/- 500,000 nucleotides from position 300 was derived from Example 3. The tables in Example 3 conclude that SNPs within this region would be in LD with the specified SNPs.
(b) The region of +/- 500,000 nucleotides from position 300 was intended to provide a proxy measurement for LD. Given the use of this proxy, he said that it would have been unworkable to imply a further requirement that there be LD between the two SNPs.
(c) There was no disclosure of other SNPs in the region that were not associated with the same trait, but he expected there to be many such non-associated SNPs. Further, he considered that there was no disclosed rationale as to why position 300 was selected. It was likely that had another SNP been chosen as the reference position, the conclusions would have been different.
(d) Consequently, the conclusion in the 253 Application that LD extended over +/- 500,000 nucleotides resulted from selective analysis and presentation of data.
333 Professor Goddard stated that there will on average be 10,000 SNPs within the +/- 500,000 nucleotide region. However, he considered that there was no way to assess which of the 10,000 SNPs were in LD with the specified SNP and which were not. In order to identify which of those SNPs would be useful for inferring a trait, he said that he would have to perform his own association study, effectively starting again. He said that all that the 253 Application would have given him was a “very large” region from which to choose a SNP.
334 Professor Goddard considered that Branhaven’s experts imputed a requirement that a non-specified SNP be in LD with a specified SNP. He stated that this requirement only made sense if the specified SNP was associated with the relevant trait and not just with a specified SNP. The non-specified SNP needed to be in LD not only with the specified SNP but also the relevant QTL. He said that just because the specified SNP is in LD with the QTL does not mean that the non-specified SNP is also. He said that this meant that the requirement imputed by Branhaven’s experts remained unclear and unworkable.
335 Further, Professor Goddard stated that a requirement that the non-specified SNP be in linkage disequilibrium with the specified SNP was “unworkable”, because there is no indication of what degree of LD was meant or needed. The degree of LD was an important factor. Further, he said that as at the priority date there was no established amount of LD that was recognised as being an acceptable degree of LD.
336 Professor Visscher’s reading of the specification of the 253 Application led him to consider that the +/- 500,000 nucleotide region was being used as a proxy for LD, such that there was no separate requirement for LD to be present. He considered that the disclosed requirement that mattered was the distance, not LD. He considered that Example 3 showed an indirect method of determining LD, such that the two SNPS may not be in LD and there could be alternative explanations for their statistical association.
337 Professor Visscher considered that it was inappropriate to make assertions about LD between SNPs based upon the statistical associations between SNPs and traits shown in Example 3 and Tables 2 to 4. He said that it was possible for two SNPs to be statistically significantly associated with the same trait but for the direction of LD and the direction of the frequency difference to be inconsistent. It did not then follow that an observation that two SNPs were associated was due to LD. He said that in order to actually estimate LD as a function of distance, a sample of individual-level genotypes was necessary and, in the absence of such data, he did not consider that there was any justification for the claim that SNPs within the +/- 500,000 nucleotide region could be utilised in the claimed method.
338 Professor Visscher considered that it was necessary when discussing LD to specify the strength of the association. To merely say that two markers were in LD disclosed no more than that they are not in linkage equilibrium. If, contrary to his opinion, there was a requirement in the claim for LD, it remained unclear what degree of LD was required. If the degree of LD was low, then a very large sample size would be necessary to determine whether it was present or not, a difficult thing to accomplish experimentally.
339 Professor Hayes stated that using a +/- 500,000 nucleotide region to determine the extent of LD was flawed, because it failed to account for there being multiple QTLs in the region. For example, if there were two QTLs within the region and the specified associated SNP was in LD with the first QTL, one could select a non-specified SNP and show an association, but that non-specified SNP might be in LD with the second QTL. Only where the degree of LD between the two SNPs was 100% could one conclude that the non-specified SNP tracked to the first QTL. In terms of the 253 Application, Professor Hayes said that Example 3 didn’t show that three SNPs selected within claim 1 were picking up the same QTL.
340 Reading the claims, Professor Hayes noted that there is no mention of LD. However, he acknowledged that [0126] and [0127] of the specification suggested there to be a requirement that the non-specified SNP must be shown to be in LD with the specified SNP, albeit going to an embodiment of the invention.
341 Professor Plastow said that when discussing LD in terms of the method in the 253 Application, the degree of LD sought was dependent on the method’s practical application, i.e. whether a non-specified SNP would capture the same value that the method was trying to generate as applied. He said that even if the application of limb (b) only captured 90% of the value it would still be valuable to the customer. Consequently, if the non-specified SNP had a sufficient degree of LD for the purpose of its application, then Professor Plastow considered limb (b) to be a satisfactory substitute for the primary method using specified SNPs.
342 Professor Plastow considered that one could determine infringement of the 253 Application based upon the degree of LD. That degree need not be total and the skilled person would make a judgement call on infringement based on the strength of LD and the closer it was to 1 (or 100%).
343 Professor Plastow acknowledged that the words of the claims did not refer to LD. Nonetheless, he understood limb (b) to require that the non-specified SNP be in LD with one of the specified SNPs at position 300 and read that requirement in.
344 Dr Sonstegard understood the reference to +/- 500,000 nucleotides to mean that if the non-specified SNP was in LD with the specified SNP at position 300, that non-specified SNP would be associated with the relevant trait.
345 Professor Taylor understood that the reference to +/- 500,000 nucleotides required the non-specified SNP to be in LD with a specified SNP at position 300. Referring to [0208] of the specification (i.e. Example 3), Professor Taylor explained that the inventors concluded that disequilibrium in cattle extended over a region of +/- 500,000 nucleotides from a specified SNP. Consequently, he explained that when an SNP associated with a trait was identified, other markers within the region would be in LD with the specified SNP and the relevant trait, though not every SNP in the region would be of use.
346 Professor Taylor explained that LD was a population specific phenomenon, by which he meant that a relationship of LD in one population may not appear in another population because the degree of LD, as a function of allele frequencies at the relevant loci, is different.
Analysis – construction questions
347 Much of MLA’s concerns are related to limb (b) of claim 1. Let me deal with this directly.
348 The first issue to consider is whether limb (b) is just concerned with the distance of the SNP referred to from a specified SNP (limb (a)) or whether it requires that the two SNPs be in LD with each other. Branhaven contends that the skilled person would understand the claim in the latter sense.
349 Now in my view it is clear that the claim does not expressly refer to a requirement of LD. Nevertheless, Branhaven contends that the body of the specification makes it clear that this is what is entailed by the provision for the use of other SNPs within the +/- 500,000 nucleotide region. Branhaven says that the specification explains that SNPs within that region may be in LD with the specified SNP, and that where they are, they can be used in lieu of the specified SNP in the method of the invention (see at [0035], [0126], [0127], [0128] and Example 3 at [0198] to [0208]). Branhaven contends that adopting a purposive and common sense construction, the requirement of the claim is that LD is required between the limb (a) SNP and the limb (b) SNP.
350 Further, Branhaven contends that the weight of the expert evidence supports this construction. It says that each of Dr Sonstegard and Professors Plastow and Taylor understood the claim in this way. Further, it says that Professor Hayes’s response to the expert questions revealed a similar understanding of the claim when read in the light of the description in the body of the specification. Further, it is said that Professor Taylor did not change his view in the course of the hearing. Further, it contends that Professor Visscher read the claim too literally and Professor Goddard refused to engage with it.
351 Further, Branhaven says that it is an important matter of context that the skilled person would understand that not all SNPs within the region of +/- 500,000 nucleotides would be in LD with the specified SNP. It is said that this was clear on the evidence. It is said that the 253 Application does not say so, and references to the identification of “other markers” within the region being in LD with the specified SNP should be understood accordingly. It is said that there is no statement in the 253 Application that all SNPs within the region are in LD with the specified SNP. It is said that the skilled person simply would not expect this to be so.
352 Further, Branhaven says that MLA’s contention that the references to “other markers” in the specification should be understood, contrary to the accepted scientific fact, to indicate that all other SNPs within the region are in LD involves misreading the description and interpreting the claim in a vacuum, without regard to the background knowledge and understanding of the skilled person.
353 On the question of the construction of limb (b) and whether LD is implicit, I would reject Branhaven’s arguments.
354 I do not accept that there is an additional requirement in the claim with respect to limb (b) that LD must be demonstrated for a SNP within the about +/- 500,000 nucleotide region.
355 Professor Plastow admitted that there was no reference to LD in the words of the claim. Further, the importation of such a requirement into the claims would be adding an impermissible gloss. The specification does not define the words of the claim to have something other than their plain meaning. If LD had been a requirement, one would have expected express words to that effect.
356 Moreover, in my view if LD were a requirement of the claim, there would have been no need for the region in which the non-specified SNPs are to be found to be defined in the claim at all. It would mean that those words were redundant. I agree with MLA that a construction that would lead to such redundancy is not to be preferred.
357 Furthermore, and as MLA correctly pointed out, such a construction would be inconsistent with the specification, which explains that SNPs within +/- about 500,000 nucleotides of the specified SNPs are expected to be associated with the same traits as the specified SNPs. Such an “expectation” avoids or negates the need for SNPs within that region to be shown to be in LD with a specified SNP. Indeed, depending on the parameters by which LD is assessed, including population size, all SNPs in the genome can have some degree of LD with each other SNP, as Professor Goddard explained.
358 Further, even if there were such an LD requirement, the skilled addressee would not know how to measure it. Neither the claims, nor the specification, provide guidance as to the degree of LD required between a non-specified SNP (limb (b)) and a specified SNP (limb (a)).
359 As the experts explained to me, LD is determined by measuring the frequency at which the presence of two markers (such as SNPs) are found together within individuals in a population. LD is not an all or nothing concept, but a matter of degree. LD is measured between 0 and 1, with 0 indicating no LD (i.e. the SNPs are only found together in individuals at the expected rate) and 1 indicating perfect LD (i.e. the SNPs are always found together). If the degree of LD between a specified SNP and a non-specified SNP is very low, the non-specified SNP would not be expected to be a useful surrogate for the specified SNP.
360 In my view, Professor Plastow’s attempt to explain how LD would be assessed showed that it was dependent on what approach and outcome the research team wanted. For example, Professor Plastow said LD did not need to be 100% linkage, but could depend on “how much value you’ve captured”. He later explained that it meant “something that looked pretty in linkage disequilibrium” and “it would be a guess or using my experience to determine”. And he ultimately agreed that someone else might make a different judgment call as to what was in LD.
361 I quite agree with MLA that such a requirement cannot provide a workable standard by which a third party can assess whether or not a SNP falls within limb (b) of claim 1.
362 In my view, limb (b) fails for lack of clarity absent amendment. First, the LD requirement should be explicitly stated. Second, a meaningful degree of LD should be explicitly stated, so that a non-specified SNP (limb (b)) can be considered to be a useful surrogate for the specified SNP (limb (a)). I would also note that the limb (b) SNP (after appropriate amendment) would, of course, need to also satisfy the association integer that I have previously discussed.
363 For completeness I would note the following. The delegate construed claim 1 as describing two alternative methods. The first defined specific SNPs by their specific location, requiring the at least one SNP to be posited at nucleotide 300 of any one of SEQ ID NOS: 19473 to 21982. The second required the SNP to be about 500,000 nucleotides or less from the SNP at position 300 in those sequences. The delegate also accepted the specification’s definition of “about” as providing a 10% variance, such that the second alternative required the SNP to be located within 550,000 nucleotides of the SNP at position 300. In relation to the second alternative (what I have described in this judgment as limb (b)), the delegate was satisfied that, although claim 1 does not recite that the SNP within about 500,000 nucleotides of the SNP at position 300 is in linkage disequilibrium with the latter SNP, the skilled person would understand within the context of the specification that a SNP within about 500,000 nucleotides was equivalent to a SNP in linkage disequilibrium with the SNP at position 300.
364 But as I have indicated earlier, I have taken a different view concerning limb (b). It will need to be amended to expressly address the required LD and the meaningful degree of LD.
Analysis – perceived difficulties
365 Let me now deal with another matter, which more goes to questions of lack of sufficiency and lack of fair basis, although it is convenient to say something at this point.
366 MLA says that if a third party wanted to determine whether they fell within limb (b) of claim 1, they would need to take the step of determining whether any one of their SNPs was within 500,000 nucleotides of any of the 2,510 specified SNPs.
367 MLA pointed to the following matters. In Table 1A, the patent applicants effectively identified an average of approximately 16 “contig” sequences that were “nearby” (meaning within 500,000 nucleotides: [0036]) each specified SNP. Each contig averages only approximately 1484 nucleotides long (ascertainable from the Sequence Listings and Table 1A). Consequently, (in combination with the 600 nucleotides provided in SEQ ID NOS: 19473 to 21982) only a small amount (approximately 2.4%) of the +/- 500,000 nucleotide sequence surrounding each specified SNP was provided. Furthermore, the contigs within 500,000 either side of the specified SNPs were not provided, and the location of the contigs was only putative since it was based on the human genome as a scaffold. According to MLA, the work of Professor Hayes has shown this created many errors. In total, the contigs in the Sequence Listings only provide approximately 1.5% of the total bovine genome. Accordingly, a third party would need to conduct the “non-trivial” exercise of creating an accurate map, identify true SNPs in the map, and map their SNPs against all of the specified SNPs. Should a SNP fall within the range, the third party would then need to find a new SNP and go through the process again.
368 MLA has contended that by not providing the genomic sequence 500,000 nucleotides around each specified SNP, but having a claim that extends to capture any SNP in that region, claim 1 does not provide a workable standard by which a third party could determine whether it infringed the claim.
369 Further MLA says that although Branhaven has submitted that the invention is that panel of SNPs which is employed in the methods, systems and compositions of the claimed invention, the claims are not limited to that panel of SNPs, but extend to the use of one SNP within 500,000 nucleotides either side of any one of the panel of 2,510 SNPs and at least two other SNPs, anywhere in the genome. MLA has said that this leaves the skilled addressee to conduct the very same work the inventors had conducted to identify such SNPs and determine whether they are associated with a trait, which is not limited to the five traits considered by the inventors.
370 Now these are more arguments that go to MLA’s suggestions of a lack of sufficiency and a lack of fair basis. But it is appropriate to observe the following.
371 In my view, subject to what I have said concerning amendment to limb (b), I do not accept the suggested overwhelming difficulties that would be experienced by the skilled person in applying limb (b) of claim 1. On the evidence, the skilled person could readily identify other SNPs within limb (b).
372 It would appear that the 253 Application provides approximately 25,000 nucleotides of sequence that is within 500,000 nucleotides of each specified SNP. And as MLA’s position appears to be that one SNP occurs every 100 nucleotides in the bovine genome, there would be an abundance of additional SNPs that could be identified from within the 25,000 nucleotides of sequence provided by the 253 Application. And I agree with Branhaven that MLA’s position that finding SNPs within the 500,000 nucleotide regions defined in the claims would not be a major problem. To identify those SNPs, a skilled person could take a DNA sample from a single heterozygous animal or from multiple animals and sequence a small section of the 25,000 nucleotides. Traditional sequencing technology was able to sequence stretches of 100 to 1,000 nucleotides with a high degree of accuracy. On that basis, a single sequencing reaction could potentially identify 10 additional SNPs with a high degree of accuracy. Those additional SNPs would necessarily lie within 500,000 nucleotides of a specified SNP. No mapping would be required.
373 In the alternative, as Branhaven submitted on the evidence, if the skilled person wished to go further, the specified SNPs could be mapped to the human genome. That genome is highly conserved with the bovine genome. This could be done with comparative mapping approaches that were available at the priority date. Additional SNPs could then be identified by sequencing regions of DNA that were within about 500,000 nucleotides of the specified SNP.
374 I do not agree with MLA that the work needed in order to assess whether a SNP is in LD with a specified SNP would be very difficult. In order to assess LD in this context, it would not be necessary to conduct an association study, but rather to genotype the SNPs in the target population and assess whether they appear to be associated with each other in a non-random fashion across the population. I agree with Branhaven that such an approach was routine and well within the skill of the calling.
375 I do not consider that MLA has established these matters such that I could be clearly satisfied, but I will return to the question of insufficiency and lack of fair basis later.
(vii) Other construction findings by the delegate
376 The delegate stated that her findings in relation to the construction of claim 1 also applied to independent claims 6, 7, 8 and 12 and dependant claims that recited at least one SNP corresponding to position 300 of any one of the defined sequences or alternatively within 500,000 nucleotides of the SNP positioned at 300 in the defined sequences.
377 Before the delegate, MLA contended that claim 2 ought to be construed to include SNPs within 500,000 nucleotides of position 300 as defined in claim 1. The delegate did not consider that claim 2 imported this feature, because claim 2 limited relevant SNPs to the occurrences of nucleotides at position 300 described in Table 1A.
378 The delegate considered that claims 3 to 5 depended upon claims 1 and 2, only increasing the number of SNPs to be identified from at least three to at least five, seven or ten respectively.
379 In respect of claim 6, the delegate accepted that it embodied a method of sorting “one or more” bovines on the basis of identified SNP occurrences as per claim 1 and that, as MLA submitted, it was not possible to sort a single bovine subject from itself. Nonetheless, the delegate did not consider the claim to be ambiguous. She was satisfied that the skilled person would understand the claim to be a method of sorting and selecting bovines based on the identification of traits associated with at least 3 SNPs.
380 In respect of claims 7 and 11, MLA contended that the method claimed defined nothing more than a known process of cloning cattle, because the identification of SNPs had no working relationship with the cloning method disclosed. But the delegate considered that the skilled person would construe the claim to be a cloning method incorporating the identification of defined SNPs associated with a trait, the isolation of a progenitor cell with the trait, and the generation of a clone (the cloned animal being the subject of claim 11).
381 In respect of claim 8, the delegate concluded that the use of the term “a” instead of the term “the” in the phrase “where selective hybridization of the nucleic acid sample to the system indicates the presence of a SNP associated with a trait” did not make the claim unclear. The delegate construed the claim to be a method inferring the presence of a trait in a bovine by selectively hybridising a sample of the bovine’s nucleic acid to the system disclosed.
382 The delegate was satisfied that claims 9 and 10 narrowed the methods in prior claims to specific traits.
383 The delegate determined that claim 13 was unclear. But she considered that the intention was to define isolated and specific polynucleotides with at least one SNP in a gene where the SNP was associated with a trait and either the SNP corresponded to position 300 of any one of SEQ ID NOS: 19473 to 21982, or the SNP was about 500,000 or less nucleotides from position 300 of any one of those defined sequences. I will discuss claim 13 later in my reasons in the context of “manner of manufacture”.
384 The delegate considered claim 14 to be limited to a polynucleotide when used in any one of the methods of claims 1 to 10, and as distinct from isolated polynucleotides per se. But the claim did not extend to isolated polynucleotides with the relevant SNP at position 300. Finally, the delegate treated claim 15 as an omnibus claim.
385 For my part, having reviewed the specification and considering the evidence before me, I have reached similar views in relation to these other claims.
Manner of manufacture
386 For patentability, s 18(1)(a) of the Act requires that the invention claimed in any claim be a manner of manufacture within the meaning of s 6 of the Statute of Monopolies 1624, 21 Jac 1 c 3. Further, Schedule 1 to the Act defines “invention” as “any manner of new manufacture the subject of letters patent and grant of privilege within s 6 of the Statute of Monopolies, and includes an alleged invention”. MLA has contended that none of the claims of the 253 Application claim an invention that is a “manner of manufacture”. It asserts that each claim in substance:
(a) is to the mere discovery of a naturally occurring correlation between naturally occurring SNPs and naturally occurring traits;
(b) does not give rise to anything that is man-made or an artificially created state of affairs that has economic utility;
(c) does not fall within the boundaries of existing patentable subject matter, and in light of other matters such as the chilling effect of the grant of such claims and the desire for cohesion of the law both within Australia and with the US, such boundaries should not be extended to encompass such claims; and
(d) in any event is not patentable because on the face of the specification all that is claimed is the use of standard techniques in a manner for which the known properties of those techniques make them suitable.
(a) D’Arcy v Myriad Genetics Inc
387 In D’Arcy v Myriad Genetics Inc (2015) 258 CLR 334 (Myriad), French CJ, Kiefel, Bell and Keane JJ (at [18]) endorsed the relevant question, citing National Research Development Corporation v Commissioner of Patents (1959) 102 CLR 252 (NRDC) at 269, as: “Is this a proper subject of letters patent according to the principles which have been developed for the application of s 6 of the Statute of Monopolies?”. Section 6 of the Statute of Monopolies declared all monopolies to be void except for:
Letters Patents and Grants of Privilege for ... the sole working or making of any manner of new Manufactures within this Realm, to the true and first Inventor and Inventors of such Manufactures, which others at the time of making such Letters Patents and Grants shall not use, so as also they be not contrary to the Law, nor mischievous to the State, by raising prices of Commodities at home, or hurt of Trade, or generally inconvenient ...
388 The concept of “manner of manufacture” is to be developed on a case-by-case basis and is not susceptible to any verbal formula in lieu of the phrase “manner of manufacture”. Two necessary but not necessarily sufficient criteria for patentability of an invention are (at [28]):
1. Whether the invention as claimed is for a product made, or a process producing an outcome as a result of human action.
2. Whether the invention as claimed has economic utility.
389 The plurality observed that where the invention so far as claimed in the claim in issue falls within the existing concept of manner of manufacture, it will ordinarily be sufficient if the claimed invention satisfies those criteria. But where the claim is not within the established boundaries of what is patentable, other considerations come into play. A non-exhaustive list of these other factors was set out in the following terms (at [28]):
3. Whether patentability would be consistent with the purposes of the Act and, in particular:
3.1 whether the invention as claimed, if patentable under s 18(1)(a), could give rise to a large new field of monopoly protection with potentially negative effects on innovation;
3.2 whether the invention as claimed, if patentable under s 18(1)(a), could, because of the content of the claims, have a chilling effect on activities beyond those formally the subject of the exclusive rights granted to the patentee;
3.3 whether to accord patentability to the invention as claimed would involve the court in assessing important and conflicting public and private interests and purposes.
4. Whether to accord patentability to the invention as claimed would enhance or detract from the coherence of the law relating to inherent patentability.
5. Relevantly to Australia’s place in the international community of nations:
5.1 Australia’s obligations under international law;
5.2 the patent laws of other countries.
6. Whether to accord patentability to the class of invention as claimed would involve law-making of a kind which should be done by the legislature.
390 Factors 3, 4 and 6 were said to be of primary importance and were not mutually exclusive. Factor 5 was said to be of secondary significance. Further, it was said that such factors might also inform the “generally inconvenient” limitation in s 6 of the Statute of Monopolies. Now I would note at this point that the separate judgments of Gageler and Nettle JJ and of Gordon J did not expressly adopt what I will describe as the “other factors” approach.
391 Now various questions might be said to suggest themselves concerning the “other factors” approach. First, is this a policy-driven approach to the assessment of patentability for cases on or beyond the existing boundaries? Second, is this approach properly characterised as purposive or consequentialist or both? Third, is there a clear threshold to justify moving into such a space, and if so what? In some cases reasonable minds might differ as to whether a case is within or without existing boundaries. Fourth, has the plurality just been more transparent about the considerations to be taken into account in assessing whether new or difficult subject matter is a proper subject matter for the grant of letters patent? Fifth and further, various issues concerning the priority ranking and weighting to be given to these factors remain to be explored. For example, how are factors 3.1 and 3.2 to be ranked and weighted with factor 3.3? And what is the scope of factor 3.3? How are factors 3, 4 and 6 ranked and weighted as between themselves? How is factor 5 to be weighted with the other factors, even if it is only of secondary significance? And am I obliged to consider each and all of the factors or only some of them? For the purposes of the present case it is not necessary to answer any of these questions. That is because I do not consider that I am dealing with a new class of claim involving a significant new application of or extension to the concept of “manner of manufacture” (Commissioner of Patents v RPL Central Pty Ltd (2015) 238 FCR 27 at [118] and [119] (per Kenny, Bennett and Nicholas JJ)). But if I am wrong, I have been able to apply this “other factors” approach without answering any of these questions, and in doing so have fortified my conclusion on patentability in any event, which is perhaps unsurprising. All of these “other factors” applied to the case before me are uni-directional. They all point to patentability. Given that conclusion, the characterisation of these other factors, their priority and their weighting is of academic interest only in the present case.
392 Now before proceeding further and for the purposes of my later analysis, it is appropriate to consider a number of topics and concepts analysed in Myriad, which the parties before me rightly focused on.
What in substance is claimed?
393 All members of the Court were concerned with what in substance the claim was for when determining whether there was a manner of manufacture.
394 The claims in issue in Myriad were to an “isolated nucleic acid coding for a mutant or polymorphic BRCA1 polypeptide…”. The patentee submitted that the Court ought to treat the impugned claims as claims for a chemical compound and should not treat them as any different from any other product claims (at [27]). But the plurality in Myriad at [6] explained that “[d]espite the formulation of the claimed invention as a class of product, its substance is information embodied in arrangements of nucleotides. The information is not ‘made’ by human action. It is discerned”. This was so despite the process by which the nucleic acid sequence in Myriad was isolated involving human intervention. Further, the resulting nucleic acid sequence could not function in nature. Further, the definition of “an isolated nucleic acid” in the patent in issue in Myriad included “chemically synthesised analogs” made outside the naturally occurring environment of the nucleic acid from which the isolate was prepared. The plurality explained that it was the existence of the said information, which was the same information contained in the DNA of the person from whom the nucleic acid was isolated, which was an essential element of the invention as claimed and that the product was the medium in which that information resided (at [89]).
395 Further, Gageler and Nettle JJ considered whether the wording of the claim “properly reflects the substance of the claimed invention” (at [142]). They held that the “way in which a claim is drafted cannot, however, transcend the reality of what is in suit” (at [144]) and that “care must be taken to examine the form of claim actually made to see if it is in fact an attempt to establish a monopoly for the manufacture of a substance for a purpose for which a monopoly cannot be claimed.” (at [145]). Their Honours observed that the patentee did not invent and could not claim to have invented the process of isolating nucleic acid or the process of amplifying for genetic testing the fragment comprising the BRCA1 gene (at [146]). Accordingly, it was said to be beside the point that the isolated nucleic acid was chemically, structurally and functionally different from naturally occurring DNA from which it was isolated because the patentee had not invented or claimed a new method for isolating nucleic acid (at [158]).
396 Gageler and Nettle JJ concluded that in substance the claim was (at [160]):
a claim for a monopoly over the right to apply long-established methods for the isolation and amplification of specific nucleotide fragments to the isolation and amplification of a patient’s naturally occurring BRCA1 gene, where and if it is found upon subsequent examination that the patient’s BRCA1 gene happened to be afflicted by any of the specified mutations and polymorphisms
397 Accordingly it was not a valid claim of a manner of manufacture of a product (at [161]).
398 And on this aspect, Gordon J took a similar approach. Her Honour observed that the Full Federal Court’s finding that claim 1 was to a molecule which was structurally and functionally different to what occurs in nature did not take account of the words of the claim. Accordingly she observed (at [279]) that:
As a matter of substance, each of claims 1-3 focuses on the existence of one or more elements of an identified code: a code which is found in the nucleic acid isolated from a patient and which necessarily must be identical to the coding sequence in that patient. None of the asserted chemical, structural and functional differences identified by the Full Court play any part in the definition of the invention “so far as claimed” (ss 18(1)(a) and 40(2)(b)) in each of claims 1-3 or in the description (s 40(2)(a)) of the invention in the specification. (emphasis in original)
399 In substance then, the Court assessed what was claimed not through the lens of the chemical properties or the chemical potentiality (or lack thereof) of the claimed product, but rather through its genetic informational content.
Not man-made – no “artificially created state of affairs”
400 As the plurality had identified that the substance of the claim was to information rather than a chemical entity, they considered that it was a claim on the boundaries of what was patentable subject matter. But their Honours also considered the “artificially created state of affairs” criterion at [91]. Their Honours observed that engaging with that criterion places the question of patentability in too narrow a frame because it distracts from the central issue of “whether an essential integer of the claims, the genetic information, takes them outside the category of that which can be ‘made’” (at [91]). Thus, the central question was, having properly construed the claims, whether the claim in substance was something that was man-made. Further, the plurality at [91] went on to observe that even if the criterion of an “artificially created state of affairs” was appropriate, “the fact of the existence of the requisite mutations or polymorphisms is a matter of chance. It is not something ‘made’. It is not ‘artificially created’”.
401 Gageler and Nettle JJ took the approach of observing (at [125]) that an “artificial state of affairs” and “economic utility” are not the only considerations relevant to the question of manner of manufacture. At [127] they identified the question as being “whether the subject matter of the claim is sufficiently artificial, or in other words different from nature, to be regarded as patentable” (my emphasis). As to the sufficient degree of the artificiality or difference, their Honours observed (at [128]) that “it is necessary that the inventive concept be seen to make a contribution to the essential difference between the product and nature”. So, the subject matter of the claim must “have about it a quality of inventiveness which distinguishes it from a mere discovery or observation of a law of nature” (at [131]). Moreover, their Honours observed that, by definition, a manner of manufacture “is an artificial thing or state of affairs which involves an element of inventiveness” (at [161]). They concluded that the presence or absence of the mutations and polymorphisms in the isolated nucleic acid was the critical discovery and was the “antithesis of a man-made artificial state of affairs” (at [162]).
402 I will address later the question of whether Gageler and Nettle JJ can be taken to have added a separate threshold requirement of inventiveness as part of the concept of “manner of manufacture” as MLA seemed to suggest.
Economic utility
403 As to the question of economic utility, the plurality, in disposing of the idea that the isolation of the nucleic acid per se leads to an economically useful result, viz the treatment of breast and ovarian cancers, observed (at [85]):
The economic significance necessary to the patentability of an “artificially created state of affairs” in the sense used in NRDC is not demonstrated by stating that the artificially created state of affairs is a step along the way to a process or method itself claimed as an artificially created state of affairs of economic significance.
404 This rejection was reinforced by Gageler and Nettle JJ who said (at [164]):
Fifthly, it is not the isolation of nucleic acid, or even the isolation and amplification of the fragment comprising the BRCA1 gene, which leads to the “economically useful result” of treating breast and ovarian cancers. It is rather the first respondent’s discovery of a naturally occurring correlation between the presence of the specified mutations and polymorphisms in such a fragment (and thus in the DNA in the cell from which the fragment is derived) and an increased probability of actual or potential malignancy.
405 In the case before me, MLA has contended that that is precisely what Branhaven has done in contending that the methods claimed lead to cattle with better genetic potential or traits. It is said that there is not in form or substance any such claim in the specification. But even if there were a claim expressly worded to the breeding of cattle with better genetic potential or traits, such an expressly worded claim would in substance be no more than an attempt to patent breeding activities through the uninventive and unpatentable discovery of a natural correlation between a marker and trait, and thus could not in and of itself be patentable.
The “other factors”
406 The plurality observed that the “purpose of the Act would not be served by according patentability to a class of claims which by their very nature lack well-defined boundaries or have negative or chilling effects on innovation” (at [29]). Their Honours found that in Myriad, the contentious claim(s) lacked well-defined boundaries because the isolated nucleic acid sequence included the products applying any process, known or unknown, to isolate the nucleic acid and thus created an unwarranted de facto monopoly on all methods of isolating nucleic acids from the BRCA1 gene. Moreover, infringement could occur without the infringer being aware of it (at [93]). Indeed, these factors would have a “chilling effect” on innovation, which effect was apparent from the face of the claim.
407 In terms of this question of “chilling effect” that I will come to later on the alternative argument where I am required to consider the “other factors”, MLA has suggested that no evidence of “chilling effect” is necessary to make good the point and pointed to Myriad to support that proposition. In any event, it did adduce some evidence on this point that I will discuss later.
408 And as to maintaining to the extent feasible coherence of Australian law, the plurality pointed to Apotex Pty Ltd v Sanofi-Aventis Australia Pty Ltd (2013) 253 CLR 284 to the effect that given the established patentability of pharmaceutical products it would have been anomalous to exclude medical treatments using such products. Now in the case before me, MLA has contended that the converse should apply in the sense that it would be anomalous if a claim to a product was not patentable (that is, applying Myriad in all its force so it was asserted), but claims to the use of that product in a method claim were patentable especially where that involves the use of known techniques. I will dispose of this argument later. But it leads on to the next point.
The application of naturally occurring phenomena to a particular use
409 A distinguishing feature of the present case, putting to one side claim 13, that is not unimportant is that I am dealing with method claims. All judges in Myriad not only explained that they were not addressing such claims, but by implication that such claims on their face may perhaps be more readily seen as within the existing boundaries of “manner of manufacture”.
410 The plurality observed that claims to the use of the isolated nucleic acid as opposed to the product itself were not in issue in the proceeding. So at [71] they said:
A number of sections of the specification relate to methods for the use of nucleic acids in various ways and the preparation of recombinant or chemically synthesised nucleic acids and vectors. The claims in the patent relating to those matters are not in issue. Nor is there any question about the utility of the applications of isolated nucleic acids reflected in those undisputed claims.
411 And at a more general level they observed at [20]:
It is true that in Anaesthetic Supplies Pty Ltd v Rescare Ltd Lockhart J in the Full Federal Court, in a passage endorsed by Crennan and Kiefel JJ in Apotex, said:
If a process which does not produce a new substance but nevertheless results in “a new and useful effect” so that the new result is “an artificially created state of affairs” providing economic utility, it may be considered a “manner of new manufacture” within s 6 of the Statute of Monopolies.
Importantly, however, his Honour used the word “may”. Neither Lockhart J nor Crennan and Kiefel JJ should be read as holding that satisfaction of that formula would mandate a finding of inherent patentability. That is not to say that it will not suffice for a large class of cases in which there are no countervailing considerations. (footnotes omitted) (citations omitted)
412 Of course I accept that it is first appropriate to properly construe what in substance the process claim is to and, if necessary, consider the “other factors” identified by the plurality.
413 Further, Gageler and Nettle JJ also addressed whether claims to methods of using isolated nucleic acids may be patentable subject matter. Now at [134], their Honours observed that the essence of claim 1 was the “correlation between the incidence of cancer and the presence of the specified mutations and polymorphisms in the BRCA1 gene”, but that was not enough to make the claim a manner of manufacture. MLA says that it should follow that a claim to a correlation between cancer and the mutations or polymorphisms in the gene and to the use of that correlation in a known method would also lack patentability. Now I disagree and will dispose of this contention later. But such a contention also ignores other aspects of their Honours’ reasoning at [137] where they distinguished a product claim from a process claim:
Of course, as NRDC implies, the application of naturally occurring phenomena to a particular use may be a manner of manufacture if it amounts to a new process or method of bringing about an artificially created state of affairs of economic significance. Even so, the inventor cannot claim to have invented the naturally occurring product as opposed to the process of application. In NRDC, the patentee could not claim to have invented, and therefore there was no suggestion of it laying claim to a monopoly over, the commonplace herbicides which were used in the course of the patentable process. Similarly, in Shell Oil Co v Commissioner of Patents, the patentee could not claim to have invented, and therefore there was no suggestion of laying claim to a monopoly over, the known compounds which were applied as part of the patentable process to a new use of plant growth regulation. So too here, in so far as the invention consists in the application of a naturally occurring phenomenon to a particular use, the inventor cannot claim to have invented the naturally occurring phenomenon as opposed to the method of use and has no claim to a monopoly over the naturally occurring phenomenon as opposed to the method of use. (footnotes omitted)
414 Their Honours observed that “the application of naturally occurring phenomena to a particular use may be a manner of manufacture if it amounts to a new process or method bringing about an artificially created state of affairs of economic significance” (emphasis added). Now I accept that the word “may” is not unimportant. The formalism of the claim cannot overcome its substance. So, if in substance all that is claimed is the identification of a natural phenomenon or correlation, it will not be a manner of manufacture. MLA contends that if all the claim employs is established technology to isolate a fragment of naturally occurring DNA comprising the naturally occurring polymorphism and determination of a naturally occurring correlation between that polymorphism and an observed naturally occurring trait by way of standard statistical methods, the use of that method does not employ any method invented. I will come to this later.
415 Now Gageler and Nettle JJ said at [147]:
It was not disputed that [the patentee] might justly lay claim to the discovery that, if an isolated fragment comprising the BRCA1 gene is found upon examination to exhibit the specified mutations and polymorphisms, their presence is or may be indicative of particular kinds of malignancy in the cell. Nor was it disputed that a process or method of using known technology to isolate a sequence of nucleic acid comprising the BRCA1 gene and examining it for the presence of the specified mutations and polymorphisms for the purpose of detecting or predicting malignancy might be patentable …
416 Of course their Honours did not accept that such process claims would necessarily be patentable. Indeed, their Honours went on to say at [147] “[b]ut, as has been observed, the discovery of a natural correlation is not patentable as such ...” and its discovery did not mean the product used in that process was patentable. MLA contends that their Honours can be properly understood to be saying that the fact that process claims using the isolated nucleic acid of claims 1 to 3 were not in issue, did not mean the process claims themselves were patentable.
417 Further, I should also note that their Honours said at [152]:
In the same way here, it is one thing to say that the first respondent has invented a process which consists in isolating and examining the fragment comprising the BRCA1 gene for the presence of the specified mutations and polymorphisms as an indicium of malignancy. It is quite another and different thing to say that the first respondent, as inventor of that process, is entitled to a monopoly over the mutated BRCA1 gene, which is used merely as an ingredient in that process. The invention claimed makes no contribution to the manufacture of the substance. At best, it takes advantage of properties in the substance hitherto unknown or unsuspected. Just as there was no difference between the process and the product in Wellcome Foundation, there is no distinction between a claim to the process of isolating the BRCA1 gene for the purpose of examining it for the presence of the specified mutations and polymorphisms and the claim to the BRCA1 gene itself.
418 Further, their Honours said at [168]:
It is not disputed that a process or method of detecting the increased likelihood of certain kinds of malignancy by isolating the BRCA1 gene and examining it for the presence of any of the specified mutations and polymorphisms may be patentable subject matter as a process (subject to considerations of novelty and inventive step when compared to the prior art base). But, to repeat, claim 1 is not a claim for any such process. It is a claim for a monopoly over such isolated fragments of naturally occurring DNA as comprise the BRCA1 gene as are found upon examination to contain the (naturally occurring) specified mutations and polymorphisms. (emphasis added) (footnote omitted)
419 Again their Honours used the qualifier “may be”, which I will return to later. Now MLA has contended that their Honours’ requirement that a claim be properly construed to identify its substance and their observations that natural correlations are not patentable, answer any suggestion that their Honours would have accepted the method claims as patentable if they had been in issue. I consider that contention to be a rose-tinted perspective of what their Honours said.
420 Finally on this aspect of the analysis, Gordon J (at [258]) observed that claim 4, being to a probe containing a fragment of the isolated nucleic acid which was usually constructed artificially and had a radioactive label attached and could be used to identify mutations that might suggest a predisposition to cancer, was an invention. But as her Honour observed at [191] and [256], claims 4 to 30 of the patent in suit were not in dispute. MLA has contended that her Honour’s obiter conclusion must be confined to the detailed description her Honour gave of the “invention”, and in any event had to be contrasted with requirements elucidated by other members of the Court that any such claim be considered as a matter of substance to determine whether it was merely a claim to a natural correlation. I am not convinced that the implications from her Honour’s observations should be confined so narrowly and will return to this question later as to what should be drawn from Myriad concerning process or method claims.
Express exclusions in section 18
421 Before dealing with my analysis of the parties’ arguments before me, I should say something about s 18 of the Act.
422 Section 18(2) excludes as patentable inventions for standard patents “Human beings, and the biological processes for their generation”. Now an exclusion of that kind does not mean that everything outside that exclusion is patentable subject matter or, in particular, a manner of manufacture. This is plain from the finding in Myriad itself where the exclusion in s 18(2) was not the basis upon which Myriad was decided (at [11]). Of course, s 18(2) has no direct application to the 253 Application before me in any event as it only concerns aspects of the cattle genome. But I do not consider that I can draw any necessary implication in favour of Branhaven to the effect that the claims are patentable subject matter.
423 Let me also dispose of s 18(3) which provides that for the purposes of innovation patents, “plants and animals, and the biological processes for the generation of plants and animals, are not patentable inventions”. Now that provision does not apply to standard patents. But I accept that s 18(3) does not provide any necessary implication such as to conclude that a standard patent for those things is patentable. In any event, only the claims relating to the cloning of cattle (claims 7 and 11) would fall within this exclusion if they were in an innovation patent. For completeness, I should also mention s 18(4) that goes on to provide that s 18(3) does not apply if the invention “is a microbiological process or a product of such a process”. That provision limits the s 18(3) exclusion, but I accept that it does not necessarily entail that any such invention is a manner of manufacture.
(b) The present case
424 In essence, MLA has made two objections. The first objection relates to patentable subject matter, namely, whether the invention is a manner of manufacture in the sense of a proper subject of letters patent within developed principles. The second objection relates to the so-called “threshold requirement” of inventiveness on the face of the specification.
425 In my view neither type of objection is made good in relation to the claims of the 253 Application, save the first type of objection concerning claim 13. The rest of this section will address the first objection. In my view it has not been made out. But before addressing the claims in detail, let me set out my principal reasons for distinguishing Myriad.
426 First, the claims of the 253 Application are not directed to “naturally occurring genetic information per se”, as isolated nucleic acid sequences or otherwise, save claim 13 that is in a different category to the other claims in this respect. But even then, claim 13 is not entirely on all fours with the claims that were held not to be patentable in Myriad. Claim 13 has the rider “an isolated polynucleotide identified according to the method of claim 8” (my emphasis). The significance of that rider is something I will return to later.
427 Second, it is apparent from each of the three sets of reasons in Myriad that although claims 1 to 3 were held not to be patentable on the basis indicated, no member of the Court gave any significant indication that claims 4 to 30 were not or did not give rise to a “manner of manufacture”. The plurality emphasised that the Court was not concerned with gene patenting generally, but only with three claims encompassing isolated nucleic acids coding for a mutant or polymorphic BRCA1 polypeptide. Their Honours found that although the claims were product claims, they were in substance claims to the genetic information embodied in, and conveyed by, the nucleic acid sequence. And as the genetic information was naturally-occurring and had not been “made” by human action, it followed that claims 1 to 3 did not define a manner of manufacture. But it is apparent that their Honours’ reasoning did not rule out patentability to the products and methods of claims 4 to 30.
428 Third, it is also apparent that the plurality considered that if claims 1 to 3 were held to be a manner of manufacture, this would have involved an extension of the categories of patentable subject matter to a new class of case. It was in that context that their Honours discussed the need to consider the potential chilling effect and “other factors” before recognising such an extension. But the “other factors” do not arise unless the claims in question require an extension of the existing concept of manner of manufacture to a new class of case or are on the border. But that is not the present case before me, save for claim 13. The claims of the 253 Application (other than claim 13), directed to novel and inventive methods and processes as I have ultimately found, are within the plain vanilla concept of manner of manufacture as outlined in NRDC and Myriad.
429 Fourth, as I have already indicated, Gageler and Nettle JJ noted the recognition in NRDC that the application of a naturally occurring phenomenon to a particular use may be a manner of manufacture if it amounts to a new process or method of bringing about an artificially created state of affairs of economic significance. In essence, their Honours acknowledged that a process of using known technology to isolate a sequence of nucleic acid comprising the BRCA1 gene and examining it for the presence of the specified mutations and polymorphisms to detect or predict malignancy, might be patentable (at [147]), a point that they later reiterated (at [168]). Further the words “subject to considerations of novelty and inventive step when compared to the prior art base” in [168] are not to be overlooked (as MLA conveniently did so). They elucidate the qualification intended by the use of the word “may” in the preceding line. Their Honours implicitly recognised that a method involving the use of naturally occurring nucleic acid sequences for a particular purpose, such as a method of detection or diagnosis, may be within the established concept of a manner of manufacture and may be patentable if it satisfies the other requirements for patentability in the Act, including novelty and inventive step.
430 Fifth, Gordon J recognised the significant difference between claims 1 to 3 of the patent in suit and the remaining claims 4 to 30. Her Honour outlined the subject matter of the latter claims at [191] and later reasoned (at [256] to [258]):
What then did Myriad do? It took the idea, concept or principle that specific mutations or polymorphisms in that sequence suggest a predisposition to breast cancer and ovarian cancer and moved to carry out that idea, concept or principle, or embody it in a manner of new manufacture, in claims 4-30. The validity of those claims is not in issue.
Claim 4 may be taken as an example. In simple terms, it comprises a nucleic acid probe in which the nucleotide sequence is a portion of an isolated nucleic acid with the characteristic identified in claim 1 …
The invention in claim 4 carried into effect the idea that specifically identified mutations or polymorphisms in a sequence of the BRCA1 gene suggest a predisposition to breast cancer and ovarian cancer by testing for the presence of one or more of the specifically identified mutations or polymorphisms. That is an invention.
431 On her Honour’s reasoning, it would also appear that the invention in claims 5 to 30 was no different in principle. These included preparations and uses of polypeptides (claims 10 to 16) and various methods of diagnosis (claims 17 to 30). Now for completeness, I would note that claim 17 of the Myriad patent, which was tendered before me, was expressed in the following terms:
17. A method for diagnosing a predisposition for breast and ovarian cancer in a human subject which comprises determining whether there is a germline alteration in the sequence of the BRCA 1 gene in a tissue sample of said subject compared to the nucleotide sequence set forth in SEQ.ID No:1 or a wild-tyle [sic] allelic variant thereof, said alteration indicating a predisposition to said cancer being selected from the mutations as set forth in Tables 12, 12A and 14.
432 Such a claim has some analogy with claim 1 of the 253 Application. And as already indicated, claim 17 of the Myriad patent was not considered not to be or not to involve a “manner of manufacture”. But I accept of course that these other claims (claims 4 to 30) were not in issue in Myriad.
433 In summary and generally speaking, the reasoning in Myriad does not assist MLA, save for claim 13.
The claims
434 Let me now discuss the specific claims of the 253 Application. But before addressing the “other factors” analysis of the plurality in Myriad, let me address each claim in turn in dealing with MLA’s contention that each of the claims does not give rise to an artificially created state of affairs, being a necessary requirement for patentable subject matter. It is also in this context that I would prefer to deal with the suggestion that seems to have been made by MLA that Gageler and Nettle JJ have introduced into the concept of manner of manufacture a separate threshold requirement for inventiveness. Certainly, many of MLA’s submissions relying upon Gageler and Nettle JJ’s reasoning seem to carry that implicit assumption.
Claim 1
435 MLA contends that despite the definition of the claimed invention (claim 1) as a process of identifying a trait in a bovine subject from a nucleic acid sample of the bovine subject, its substance is information embodied in the natural correlation that exists between the nucleotide sequences of the nucleic acid sample (in particular, at least three SNPs in those sequences) and a bovine trait.
436 It is said that that is analogous to the information embodied in arrangements of nucleotides in Myriad (at [6]) and claim 13 of the 253 Application. It is also analogous to the discovery of the natural correlation between the polymorphisms in the BRCA1 gene and the propensity for developing cancer that Gageler and Nettle JJ observed was not patentable in Myriad at [134] and [147].
437 MLA says that the identification of a trait of a bovine subject from the information gleaned from a nucleic acid sample of the bovine subject is not information that is “made” by human action – it is “discerned”. Even if claim 1 is understood as pertaining to identifying the propensity of the bovine animal for the trait, that propensity is not changed by the human who identifies that propensity through discovery of the existence of SNPs in a nucleic acid sample from the bovine subject: Myriad at [6]. It certainly is not sufficiently artificial or different from nature as to be regarded as patentable (to paraphrase Gageler and Nettle JJ at [127]).
438 MLA says that in substance, there is no invention claimed, but merely the discovery and use of naturally occurring polymorphisms (SNPs) in nucleotide sequences that are associated with naturally occurring traits.
439 MLA says that claim 1 does not expressly or in substance result in an “artificially created state of affairs” because no product is made and there is no process producing an artificial outcome as a result of human action. It says that an “artificially created state of affairs” requires:
(a) “physical phenomenon in which the effect, be it creation or merely alteration, may be observed”: NRDC at 276;
(b) “[a] physical effect in the sense of a concrete effect or phenomenon or manifestation or transformation ...”: Grant v Commissioner of Patents (2006) 154 FCR 62 at [32] per Heerey, Kiefel and Bennett JJ; and
(c) “something of a corporeal and substantial nature”: Lockwood Security Products Pty Ltd v Doric Products Pty Ltd (No 2) (2007) 235 CLR 173 (Lockwood (No 2)) at [66].
440 MLA says that the arbitrary limitation in claim 1 to at least three SNPs, where two of the SNPs occur in different genes, cannot avoid a finding that the claims are not to patentable subject matter. The limitation does not change the fact that in working the method of at least claims 1 to 6, 9 and 10 there is no concrete, tangible, physical, or observable effect produced (Grant v Commissioner of Patents at [30] to [32]). Rather, the claimed method is directed to inferring the likelihood of a bovine subject having a particular trait. This is merely an intellectual exercise, based on the natural relationship between the bovine’s genotype (the presence or absence of particular SNPs) and its phenotype (the presence of a trait), and manifests itself only in the form of added knowledge about the inherent nature of the animal.
441 Now before me Branhaven said that the artificially created state of affairs arising from the claimed method is the improvement, over time, of the genetic potential of cattle for a desirable trait. But MLA says that this is not an aspect of the alleged invention. The invention claimed in claim 1 ends at the point of identifying a trait. In fact, none of the claims give rise to an improved animal. Claims 2 to 5, 9 and 10 are all dependent on 1 and do not add any limitation of improvement. Claim 6 claims sorting cattle but that does not change the cattle. Claim 7 is to a cloned animal but that merely produces the same animal as was tested for the trait – not an improved animal. Claim 11 is dependent on claim 7 and does not result in an improved animal. Claim 8 is in substance the same as claim 1. Claim 12 is in essence the process of claim 1 for a population of bovine. Claims 13 and 14 do not involve improvement of an animal. Claim 15 is dependent on claims 1 or 6 or 8 and so does not add anything further. MLA says that consistently with the plurality’s observation in Myriad at [85], whatever may occur subsequent to these points is separate and remote from the claimed invention and is not relevant to whether the claimed invention is a manner of manufacture.
442 Further, MLA says that Branhaven’s analogy with the improvement of sown land in NRDC does not withstand scrutiny. In NRDC the claims were, relevantly, to the application of the herbicide to crop areas, and the artificially created state of affairs was discernible by observing over a period the growth of weeds and crops respectively on sown land on which the method had been put into practice. That is, there was a discernible difference between the crop area in terms of the comparative growth of weeds and crops on sown land. But in the present case, MLA says that nothing in the bovine subject or the nucleic acid sample of the bovine subject is changed. There is no “better cow”. The claim is to identification of a trait in a bovine subject, but there is no physical effect arising from the identification per se of a trait in that bovine subject. The person applying the method of claim 1 merely discovers the presence of a trait in the bovine subject based upon some natural correlation between at least three SNPs in a nucleic acid sample of the bovine subject and the trait of that bovine subject.
443 Further, MLA says that the specification acknowledges that the invention allows “discovery” of SNP markers that associate with traits throughout the genome. But it says that association of genetic markers and genes with traits has long been a well-known natural phenomenon. It says that the SNPs and their association with traits is not something that is “made” or “artificially created”. All that the 253 Application does is to claim the use of these discoveries or principles of nature to predict genetic potential.
444 MLA says that in NRDC, after acknowledging the difficulty in delineating discovery from invention, it was said (at 264):
There may indeed be a discovery without invention – either because the discovery is of some piece of abstract information without any suggestion of practical application of it to a useful end, or because its application lies outside the realm of “manufacture”.
445 MLA says that to the extent that the method of “identifying” in claim 1 constitutes the application of Branhaven’s discovery, that application lies outside the realm of manufacture. There is no “product” or no “artificially created state of affairs” or not one which is sufficiently artificial or man-made. It is said that none of the claims result in any alteration to a naturally existing animal, and there are no claims to an animal that is altered in any way via this process.
446 MLA says that in the words of Gageler and Nettle JJ, the subject matter of claim 1 does not have “a quality of inventiveness which distinguishes it from a mere discovery or observation of a law of nature” (at [131]). That is, it is not “sufficiently artificial” and the “inventive concept” identifying a trait (if there is such a thing) does not make a contribution to the essential difference between product and nature (at [127] and [128]). I will discuss the question of whether Gageler and Nettle JJ have added an additional requirement of inventiveness later. I would observe though at this point that if they have, it was not endorsed by any of the other judges in Myriad.
447 Further, MLA says that sufficiency of artificiality is a concept that has also been used when considering whether a business method that uses a computer is a “manner of manufacture”. As Commissioner of Patents v RPL Central Pty Ltd (2015) 238 FCR 27 observed at [96] (per Kenny, Bennett and Nicholas JJ):
A claimed invention must be examined to ascertain whether it is in substance a scheme or plan or whether it can broadly be described as an improvement in computer technology. The basis for the analysis starts with the fact that a business method, or mere scheme, is not, per se, patentable. The fact that it is a scheme or business method does not exclude it from properly being the subject of letters patent, but it must be more than that. There must be more than an abstract idea; it must involve the creation of an artificial state of affairs where the computer is integral to the invention, rather than a mere tool in which the invention is performed. Where the claimed invention is to a computerised business method, the invention must lie in that computerisation. It is not a patentable invention simply to “put” a business method “into” a computer to implement the business method using the computer for its well- known and understood functions.
448 MLA says that as a result of Myriad, naturally occurring genetic information is not patentable per se, even when it is isolated from its natural state. It says that in the present case the alleged invention is in identifying an association between certain traits and the specified SNPs that are naturally occurring genetic information. But the addition of identifying the trait that correlates to the naturally occurring genetic information does not involve changing any commonly used tool. And the same is true for the remaining claims of sorting, cloning, hybridising etc. It says that the invention does not lie in those acts. It says that it is not an invention to simply put the non-patentable discovery of information into methods for their well-known and understood purposes.
449 Further, MLA says that to the extent that the invention claimed in claim 1 is considered in substance to be a method, that method is not new and it has not been invented by the inventors named in the 253 Application. It says that the use of DNA markers, including SNPs, associated with traits to predict genetic potential has been well-known since before December 2002. I would note at this point that this high and wide submission travels well beyond the two types of objection, in the context of “manner of manufacture”, that I am considering, however much MLA would seek to finesse from its optimistic reading of the reasoning of Gageler and Nettle JJ on the question of “inventiveness”.
450 Further, MLA says that considering claim 1 on its terms, the claimed method is “[a] method for identifying a trait of a bovine subject from a nucleic acid sample of the bovine subject, comprising identifying in the nucleic acid sample an occurrence of at least three [SNPs] …”. But MLA says that the method of identifying traits is nothing more than identifying the occurrence of certain SNPs. It is said that this is apparent because identifying the occurrence of certain SNPs is all that the claimed method “compris[es]”. But MLA says that the patent applicants have not invented a method of identifying the occurrence of SNPs, and claim 1 covers any and all methods of identifying the occurrence of SNPs, whether those methods are presently known or unknown.
451 Further, MLA says that the fact that the claimed method may be characterised by what is identified (as opposed to how it is identified) does not assist Branhaven. It says that to the contrary, the fact that the apparently defining characteristic of the claimed method is what is identified, discloses the true substance of the claimed invention as merely the discovery and use of naturally occurring polymorphisms in gene sequences (SNPs) that are naturally associated with naturally occurring traits.
452 Generally speaking, I would reject MLA’s arguments.
453 First, the claims in suit are not directed purely to genetic information. Rather, the claims of the 253 Application are directed to methods and other embodiments involving the practical application of the identification of SNPs from a nucleic acid sample of the bovine subject and their association with a trait of interest. So, the claims are directed to artificial subject matter being the result of human action, rather than something that exists in nature per se.
454 Second, it is inappropriate to fasten upon individual elements of the claims, such as SNPs and their association with the trait. Of course these are naturally occurring phenomena. SNPs are naturally occurring. And the associations are objectively ascertained correlations between naturally occurring phenomena, that is, the genotype (used in the narrow sense) and the phenotype (used in the narrow sense). But the claims are not merely directed to the SNPs or their association with the trait per se. It is impermissible to disregard the wording of the claims and diminish their formal content under the guise of having regard to the “substance” of what is claimed. There is no suggestion in Myriad that claims to methods involving the practical application of nucleic acids could be dismissed as being in substance just directed to genetic information.
455 Third, claim 1 of the 253 Application on its face involves the practical application of a naturally occurring phenomenon to a particular use. It is not just the mere identification or discernment of the naturally occurring phenomenon, contrary to MLA’s assertions. The claimed method involves human interaction and the creation of an artificially created state of affairs. It requires the taking of a nucleic acid sample of the bovine subject through an appropriate procedure. It requires the identification in that sample of at least three SNPs that meet the requirements of the claim, which involves analysing the sample to identify SNPs present that are associated with the trait. It involves the creation of an artificially created state of affairs of economic significance through human interaction. Now I accept that it involves drawing an inference about the potential for the trait of interest to exist in the bovine subject, which involves discerning the natural state of affairs, but the claim is more than this. All of this is a practical application that results in an artificial state of affairs, and with undoubted economic significance. And even accepting that there are many possible applications of the method does not undermine its significance or patentability.
456 At this point it is now convenient to deal with several aspects of the reasons of Gageler and Nettle JJ that MLA has sought to draw comfort from, although I could have also dealt with it under the second type of objection that I will come to later.
457 First, MLA suggests (as I understood its argument) that Gageler and Nettle JJ have added another threshold requirement for inventiveness. By that I mean that it was suggested that in addition to the s 18(1)(b)(ii) requirement of inventive step there was a separate threshold requirement for inventiveness in the form of a back-door introduction of the so called “product of nature doctrine” favoured in the US. So MLA pointed to the following passages of their Honours’ reasoning:
(a) “The question then is whether the subject matter of the claim is sufficiently artificial, or in other words different from nature, to be regarded as patentable” (at [127]);
(b) “it is necessary that the inventive concept be seen to make a contribution to the essential difference between the product and nature” (at [128]);
(c) “the subject matter of a claim have about it a quality of inventiveness which distinguishes it from a mere discovery or observation of a law of nature” (at [131]);
(d) “there are limits on the patentability of products of nature inasmuch as products of nature do not involve human intervention and therefore are lacking in the necessary quality of inventiveness to qualify as a manner of manufacture” (at [136]); and
(e) a favourable endorsement (see at [136]) of Professor Sherman’s observation that the US product of nature doctrine and the Australian test of artificially created state of affairs are the same question asked from different perspectives.
458 Now I am not at all convinced that their Honours have introduced a new threshold requirement for inventiveness. There is always a difficulty in parsing and nuancing the language of a judgment. But let me assume for the moment in favour of MLA that this is how their Honours’ reasoning should be read. I have not applied such a new threshold for inventiveness because there are two binding authorities against me doing so. The first authority is Myriad itself. Neither the plurality nor Gordon J endorsed such an approach. The second authority is Lockwood (No 2) at [106] where the Court stated, with reference to Commissioner of Patents v Microcell (1958) 102 CLR 232 that Microcell:
stands for a narrow proposition that a Commissioner of Patents, or his or her delegate, may refuse an application for patent protection where a specification “on its face” shows the invention claimed is not a manner of new manufacture. This may arise, for example, from admissions concerning novelty. The decision in Microcell has not always been properly understood; it does not involve a separate ground of invalidity or a discrete “threshold” test. (footnote omitted) (my emphasis)
459 In other words, Lockwood (No 2) rejected any separate threshold test, whatever it might be said that NV Philips Gloeilampenfabrieken v Mirabella International Pty Ltd (1995) 183 CLR 655 at 663 and 664 stands for. Moreover, the plurality in Myriad at [12] did not refer to Philips to support any separate threshold requirement for inventiveness, but rather cited Philips in a manner consistent with how Lockwood (No 2) had described how Microcell should be read; this is consistent with the conjunction of the two references in footnote 26 to the plurality’s reasons in Myriad.
460 To complete the circle, Gageler and Nettle JJ applied Lockwood (No 2) (see [131] and footnote 144), thereby confirming my doubts as to whether their Honours intended to introduce a new threshold requirement.
461 Do I need to precisely resolve whether there has been such an introduction? No. As I have said, I am bound to apply the plurality’s approach in Myriad and also Lockwood (No 2); I will return to Lockwood (No 2) later when I discuss the second type of objection.
462 Second, MLA has persistently made much of the observations of Gageler and Nettle JJ that the discovery of a correlation between naturally occurring phenomena is not a manner of new manufacture. I am not sure where any of this takes MLA. It may be accepted that an idea is not patentable per se. It may be accepted that a “natural law” or “law of nature”, whatever that means (the concept is a contestable proposition for a philosopher of science), is not patentable per se. It may be accepted that the discovery of an objectively ascertained statistically significant correlation between two physical phenomena is not patentable per se. But so what? In the present case, what is sought to be patented is a method. The questions are whether that method is a manner of manufacture, is novel, and involves an inventive step. Moreover, no authority requires me to exclude from the inventive step analysis, the ingenuity and very extensive research that was undertaken by the patent applicants to identify the correlation between genotype and phenotype as part of determining whether the method (not just the discovery of a correlation) involved an inventive step.
Claims 2 to 5
463 Claims 2 to 5 are dependent on claim 1 and none introduce a limitation that warrants a different view as to patentability from claim 1.
Claim 6
464 Claim 6 is in substance the application of claim 1 to a method of sorting “one or more bovine subjects”.
465 MLA contends that sorting one bovine subject simply involves no more than claim 1, namely, identifying a trait in that bovine subject. Similarly, as to multiple bovine subjects, MLA says that no change in the bovine subjects is caused by the application of sorting them according to a trait. Furthermore, MLA says that the added limitation of sorting the cows by the identified trait does not involve any invention by the inventors. It simply involves the use of known techniques to identify the trait (by DNA sampling and genotyping and phenotyping, which are not described in the specification as new or inventive) and sorting, which is exemplified in the 253 Application as sorting to a pen but captures simply sorting animals in one’s mind and thus has very similar characteristics to a business method. Further, MLA says that as a matter of substance, the sorting integer does not change the fact that the claim is to the mere discovery of a correlation between the genetics of bovine subjects and their traits. Again it prays in aid the words of Gageler and Nettle JJ that the subject matter of claim 6 does not have “a quality of inventiveness which distinguishes it from a mere discovery or observation of a law of nature” (at [131]). That is, it is not “sufficiently artificial” and the “inventive concept” of sorting does not make a contribution to the essential difference between product and nature (at [127] and [128]).
466 In my view, for reasons already stated, claim 6 has not been shown not to satisfy the criterion of a manner of manufacture. It is dependent on claim 1 which has not been shown not to satisfy that criterion.
467 Further, I agree with Branhaven that if anything more was required for patentability over and above what is expressly set out in claim 1, it is present here. Claim 6 in terms involves the sorting of bovine subjects based on the drawing of inferences as to the existence of the trait of interest. Now of course it involves using information or discovery. But it is a practical application involving human intervention and resulting in an artificially created state of affairs of economic significance, viz, bovine subjects sorted according to the potential for the trait to exist.
Claims 7 and 11
468 Claim 7 is to a method for cloning a bovine subject with a desired trait which involves the application of identifying an occurrence of at least three SNPs associated with the trait in essentially the same way as claim 1. Claim 11 is to a bovine resulting from the cloning method in claim 7.
469 Now I accept that a cloned animal will have the same genetic code as the animal it is cloned from. Accordingly, in one sense nothing artificial is created, just as the isolated nucleic acid was not a manner of manufacture in Myriad. And I also accept that the patent applicants have not invented any method for cloning bovine subjects and do not assert as such. And I also accept that the claims in part use known techniques of identifying SNPs in a bovine subject and cloning an animal from a progenitor cell of the bovine subject.
470 But claim 7 is directed to a method for cloning a bovine subject. Further, some of the steps are analogous to the method of claim 1. Accordingly, the claim is directed to a practical application, which undoubtedly involves human intervention, and which results in an artificially created state of affairs of economic significance. The practical application is the cloning of a bovine subject, with the cloned subject the subject of claim 11. Now of course the cloned cow in one sense is the same as that which it clones. MLA says superficially that it is “mere genetic information on a grander scale” and accordingly Myriad is directly applicable. The submission has a superficial allure, but I reject it. An artificial object of economic significance is produced for its own sake, not merely as a receptacle for its informational content.
471 In my view, claims 7 and 11 are not shown not to satisfy the criterion of being a manner of manufacture. For completeness, I would note that Branhaven has invited me to draw various inferences in support of its position from ss 18(2) and 18(3) of the Act. I have already expressed my views on that aspect.
Claims 8, 9 and 10
472 Claim 8 is in essence the method of claim 1, although using the word “inferring” not “identifying”, comprising hybridising the nucleic acid sample from the bovine subject to a system wherein the selective hybridisation indicates the presence of a SNP associated with the trait. MLA says that in effect, claim 8 is a known technique to achieve claim 1.
473 In my view claim 8 includes additional features relating to the technical process of hybridising the nucleic acid sample that confirm it is patentable subject matter and is also not shown to be not a manner of manufacture for the reasons given concerning claim 1.
474 Claims 9 and 10 limit the preceding claims to the identification of particular traits, and need not be considered separately for present purposes. They also are not shown to be not a manner of manufacture.
Claim 12
475 Claim 12 is to a system for determining nucleotide occurrences of SNPs in a bovine population. It has a similarity with claim 8. MLA says that it is one level of abstraction away from claims 1 and 8 in that, in substance, the claim concerns the identification of naturally occurring correlations between SNPs and a trait.
476 In my view claim 12 is directed to a system for determining nucleotide occurrences of SNPs in a bovine population. And the claimed system comprises a hybridisation substrate that includes specific binding pair members corresponding to a series of at least three SNPs, and with features similar to claim 1. In my view claim 12 is directed to a practical application involving human intervention, and resulting in an artificially created state of affairs of economic significance. Claim 12 has not been shown not to satisfy the manner of manufacture criterion.
Claim 13
477 The delegate found that claim 13 of the 253 Application was not to a manner of manufacture in light of the decision in Myriad. The delegate said (at [58] and [134] to [136]) the following:
58. Claim 13, as drafted, does not depend in a meaningful fashion from the method of claim 8 or any other claim. This claim is unclear. However, I consider the intention of claim 13 is to define isolated and specific polynucleotides per se which harbour at least one SNP in a gene where the SNP is associated with a trait, and the SNP corresponds to position 300 of any one of SEQ ID NOS: 19473 to 21982, or the SNP is about 500,000 or less nucleotides from position 300 of any one of SEQ ID NOS: 19474 to 21982.
…
134. In response to my request, on 27 October 2015, both parties filed further written submissions regarding the relevance of D’Arcy v Myriad Genetics Inc [2015] HCA 35, to the patentability of claim 13.
135. In D’Arcy v Myriad Genetics Inc [2015] HCA 35 at [89], the majority decided that if information in an isolated nucleic acid is the same information as that contained in the DNA of the subject from which the nucleic acid was isolated, then the isolated polynucleotide is not patent eligible.
136. I have already decided claim 13 does not depend in any meaningful way from claim 8, or any other of the accepted claims. However, notwithstanding this ambiguity, if the applicant’s intention was to claim isolated polynucleotides harbouring at least one of the SNPs corresponding to position 300 of any one of SEQ ID NOS: 19473 to 21982, or a SNP within 500kb of the index SNP, the claimed isolated bovine polynucleotides amount to genetic information which has not been made. Consequently, the isolated polynucleotides would not be a manner of manufacture.
478 Now Branhaven did not appeal this finding. But it contended that because this is a de novo hearing, it could now contest that finding. Now I accept that MLA’s Notice of Appeal specifically excludes the findings of the delegate as to claim 13. Further, r 34.28 of the Federal Court Rules 2011 (Cth) provides that if a respondent wants to appeal from a decision, or part of a decision, it must file and serve a notice of cross-appeal, which Branhaven did not do. Nevertheless, it is appropriate to further consider claim 13 since the finding that claim 13 is not a manner of manufacture may be a relevant consideration as to whether it would be anomalous for method claims (i.e. claim 1) involving such nucleic acid sequences to be a manner of manufacture. Accordingly, to the extent necessary I will dispense with the requirements of r 34.28 and treat Branhaven as if it had cross-appealed.
479 Now claim 13 claims an “isolated polynucleotide identified according to the method of claim 8”. But claim 8 does not in its terms claim a method of identifying an isolated polynucleotide. Claim 8 claims a method of inferring a trait in a bovine, comprising hybridising a nucleic acid sample from a bovine subject to a system wherein the hybridising comprises certain elements. Accordingly, claim 13 was held by the delegate to lack clarity (at [58]), a conclusion with which I agree.
480 Now the delegate assumed for the purposes of her consideration of manner of manufacture that the isolated polynucleotide of claim 13 was one harbouring a specified SNP or non- specified SNP. She found that in light of Myriad that could not be patentable subject matter.
481 Now Branhaven has contended that claim 13 is not comparable with the claims that were held not to be patentable in Myriad. It points out that claim 13 is not to an isolated nucleic acid per se, but rather is to an isolated polynucleotide identified according to the method of claim 8. Branhaven says that it is only when an isolated polynucleotide is identified through that method that it will be covered by the claim.
482 I reject Branhaven’s arguments. To accept them would be a triumph of form over substance and would be inconsistent with the themes of Myriad. Applying the themes of Myriad as liberally and as faithfully as I can, and without the need to justify this position further, in my view claim 13 is not a manner of manufacture. I also do not perceive any inconsistency between that conclusion and the conclusion that I have reached concerning, inter-alia, claims 1 and 8, namely, that I am not satisfied that they do not satisfy the manner of manufacture criterion.
Claim 14
483 Claim 14 is to an “isolated polynucleotide” when used in any one of the methods of claims 1 to 10. Now I accept that the isolated polynucleotide in and of itself is not a manner of manufacture. But the important distinguishing feature is that claim 14 is directed to an isolated polynucleotide when used in any one of the methods of the earlier claims. Its scope is more coordinate with that of the method claims. The invention is the product in its use. Accordingly, in my view it is a manner of manufacture. A new and inventive use of an unpatentable product can clearly be a “manner of manufacture”.
Claim 15
484 There is nothing further to add. It is not established as not being a manner of manufacture.
All claims – Myriad “other factors”
485 MLA contends that even if the claims were to products or processes that are the result of human action, the claims would still not be to patentable subject matter. It says that the claims properly construed are not within the established boundaries of what constitutes patentable subject matter. Accordingly, wider considerations than the test in NRDC come into play.
486 Now I do not consider that MLA has established that the claims (apart from claim 13) are outside or at the margin of the established boundaries of what constitutes patentable subject matter. Accordingly, MLA’s argument concerning the application of the Myriad “other factors” fails at this threshold. But let me assume for the moment that I am wrong. It is convenient for me to address these “other factors”. And applying them, I do not consider that any lack of patentability is established save for claim 13.
487 First, MLA says that the desirability of coherency in the application of the law points against the claims of the 253 Application being patentable. It contends that at their heart, they involve the use of naturally occurring genetic information which Myriad has found not to be patentable. Therefore, so it is said, there would not be any incongruity or anomaly in holding the claims of the 253 Application non-patentable. Indeed, there would be congruity in doing so. I disagree. What is claimed are method claims. It is well accepted that method claims can use known products and apply “natural laws” to their working. A method claim cannot sensibly be characterised as a claim to information. It is a claim to inter-alia the application of information. In my view to find that the method claims in suit involve a “manner of manufacture” is to enhance rather than detract from the coherency of Australian law.
488 Second, MLA says that as a matter of coherency with foreign law, the claims should be held not to be directed to patentable subject matter having regard to decisions in the US which have rejected claims to methods of diagnosis based on discoveries or principles of nature; see Mayo Collaborative Services v Prometheus Laboratories, Inc. 566 US 66 (2012) and Ariosa Diagnostics Inc. v Sequenom, Inc. 788 F3d 1371 (3d Cir 2015). But I reject this submission. I will discuss Mayo in a moment, but I propose to say nothing further about Ariosa given that the trial of patent litigation between the Ariosa parties has been set down before me in August 2018 involving the analogous Australian patent to the patent in suit in the US proceedings.
489 Now there are a number of difficulties with MLA’s submission invoking foreign law coherency.
490 The first difficulty is that I cannot determine coherency with foreign law generally by only considering cherry-picked jurisprudence from one jurisdiction. Consistency with one foreign jurisdiction might produce inconsistency with another foreign jurisdiction. I have not had the benefit of any comprehensive international survey.
491 The second difficulty is that as a trial judge I am applying an evolving conception from the Statute of Monopolies in the context of Australian legislation and Australian conditions, not any foreign law approach. As Myriad has made abundantly clear, I have to apply a common law methodology to an ever widening conception. But I hasten to add that this does not involve me in any legislative activity save in the non-pejorative sense elegantly discussed by the eminent legal realist and jurist Benjamin Cardozo in his Storrs lecture series (Yale University, 1921), particularly lecture III.
492 The third difficulty is even more fundamental. The US approach accepts that a method involving the application of a “law of nature” may be patentable. Indeed, the contrary could not be seriously argued. What workable method in its application is ever free of a “law of nature”? The US debate turns more on the question of what it takes “to transform an unpatentable law of nature into a patent-eligible application of such a law” (Mayo at 72). And everyone seems to agree that you need to do more than simply state the law of nature while adding the words “apply it”. But then one is into quite unclear territory. The exposition of the test (particularly the second stage) in Mayo is too sweeping for me to work out whether I am acting consistently or inconsistently with its spirit in the conclusion that I have reached that the claims of the 253 Application (save for claim 13) are patentable.
493 Third, MLA says that the potential claims will have (or may indeed have had) a chilling effect on future research in the livestock industry in Australia contrary to the interests of the Australian public, and will be “generally inconvenient” contrary to s 6 of the Statute of Monopolies. MLA points to the observation in Myriad (at [93]) that the isolated DNA claims had:
… the odd consequence that if the claims are properly the subject of a patent, the patent could be infringed without the infringer being aware of that fact. That consequence coupled with the very large, indeed unquantified size of the relevant class of isolated nucleic acids, all of which bear the requisite information, raises the risk of a chilling effect upon legitimate innovative activity outside the formal boundaries of the monopoly and risks creating a penumbral de facto monopoly impeding the activities of legitimate improvers and inventors.
494 MLA says that the same applies to the claims of the 253 Application, which include claims to an almost infinite class of isolated DNA (all genomes of all cattle whether existing now or coming into existence in the future) and the use of such DNA in well-known methods. It is said that Australian cattle farmers might not know that they are already doing something that falls within the scope of the claims of the 253 Application. It is said that research into breeding methods is likely to be hindered. MLA says that the grant of such claims would lead to the creation of an exorbitant and unwarranted de facto monopoly on all methods of using isolated nucleic acids containing the sequences coding for naturally occurring SNPs which are associated with any trait in bovine. It is said that the infringement of such claims would not be ascertainable until not only the SNPs were identified, and identified as falling within more than one gene, but also until an association between the SNPs and a trait was identified. It is said that such an outcome would be at odds with the purposes of the patent system. Indeed, it is said that if one chooses to use a group of SNPs which are not one of the specified SNPs, while there is a high chance one might infringe (because of the breadth of the claims) one could not find out if this was the case without conducting significant research to determine whether any one of the SNPs used is within 500,000 nucleotides of a specified SNP (and in association with a trait).
495 MLA also says that the evidence of Professor Plastow and Professor Goddard supports the view that the uncertainty of breadth of the claims would if granted have a chilling effect on future research, particularly given the scope of limb (b) of claim 1 and also the scope of claim 1 concerning the other two SNPs which are not required to be selected from limb (a) or limb (b) of claim 1.
496 I reject MLA’s contention that the claims of the 253 Application will have a substantial chilling effect on future research, particularly as narrowed by any amendment to deal with concerns that I have raised concerning claim 1 dealing with “association” and the requirement for LD in relation to limb (b). There is no cogent evidence to support the contention. Professor Goddard’s assertions have little weight on this aspect in my opinion. Indeed, the same criticisms he levelled at the 253 Application could be said to apply equally to other patents. MLA’s assertion does not sit well with the following patent applications and their grant.
497 First, take Australian standard patent application No 2007335195 titled “Artificial selection patent method and reagents” that was granted on 27 April 2017, the inventors for which are Professor Goddard and Professor Hayes (witnesses called by MLA) and the applicant for which was Agriculture Victoria Services Pty Ltd (“the AV/Goddard patent application”). Claim 1 covers both plants and animals and is of very great width. Further, claim 1 refers to “informative markers”. But it is not clear what is meant. It seems to embrace (p 17) a universe of possibilities including, in relation to DNA polymorphisms, SNPs, indels, short tandem repeats, microsatellites, mini-satellites, restriction fragment length polymorphisms and amplified fragment length polymorphisms. Further, there is little if any teaching on how to get an informative marker. One can make the jury point that if MLA’s chilling effect point was good, then the AV/Goddard patent application would be an example par excellence.
498 Second, take Australian standard patent application No 2002229406 titled “DNA markers for meat tenderness” granted on 5 March 2007, the applicants for which are notably MLA, the State of New South Wales, the State of Queensland, the CSIRO and the University of New England. Claim 1 thereof provides:
1. A method for assessing the tenderness of meat from an animal, comprising the step of testing the animal for the presence or absence of a genetic marker selected from the group consisting of:
(1) an allele of the gene encoding calpastatin (CAST) associated with peak-force variation or genetic variation located other than in the CAST gene which shows allelic association with the CAST allele; and
(2) an allele of the gene encoding lysyl oxidase (LOX) associated with variation in instron compression of the semitendinosis muscle or genetic variation located other than in the LOX gene which shows allelic association with the LOX allele.
A number of features may be noted. First, the type of animal is not constrained. Second, the type of genetic marker is hardly constrained, as compared with claim 1 of the 253 Application in terms of the type of marker, the number, and with at least one SNP that must fit within limb (a) or limb (b) (and accepting for the moment that the limb (b) SNP should be in linkage disequilibrium with the limb (a) SNP). MLA’s “chilling effect” argument before me, although put earnestly by its counsel, hardly sits well with its own patent and its effect. I accept though that this is more a jury point.
499 Further, it should be borne in mind that the breadth of claims per se is not indicative of a lack of patentable subject matter. A complaint about the breadth of claims per se is something that arises under other grounds of invalidity, such as a lack of clarity or a failure to define the invention because the breadth entails that the boundaries of the monopoly are elusive. In CCOM Pty Ltd v Jiejing Pty Ltd (1994) 51 FCR 260 in the context of a challenge to the patentability of a computer-implemented invention it was said at 294 and 295 (per Spender, Gummow and Heerey JJ):
[The submission] was that there could be no manner of manufacture in identifying “basic characteristics” or “desiderata” and “to claim all ways of achieving [them]”. Applying that to the present case, it was submitted that all that had been done was to select a desirable characteristic of a computer program, the ability to search, in the manner described, a data base of the type described, and “to claim all computers present and future possessing that characteristic”.
That submission should not be accepted. It may be that such a claim lacks novelty, is obvious, or lacks utility, or there is a failure to comply with one or other of the limbs of s 40 because, for example, the invention is not fully described or the claim is not clear and succinct. But if such hurdles all are surmounted, then in our opinion in a case such as the present there does not remain an independent ground of objection as to patentability, within the sense of s 18(1)(a) …
500 In other words, assertions about the breadth of claims per se, as opposed to the nature of the subject matter to which claims are directed, are to be assessed by reference to other grounds, each of which are legally and conceptually distinct, rather than under the rubric of manner of manufacture, save that breadth is relevant to the “other factors” 3.1 and 3.2 (as the plurality indicated at [29]). Now I have considered the observations of Gordon J in Myriad at [259] to [264], but I have endeavoured to track as closely as possible the approach of the plurality ([14], [29] and [87] notwithstanding) although her Honour’s observations on breadth can also be seen as resonating with the context in which the plurality were referring to breadth.
501 In summary, in my view none of the “other factors”, to the extent that they have meaningful application in the context that I am considering, point against patentability. They all consistently point in one direction, which unsurprisingly is a conclusion consistent with my primary conclusion that the claims of the 253 Application (save for claim 13) are well within the established boundaries of “manner of manufacture”. It also follows that for the above reasons the “generally inconvenient” limb of s 6 of the Statute of Monopolies is not engaged.
(c) The “threshold requirement”
502 The other type of objection based on the “threshold requirement” of inventiveness is a very limited one. It only arises where it can be said that there is not invention (in the sense of a lack of inventive step or possibly lack of novelty) “on the face of the specification” alone, and without regard to evidence or extrinsic facts external to the specification. Further, as I have said earlier, authority binding upon me establishes that this threshold test does not permit of an additional and separate threshold test for inventive step in addition to s 18(1)(b)(ii); accordingly I do not propose to say anything further at this point on any additional threshold concept of inventiveness that may be said to arise from the analysis of Gageler and Nettle JJ in Myriad.
503 In considering this other type of objection, the analysis is limited to what appears on the face of the specification properly construed of course by the skilled addressee in the light of common general knowledge. This type of objection has sometimes been characterised as turning upon an admission in the specification that there is no patentable invention. That is how the plurality characterised the objection in Myriad (at 12]):
That anterior exclusion may be based upon an admission, on the face of the specification, which makes clear that the invention claimed is not novel or does not involve an inventive step.
504 In Lockwood (No 2), to which I have already referred, the Court said (at [106] and [107]):
In Chapman and Cook and Lectro Linx Ltd v Deltavis Ltd, Clauson J remarked:
[I]f a Patentee, though entirely erroneously, does state by way of what I may call recital in his Specification that a particular form of thing is common and then by some oversight or some mistake claims a monopoly in that particular form of thing he will have, so to speak, recited himself out of Court and I venture to doubt whether he could possibly maintain any claim to a monopoly in a thing which he has recognised to be something which existed.
Chapman may be understood as a case which exemplifies a specification showing “on its face” that an invention did not involve an inventive step. The expression derives from Commissioner of Patents v Microcell Ltd, which stands for a narrow proposition that a Commissioner of Patents, or his or her delegate, may refuse an application for patent protection where a specification “on its face” shows the invention claimed is not a manner of new manufacture. This may arise, for example, from admissions concerning novelty. The decision in Microcell has not always been properly understood; it does not involve a separate ground of invalidity or a discrete “threshold” test.
It is also possible to imagine, as Lord Hoffmann did in Biogen Inc v Medeva Plc that there may be cases where the alleged subject-matter is “so obviously not an invention that it is tempting to take an axe to the problem by dismissing the claim”. Such cases are likely to be rare. (citations omitted)
505 By reference to the case before it, the Court said (at [111]):
[T]he question of whether the concept of adding integer (vi) to integers (i)-(v) (claim 1) or to the combination of integers (i)-(v) and (vii)-(x) (claim 13) is inventive will turn on what a person skilled in the relevant art, possessed with that person’s knowledge, would have regarded, at the time, as technically possible in terms of mechanics, and also as practical. That is the sense in which an idea can involve an inventive insight about a known product. A court cannot substitute its own deduction or proposition for that objective touchstone, except in the rarest of circumstances, such as where an expressly admitted matter of common general knowledge is the precise matter in respect of which a monopoly is claimed. Even if an idea of combining integers, which individually may be considered mere design choices, is simple, its simplicity does not necessarily make it obvious. Older cases concerning simple mechanical combinations illustrate this point, as does Haberman v Jackel International Ltd. Common general knowledge has negative as well as positive aspects. Practical and technical issues can affect the means by which a concept may be implemented in respect of an already known vendible product, and scepticism can inhibit recognition of the utility of applying a concept or idea to a known set of integers. These are matters within the knowledge of relevant witnesses. (citations omitted)
506 So, the objection is narrow. Further, what was also stressed was the importance in having regard to evidence from skilled addressees in deciding whether an invention involves an inventive step. Admissions in a specification always need to be weighed with evidence from persons skilled in the relevant art addressing such matters. Further, the context of Microcell is important. In Microcell, the objection arose in the context of proceedings that were not inter partes. In other words expert evidence was not necessarily available to assist. But in the present case, expert evidence has been adduced. As Branhaven contended, there is something to be said for having the issues decided by reference to the substantive requirements of novelty and inventive step rather than dealing with the matter as a so-called “threshold requirement” test. In Bristol-Myers Squibb Co v FH Faulding & Co Ltd (2000) 97 FCR 524, it was said (at [45] per Black CJ and Lehane J):
Indeed, there is in our view an element of unreality, in a case such as the present, even in posing the question in that form. Although Philips suggests that there may be such cases (it does not decide the question, because obviousness was not pressed), it is not easy to envisage circumstances in which a claimed invention may lack the threshold requirement of inventiveness, but yet involve (for the purposes of s 18(1)(b)(ii)) an inventive step. This is not a case, like Philips, where there was no attack on the patents on the ground of obviousness. It was, instead, a case where expert evidence, including evidence as to common general knowledge, was available (and was given). Where the Court has evidence on the basis of which it can make a finding about common general knowledge, and the other information referred to in s 7(2) and (3), and about what would or would not have been obvious to persons skilled in the relevant art, it must be only rarely that it will be appropriate to find (by resort to a “threshold test”) lack of inventiveness on the face of a specification. In our opinion this is not a case where such a finding is justified.
507 Now in terms of the threshold requirement determined on the face of the specification, it is not sufficient for the specification to assert “newness” or “inventiveness”. The specification must be properly construed in order to see whether that appears on its face or conversely clearly does not. The specification includes documents referred to therein and which are incorporated into and read as part of the specification.
508 MLA submits that the claims are not to a manner of new manufacture on the basis that there is no invention on the face of the specification because the claims are only for the use of a known material in the manufacture of known articles for the purpose of which its known properties make that material suitable. I can say now that I reject such a challenge for the same reasons that I have rejected MLA’s case on lack of novelty and lack of inventive step.
509 MLA contends that the specification makes clear that all the inventors did was to:
(a) discover naturally occurring SNPs and create a high density map of the bovine genome by known methods ([0053], [0191] to [0193]);
(b) discover naturally occurring associations between those SNPs and bovine traits ([0054] to [0057], [0121], [0122], [0194] to [0197]);
(c) identify SNPs that were in LD with the specified SNPs by known methods ([0198] to [0200]); and
(d) assert that the information would be useful in known methods.
510 MLA points out that Professor Taylor accepted that the specification does not suggest that any of the techniques used to obtain the information the subject of the claims was new or special. He agreed that the inventors followed the Venter approach to create the map of the bovine genome and a comparative mapping approach that was widely accepted as at the priority date.
511 MLA says that the claims are to well-known methods of predicting genetic potential based on the presence of DNA markers associated with known desirable traits, methods of cloning and methods of using isolated DNA.
512 MLA points to what the Full Federal Court concluded in Merck & Co Inc. v Arrow Pharmaceuticals Ltd (2006) 154 FCR 31 (at [75]) that :
The Patent specification discloses no new substance, no new characteristic of a known substance, no new use and no new method. There is, therefore, no manner of new manufacture.
513 I reject MLA’s threshold argument based upon the face of the specification.
514 As I have already discussed, a fundamental aspect of this type of objection is that it must be made out on the face of the specification itself. But in the present case it cannot be clearly said that the specification “on its face” shows that the invention claimed is not a manner of new manufacture. The specification does not anywhere admit that a method, as defined by claim 1, is not new and inventive over the prior art. To the contrary, the specification provides a detailed description of the work undertaken to develop the invention. And the specification positively asserts that the resulting method is new and inventive. Moreover, and contrary to MLA’s assertions, even though there are some references to the use of known techniques or methods, they do not amount to an admission that what is claimed is not inventive.
515 Further, I agree with Branhaven that this is not a case of the “rarest” kind contemplated in Lockwood (No 2), where “an expressly admitted matter of common general knowledge is the precise matter in respect of which a monopoly is claimed” (at [111]).
516 Further, it is quite inappropriate for me to substitute my own deductions for the objective touchstone of inventive step assessed in accordance with s 7(2) of the Act. I quite agree with Branhaven that the present case should be decided based on the evidence before me relevant to the grounds of novelty and inventive step, rather than on MLA’s assertions about what is said to be disclosed on the face of the 253 Application. But even if I were to confine myself to the face of the specification properly construed, including with the assistance of the expert evidence called before me concerning questions of construction and how a skilled addressee would read and understand the specification, MLA does not come close to substantiating this type of threshold objection.
Lack of novelty
517 The legal principles are not in doubt. For a patentable invention so far as claimed in a claim, the question is whether it is novel when compared with the prior art base before the priority date. An invention is taken to be novel when compared with the prior art base unless it is not novel in the light of, inter-alia, prior art information made publicly available in a single document or prior art information made publicly available through doing a single act. The onus is on MLA to establish that such a single document or single act of prior art overcomes this presumption.
518 For there to be anticipation, the prior art, whether it is a prior publication or prior use of a product must constitute a clear and unmistakeable disclosure of each and every integer of the relevant claim the subject of the challenge. If the prior art is a document, it should be read through the eyes of the skilled addressee; terms in the prior art are to be given the meaning which the person skilled in the art would attach to them having regard to relevant common general knowledge. It is a question of the disclosure to the skilled reader. Such a disclosure may be explicit or in certain circumstances implicit. This may occur where the prior art information is a publication which does not specify an integer but the skilled reader would understand that integer to be present. If the prior art does not expressly specify each and every essential integer of the claimed invention, the evidence must clearly establish that to the skilled reader each and every essential integer is included.
519 Where the prior art is a document, to constitute anticipation the skilled addressee must be given clear and unmistakeable directions to make or perform the invention. More colloquially expressed, “the prior inventor must clearly be shown to have planted his flag at the precise destination” (General Tire & Rubber Co v The Firestone Tyre & Rubber Co Ltd (1971) 1A IPR 121 at 137 and 138). Even more colourfully expressed, “anticipation is deadly but requires the accuracy of a sniper, not the firing of a 12 gauge shotgun” (Apotex Pty Ltd v Sanofi-Aventis (2008) 78 IPR 485; [2008] FCA 1194 at [91]; H Lundbeck A/S v Alphapharm Pty Ltd (2009) 177 FCR 151 at [170]).
520 It is not sufficient to demonstrate that a prior publication is capable of being carried out in a manner which would equally infringe or not infringe the particular claim. In such a case there would not be the relevant anticipation. To elaborate, if the prior art is a document and there is ambiguity in the sense that the disclosure can be read in two or more ways, such that one way would, if carried out, infringe, and one or more other ways would not, then there has been no anticipation. Anticipation must not merely be a possibility or even a likely consequence of performing the invention disclosed by the prior art, but it must necessarily be entailed in or an inevitable result of carrying out the disclosure.
(a) Summary
521 In my view none of the prior art documents relied upon by MLA provides a disclosure, let alone a disclosure with sufficient clarity, of all of the essential features of the claimed invention. Alternately expressed, MLA has come nowhere near to establishing the requisite threshold applicable in the context of the present appeal concerning this ground of opposition.
522 MLA has relied upon 4 prior art documents. These are:
(a) Meuwissen THE, Hayes BJ and Goddard ME, “Prediction of Total Genetic Value Using Genome-Wide Dense Marker Maps” (2001) 157 Genetics 1819 (Meuwissen) (claims 1 to 6 and 8 to 10);
(b) Moody DE, Pomp D, Newman S and MacNeil MD, “Characterization of DNA Polymorphisms in Three Populations of Hereford Cattle and Their Associations with Growth and Maternal EPD in Line 1 Herefords” (1996) 74 J Anim Sci 1784 (Moody) (claims 1, 6 and 14, with leave sought to amend to include claims 3 to 5);
(c) Grosse WM, Kappes SM, Laegreid WW, Keele JW and Chitko-McKown CG, “Single Nucleotide Polymorphism (SNP) Discovery and Linkage Mapping of Bovine Cytokine Genes” (1999) 10 Mammalian Genome 1062 (Grosse) (claims 1, 3, 4, 5, 6, 9, 14 and 15); and
(d) Australian Patent Application No 2003226666 titled “Method for Identifying Animals for Milk Production Qualities by Analyzing the Polymorphism of the Pit-1 and Kappa-Casein Genes”, filed 11 March 2003 claiming priority from FR0203026 filed 11 March 2002 (the Gengler Patent) (claims 1, 6, 8, 9, 12 to 14, with leave sought to amend to include claims 3 to 5).
523 But these prior art documents do not disclose the 2,510 SNPs described and claimed in the 253 Application. They also do not disclose any of the SNPs of limb (b) of claim 1 assuming for the moment that limb (b) requires the necessary linkage disequilibrium or that an amendment is made to limb (b) to make this clear. And it also follows that none of the prior art documents disclose the use of at least 3 of the SNPs in the claimed method, where at least one of the SNPs includes a limb (a) SNP or a limb (b) SNP. Indeed, apart from arguably the Gengler Patent that I will discuss later, the other prior art documents do not make any disclosure of the requirement to use at least 3 SNPs and from more than one gene.
524 Further, MLA has failed to establish that following the clear and unmistakable directions of any of the prior art documents would inevitably result in the claimed method.
525 Further, MLA’s attempt to elide integers of the claims by recourse to “parameteritis” should also be rejected. These integers, such as the identification of at minimum number of SNPs (or the use of more than one gene) to be used in the claimed method, or the particular traits to be inferred, are not just parameters that have arisen from new measuring techniques or a statement of an underlying scientific theory. They are essential and limiting integers of the claimed method. As I have previously explained, I have rejected MLA’s misplaced diagnosis of parameteritis.
526 Further, as I have already explained in that part of my reasons dealing with construction, MLA’s contention that the requirement that the method be used for identifying a trait of a bovine subject may be ignored based on the use of “for” is rejected. In the context of the specification and as I have discussed earlier on the question of construction, it is apparent that the method must be used for the purpose of identifying a trait.
527 MLA’s reliance on Otsuka is misplaced as I have previously said. Claim 1 of the impugned patent in Otsuka was a Swiss type claim limited by a particular therapeutic use; it was a method or process claim. In relation to infringement, Yates J rejected the proposition that importation of a drug containing the active ingredient would be an infringement because the drug would be “effective” for that therapeutic use. His Honour explained that the therapeutic use limitation could not be ignored as it qualified and confined the scope of the monopoly claimed. The passage at [174] relied on by MLA merely stated that his Honour was not prepared to rely on how the medicament had been marketed, but to objectively determine the therapeutic use of the medicament from the marketing approval and registration that had been granted. Otsuka does not support the proposition that for a process claim limited by a use, an alleged infringement (and by parity of reasoning, prior art anticipation) does not have to disclose that use but merely be capable of that use. Rather, Otsuka supports the contrary proposition. I agree with Branhaven that for a method claim limited to a use, an alleged anticipation has to objectively disclose that use, not merely be effective (or capable) for that use.
528 Further, MLA’s reliance on Pharmacia & Upjohn AB (opposition by CSL Limited) [2000] APO 58 is also misplaced. As Branhaven correctly points out, the Patent Office Manual of Practice and Procedure at 2.11.2.3.3 states that “[i]n general, method or process claims using words of purpose are construed as being restricted to that purpose …” and notes Pharmacia as being an exception to that general rule.
529 Let me now deal with each prior art reference separately. For the following reasons, in my view MLA has failed to establish that any of the specified claims have been anticipated. I would make one further point. Although the various experts gave evidence on how the prior art should be read and what was disclosed, at the end of the day I was in a good position to read the prior art and make my own assessment of how the skilled addressee would so read and understand the relevant prior art.
(b) Meuwissen
530 MLA contends that claims 1 to 6 and 8 to 10 of the 253 Application are anticipated by Meuwissen.
Introduction
531 Meuwissen, as its title suggests, deals with the subject matter of predicting total genetic value using genome-wide dense marker maps. It forecasts that the advent of DNA chip technology may make genotyping of many animals for vast numbers of SNPs feasible and perhaps even cost effective.
532 But the method and simulation it describes is highly theoretical.
533 First, it is not limited to any type of animal.
534 Second, the theoretical genome modelled was assumed to consist of only 10 chromosomes (cf the bovine genome of 3 times the size).
535 Third, the 1,010 markers used were multi-allelic (7 alleles per marker) resembling microsatellites, rather than bi-allelic (2 alleles per marker) resembling SNPs. Indeed, this is confirmed within Meuwissen itself (at 1825). The statement at 1825 is not unimportant and it is appropriate that I set it out:
The markers used in the simulations more closely resembled microsatellite markers than SNPs, which are biallelic and have a much lower mutation rate. However, three to five closely linked biallelic SNP markers may be pooled to obtain ~23 different haplotypes, which resemble the about seven alleles per marker that were used in the simulation. If the closely linked markers are within a region of ~0.25 cM, their recombination rate would resemble the mutation rate of the simulated markers (Table 1). The construction of haplotypes from the SNP markers, however, requires knowledge about the linkage phase of the markers. This requires at least two generations of typed individuals (i.e. generations 1001 and 1002 here) and should be possible with high precision when the markers are closely linked; i.e. (double) recombinations are very unlikely. In situations where the linkage phases are still uncertain, this uncertainty may be accounted for in the design matrix of the haplotype, Xi, by having p and (1 – p) at the elements that belong to haplotypes A and B instead of 0 and 1 (or 1 and 0).
536 So the resemblance of the markers with microsatellites is confirmed. Further, the paper more points to using microsatellites than SNPs. Further, the paper accepted that if SNPs were used, information concerning the linkage phase would be required.
537 Fourth, claim 1 of the 253 Application used a genome wide association study (GWAS). But contrastingly, Meuwissen taught genomic selection (GS).
538 Fifth, Meuwissen presented 4 different types of statistical analysis that could be used. The first statistical approach is a least squares analysis. This is well known in terms of regression analysis and I have described it in another field in simple terms. The approach is more complicated here, as Meuwissen described at 1821. The second statistical approach, namely, Best Linear Unbiased Prediction (BLUP) involves in one sense optimising phenotypic information by correcting phenotypes for systematic effects. The third and fourth statistical approaches used Bayesian statistics: Bayes A and Bayes B. This is not the occasion to explain in detail Bayesian statistics. But some observations should be made. Classical statistics is based upon what has been described as the frequentist paradigm. Probability is associated with long-run frequency (i.e. tossing a coin infinite times). In this framework, the parameter of interest is assumed to be unknown but fixed. There is assumed to be say one true mean or one true regression co-efficient for the population parameter. The Bayesian paradigm interprets probability as the subjective experience of uncertainty. There are three essential ingredients underlying Bayesian statistics. The first ingredient is the background knowledge on the parameters of the model being tested. This first ingredient refers to all knowledge available before seeing the data and is captured in the so-called prior distribution, for example, a normal distribution. The variance of this prior distribution reflects the level of uncertainty about the population value of the parameter of interest: the larger the variance, the more uncertainty. The second ingredient is the information in the data itself. It is the observed evidence expressed in terms of the likelihood function of the data given the parameters. In other words, the likelihood function asks: given a set of parameters, such as the mean and/or the variance, what is the likelihood or probability of the data in hand? The third ingredient is based on combining the first two ingredients, which is called posterior inference. Both the first and second ingredients are combined via Bayes’ theorem and are summarised by the so-called posterior distribution, which is a compromise of the prior knowledge and the observed evidence. The posterior distribution reflects one’s updated knowledge, balancing prior knowledge with observed data. In the Bayesian approach, all unknown parameters are treated as uncertain and to be decided by a probability distribution. Meuwissen used two different Bayesian methods (Bayes A and Bayes B) which are described at 1822 and 1823.
539 Now what is significant to note for present purposes is that Meuwissen reported (at 1826) that of the four methods of statistical analysis used, the least squares method performed “badly”. It was reported that the least squares method “greatly over-estimate[d] some haplotype effects and underestimate[d] others”. Now it is to be noted that Bayes A and BLUP did not require association, but least squares does require association (and so too may Bayes B).
MLA’s contentions
540 MLA says that Meuwissen describes the prediction of total genetic value using genome-wide dense marker maps. The authors state that the total number of SNPs is estimated at many millions (citing Halushka MK, Fan JB, Bentley K, Shen N, Weder A, Cooper S, Lipshutz R and Chakravati A, “Patterns of Single-Nucleotide Polymorphisms in Candidate Genes for Blood-Pressure Homeostasis” (1999) 22 Nat Genet 239). It is said that the advent of DNA chip technology may make genotyping of many animals for many of these markers feasible and perhaps even cost effective (at 1819).
541 The authors simulated estimation of the effects of approximately 50,000 marker haplotypes simultaneously from a limited number of phenotyped animals, each with a proposed genome of 1,000 cM with a marker spacing of 1 cM.
542 The following conclusions were drawn by the authors (at 1828):
(a) “By using a dense marker map covering all chromosomes, it is possible to accurately estimate the breeding value of animals that have no phenotypic record of their own and no progeny”.
(b) “Selection on breeding values predicted from markers could substantially increase the rate of genetic gain in animals and plants especially if combined with reproductive techniques to shorten the generation interval”.
543 Professor Goddard and Professor Hayes, who gave evidence for MLA, are co-authors of Meuwissen. This paper was published in the Journal of the Genetics Society of America, Genetics, which is said to be a very reputable journal in the broad field of genetics, which includes cattle breeding.
544 All experts who gave evidence on the subject before me agreed that Meuwissen describes a theoretical approach to identify DNA markers across the whole of the bovine genome and the use of statistical models and methods to determine associations between those markers and traits. MLA contends that that is the approach that was used for the human genome and is broadly the same approach that the 253 Application describes and claims.
545 MLA contends that although there is some disagreement as to whether Meuwissen teaches the reader to use the method for the purposes of identifying the genetic potential of animals, it clearly describes the potential for selective breeding based upon predicted breeding values as a use for the method (see e.g. the Abstract, last sentence, at 1819 and the Conclusions, at 1828). Indeed, Meuwissen indicates that the ultimate purpose is selective breeding (Conclusions, paragraph 4, at 1828). I would note that Dr Sonstegard for Branhaven did not consider that the claims of the 253 Application were limited to selective breeding, although they included the purpose of selective breeding. In any event, so MLA contends, it is sufficient for the purposes of claim 1 of the 253 Application that the method is suitable for that purpose.
546 MLA contends that a person skilled in the relevant art and following the approach in Meuwissen would have inevitably identified at least some SNPs covered by the claims of the 253 Application, discovered any natural associations of those SNPs with a trait, and used a standard mathematical algorithm to analyse that information in order to predict the genetic potential of animals.
547 MLA points to Professor Visscher’s explanation that given that the aim of the method of Meuwissen is to cover the whole genome with SNPs, and given the number of SNPs required to do so, there would be significant overlap between the region covered by claim 1 of the 253 Application and the SNPs identified by implementation of the method described in Meuwissen. It is said that that is because claim 1 of the 253 Application extends to about +/- 500,000 nucleotides of the 2,510 specific SNPs. It is said that this region encompasses around two thirds of the bovine genome. Therefore, so it was argued by MLA, Meuwissen inherently directs the reader to identify and use at least three markers (e.g. SNPs) within the region covered by claim 1 of the 253 Application.
548 MLA contends that in any event, as the requirement that there are at least 3 SNPs occurring in more than one gene is an arbitrary parameter, this feature can be ignored for the purpose of novelty. I have found against MLA on this point as I have explained in these reasons.
549 Further, MLA contends that Meuwissen is directed to selection for economically important quantitative traits in livestock and crop species (Meuwissen at 1819, first column, see also at 1829, conclusion 4). Livestock includes cattle. Further, MLA points out that Meuwissen references other articles that in turn mention the cattle genome or genomic improvement in cattle.
550 Further, MLA says that contrary to Dr Sonstegard’s evidence, Meuwissen does teach the use of SNPs. MLA points for example to 1825, right column, of Meuwissen, where it is stated that three to five closely linked biallelic SNP markers may be pooled to obtain ~23 haplotypes which resemble the ~7 alleles per (microsatellite) marker used in the simulation. Furthermore, Professor Plastow accepted that Meuwissen discloses that SNPs could be used in a whole genome scan. Further, Professor Hayes explained that the method in Meuwissen would in fact be simpler to use with SNPs than with microsatellites.
551 MLA also contends that the expert evidence demonstrates that following the approach in Meuwissen it is inevitable that more than three SNPs occurring in more than one gene, i.e. in at least two genes (understood as protein coding regions), would be identified. Meuwissen states at 1820, left column, that on the basis of comparative mapping, all 50,000 or so genes in the cattle genome will be identified with many of those genes having SNPs within them. Professor Visscher explained that tens of markers utilised in the approach of Meuwissen will be located within a gene simply based on the assumption that genes make up around 2% of the bovine genome. Further, Professor Hayes indicated that 25.3% of the nearly 26.72 million SNPs he and his colleagues had considered as part of his “1000 bulls genome project” were within genes. Therefore, so MLA contends, the application of Meuwissen using SNPs would likely result in about a quarter of the SNPs being in genes. Further, as Professor Hayes explained, he would expect about 25% of SNPs in the bovine genome would appear within genes if SNPs were selected randomly. Therefore, so MLA contends, the method of Meuwissen inherently directs the reader to identify and use at least three markers (e.g. SNPs) that are associated with a trait wherein “the at least three markers (e.g. SNPs) occur in more than one gene”.
552 Further, MLA rejects Branhaven’s criticism (and those of Branhaven’s experts) that the simulation in Meuwissen is only theoretical. It says that such simulations are commonly published in the area of breeding because actual breeding experiments can take a long time; it relies upon Professor Plastow’s evidence in support. Further, it says that the way the simulation was carried out in Meuwissen mirrored what happens in real world cattle populations, as Professor Goddard indicated. It says that there was confidence amongst quantitative geneticists that the method in Meuwissen would work, as Professor Visscher indicated. In any event, MLA says that an invention can be patentable even if it was only conceived in the mind (it can be “remembered from a dream” (Wellcome Foundation Ltd v VR Laboratories (Aust) Pty Ltd (1981) 148 CLR 262 at 286)), if it otherwise satisfies the requirements of the Act. Likewise, a publication can anticipate even if it is purely theoretical. In Merck & Co Inc v Arrow Pharmaceuticals Ltd (2006) 154 FCR 31, the Full Federal Court (Heerey, Kiefel and Dowsett JJ) upheld the primary judge’s decision that Lunar News anticipated the claims of a patent to a method of treatment despite the fact that the article said that the proposed new dosing regimen needed to be tested (at [104]). The Court rejected the notion that the need for routine testing and experimentation meant that a prior suggestion could not be novelty defeating (at [104] to [112]).
553 Further, MLA says that the evidence establishes that implementation of the approach described in Meuwissen could be performed using well known techniques, although there were practical difficulties such as time and cost. But such practical difficulties are not relevant to the question of anticipation.
554 Further and apparently for good measure, MLA also contends that the 253 Application is also in one sense only theoretical, because it did not validate any associations of SNPs with traits in new cattle, i.e. in cattle breeds other than the specific breeds used in the study.
555 Further, MLA also takes issue with the position of Branhaven and its experts that Meuwissen is different from the 253 Application because Meuwissen teaches a GS approach, rather than a GWAS approach as used in the 253 Application. MLA says that there is nothing in this point. It says that the steps described by Meuwissen are the same steps as claimed in the 253 Application. In essence, Meuwissen teaches GS and the 253 Application teaches GWAS plus predicting genetic potential based on multiple markers, which MLA says is equivalent to GS. For this purpose, MLA relies upon the evidence of Professor Goddard and Dr Sonstegard.
556 Further, MLA contends that the approach described in Meuwissen was based on the knowledge that the associations between DNA markers and traits can be used to infer traits. It was also based on the knowledge (including from analysis of the human genome) that if there were sufficient DNA markers (such as SNPs) in the genome, some of the DNA markers will be close enough to the linked gene governing or affecting a trait to be associated with that trait (i.e. the closer together the DNA marker and the gene, the stronger the linkage disequilibrium (LD)). This meant that each individual marker did not need to be tested individually for association, but could be tested at the same time using the statistical model. That model made certain assumptions, including the number of markers that would be required to detect associations, the degree of LD in cattle, and the population size required. The data created by the model were then analysed, using four standard statistical algorithms, to determine if any of the DNA markers were significantly associated with a trait. The DNA markers associated with a trait could then be used to predict genetic potential.
557 MLA contends that the 253 Application describes the creation of a genome wide map of DNA markers in order to do a GWAS, as does Meuwissen (with respect to the map). The SNPs in that map were each then tested individually for association with a known trait using a standard algorithm, as was done in Meuwissen. The 253 Application claims a method of inferring traits from associations between multiple SNPs and traits with the aim (not claimed) of selectively breeding cattle, which was taught in Meuwissen.
558 In summary, MLA contends that both Meuwissen and the 253 Application describe:
(a) The creation of a genome wide map of DNA markers;
(b) Using genotypic and phenotypic data of cattle to measure association between markers and traits; and
(c) Using statistical analysis to work out which markers were useful for predicting genetic potential.
559 MLA says that the only difference in approach is the statistical method used. In Meuwissen, two of the four algorithms assumed all markers potentially have a degree of association with a QTL or gene that influences a trait (Bayes A and BLUP). The other two algorithms (the least squares and Bayes B approaches), looked at each SNP individually and tested for association on a level of stringency; I would note that a least squares approach also resonates with a GWAS style approach as Professor Taylor suggested. Further, MLA says that while Meuwissen says that the least squares approach is the least effective, it does not say it does not work. In fact, so MLA contends, it did work, just not as well as the others; MLA relies upon Professor Hayes’ evidence to this effect. In any event, so MLA contends, Meuwissen discloses no such reservations about the Bayes B method, which only uses markers which are statistically significant. MLA says that the 253 Application tests each SNP individually using an algorithm that is close to the least squares approach used in Meuwissen. In any event, MLA says that the fact that the algorithms used in Meuwissen and the 253 Application were not identical is not relevant, as the claims do not require association to be determined in any particular way, and therefore encompass all methods of determining association.
560 Accordingly, MLA contends that there is no relevant difference in approach between Meuwissen and the 253 Application.
561 Further, MLA points out that Professor Visscher explained that, if the method of Meuwissen was performed in cattle, a number of the specified SNPs (i.e. SEQ ID NOS: 19473 to 21982) might be identified and used. Therefore, MLA says that Meuwissen discloses the features of claim 2 of the 253 Application.
562 Further, MLA points out that claims 3 to 5 of the 253 Application require the use of at least five SNPs (claim 3), at least seven SNPs (claim 4), or at least ten SNPs (claim 5). MLA says that these features are arbitrary parameters, a contention that I have rejected in these reasons. Alternatively, it says that they are inherently disclosed in Meuwissen because the approach of Meuwissen will result in the use of many markers.
563 Further, as to claim 6, MLA says that with respect to step (a) of claim 6, these features are identical to those of claim 1. With respect to step (b) of claim 6, MLA says that this step is implicit in the disclosure of Meuwissen in that once an animal has been identified as having a polymorphism associated with a trait, one would sort those animals with the trait from those that do not display such a trait. Repeating steps (a) and (b) of claim 6 would also be performed to identify and sort additional animals. MLA points out that Professor Visscher explained that the implementation of the method in Meuwissen would inherently involve the ranking or sorting of cattle and therefore it discloses a method as defined by claim 6.
564 Further, as to claim 8 MLA says that Meuwissen discusses the future use of SNP-chip or microarray technology, which is required by the method of inferring a trait in a bovine as defined by claim 8 of the 253 Application.
565 Further, as to claims 9 and 10 of the 253 Application, MLA points out that Professor Visscher explained that the selection for traits as discussed in Meuwissen:
include[s] selection for those traits that are economically important in cattle, such as traits relevant to production, reproduction, health, management or physical appearance. Such traits would include those traits set out in claims 9 and 10 of the Opposed Application and indeed any other traits of economic importance for beef or dairy cattle.
566 Further, MLA says that Meuwissen specifically refers to disease resistance (at 1827, second column), which is included in the list of traits in claim 9. Further, MLA says that although Meuwissen does not specifically refer to one of the carcass traits in claim 10, there can be no novelty or invention in the selection of an economically significant carcass trait to use in the method. It says that a claim limited to a particular desirable trait or group of desirable traits is mere parameteritis.
Analysis of Meuwissen
567 Let me now analyse Meuwissen and set out my principal conclusions bearing in mind that in the context of the present appeal I must be clearly satisfied that Meuwissen anticipates for MLA to succeed on this ground of opposition. I should say at the outset that I am not so satisfied to the requisite degree.
568 First, Meuwissen is not specifically directed to cattle. And it describes a theoretical genomic selection model for analysing data generated from large numbers of simulated markers spread throughout the genome of an organism. Meuwissen was based on a hypothetical genome consisting of 10 chromosomes and hypothetical markers.
569 Second, and contrary to MLA’s contention, I am not satisfied to the requisite degree that a skilled person following Meuwissen would have identified some SNPs covered by the claims of the 253 Application. Indeed, in my view it is not inevitable that a skilled person at the priority date would have identified any SNPs at all (let alone in bovine). The skilled person might well have identified other markers, such as microsatellites.
570 Third, as Dr Sonstegard and Professor Taylor have explained, the markers disclosed in Meuwissen are hypothetical. And they resemble microsatellites more than they resemble SNPs. The markers modelled in Meuwissen had 7 alleles at each locus whereas SNPs are generally bi-allelic. This was accepted by Professor Goddard and Professor Hayes, who also accepted that one way of applying Meuwissen would be to use microsatellites. Now Meuwissen discloses that a number of SNPs together could be used to substitute for microsatellites. But Meuwissen assumed phase information would be available. But such information may not have been available as Professor Taylor explained. Further, there was evidence suggesting that microsatellites are more informative than SNPs as they have multiple alleles at each locus compared with SNPs which are usually biallelic. Further, there was evidence from Dr Sonstegard and Professor Taylor to the effect that microsatellites were the most widely used marker in cattle genetics as at December 2002. They therefore produced results that were more reproducible across laboratories. Further, there were also on the evidence before me far too few SNPs known at the time to make a dense marker map.
571 I agree with the submission of Branhaven that a person skilled in the art seeking to apply Meuwissen at December 2002 to cattle with real markers was far more likely to have used microsatellites instead of SNPs, or at least as likely to have used microsatellites. And such a conclusion is consistent with the evidence of Professor Hayes, Professor Goddard and Dr Sonstegard that there were a sufficient number of known microsatellites, but not SNPs, at the priority date to conduct a genome wide study. Accordingly, Meuwissen did not give clear and unmistakeable directions to use SNPs. The directions given in Meuwissen were at least as capable of being carried out using microsatellites. And it is not to the point that Meuwissen indicated that you could use SNPs or that some of the experts may have had a preference for using SNPs. The question is what was the clear and unmistakeable direction of Meuwissen.
572 Fourth, it was accepted by the experts before me that the authors in Meuwissen accurately stated that:
[t]he aim of this article is to compare least squares, BLUP and Bayesian analyses for their accuracy of predicting total breeding value of individuals in a situation where a limited number of recorded individuals are genotyped for many markers with many alleles per marker (1820, left hand column).
573 Both Professor Goddard and Professor Hayes accepted that the BLUP and the Bayes A methods do not identify particular markers associated with the trait. Rather, such methods use all the markers in the dense marker map in the analysis. Further, Professor Goddard accepted that one way of applying Meuwissen to real animals and real markers was to use the BLUP and Bayes A method of analysis. But as I have said, these would not identify particular markers with a trait. Contrastingly, it was accepted that the other two methods, Bayes B and least squares do identify specific markers associated with a trait. Moreover, Professor Goddard and Professor Hayes also accepted that of the 4 methods, the least squares method did not work as well as the others.
574 Now given just that evidence alone, and my own review of Meuwissen, it follows that the direction in Meuwissen (if applied to real animals and real markers) is at least as likely to be carried out in a way by using BLUP and Bayes A that does not identify specific markers associated with a trait. But that is not what is required by the claims of the 253 Application.
575 Fifth, Meuwissen does not disclose the 2,510 SNPs of the 253 Application. It also does not disclose SNPs within about 500 kb and in relevant linkage disequilibrium, assuming for the moment that this is what is implied in limb (b) of claim 1 or what I stipulate I may permit by way of amendment. I agree with Branhaven, that no sufficiently probative evidence has been adduced before me that would clearly establish the proposition that a skilled person who identified a set of SNPs from the bovine genome would inevitably identify any of the identified limb (a) SNPs and the limb (b) SNPs in linkage disequilibrium with them. They would just as likely identify different SNPs and different SNPs in linkage disequilibrium therewith.
576 Sixth, I also accept Branhaven’s point that in relation to the claims of the 253 Application, even if it were assumed that a skilled person had chosen to use SNPs rather than microsatellites and had chosen to do a GWAS to identify SNPs associated with a trait, rather than applying the BLUP and the Bayes A methods which use all the SNPs in a dense marker map, it is clearly not inevitable that such an approach would involve identifying 3 SNPs whereby one was either one of the specified SNPs or within about 500 kb of a specified SNP and in linkage disequilibrium with it (assuming that limb (b) requires this).
577 In summary, MLA’s case based upon anticipation by Meuwissen must be rejected in respect of claim 1. Further, given that claims 2 to 5 are dependant claims, MLA’s challenge also fails based upon Meuwissen. As for claims 6 and 8 to 10, limb (a) of claim 6 and claims 8 to 10 have as essential integers in substance limbs (a) and (b) of claim 1. The novelty challenge based upon Meuwissen must also fail in relation to these claims for reasons that I have already explained.
(c) Moody
Introduction
578 Moody involved a study of three populations of Hereford cattle that were genotyped for seven DNA polymorphisms; SNPs were used (see at 1786). Differences in allele frequency among the populations were found at six of the seven polymorphisms genotyped.
579 But only one SNP (substitution of a B allele for an A allele of the kappa-casein polymorphism) was considered to be a potentially useful predictor, and in that case of two traits.
580 At 1786 it was stated:
The loci investigated are all located on different bovine chromosomes (Barendse et al., 1994; Moody et al., 1995a,b). Polymorphisms within the GH, K-Cas, and B-Lac genes are in coding regions and represent different forms of the proteins coded by those genes. Thus, the K-Cas, B-Lac, and GH polymorphisms have the potential of identifying a direct physiological effect resulting from differences in the amino acid sequences of their resulting proteins, as well as being markers of linked QTL. In contrast, polymorphisms in the PIT-1, IGF-I, GHR, and PRL genes, as well as the BM2113 microsatellite, are located in non-coding regions and should only be considered as potential markers of linked QTL.
581 Table 3 presented data for K-Cas, B-Lac, GH, IGF-I and PIT-1. But only significant R2 values were given for K-Cas and IGF-I. The R2 values reflected the percentage of variability of expected progeny differences explained by genotype.
582 At 1791 and 1792, it was concluded:
The structure of the Line 1 population, which included several small half-sib families, is far from ideal for a study designed to identify QTL, especially when using non-coding region markers such as IGF-I and PIT-1. More efficient and appropriate experimental designs have been proposed (Weller et al.. 1990; Keele, 1994) but were not possible with existing resources. As a result, the power of the statistical analysis in this study was limited and significant effects explaining small amounts of variation are likely spurious. The large effect of IGF-I on BW-d should also be interpreted with caution because this study was unable to follow the segregation of IGF-I alleles in large half-sib families. However, the effects of K-Cas genotype on BW-d and PR-m may indicate a direct effect of the K-Cas gene or of a very closely linked gene. The magnitude of these effects, as well as evidence that the K-Cas polymorphism is linked to a QTL influencing milk production in dairy cattle (Bovenhuis et al., 1992: Cowan et al., 1992: Bovenhuis and Weller, 1994), indicates K-Cas genotype may be a useful marker for BW-d and (or) PR-m in future marker-assisted selection programs in Hereford cattle.
Implications
Significant differences in allele frequencies were found among three populations of Hereford cattle at six of seven DNA polymorphisms evaluated, indicating potential differences in genetic backgrounds exist among populations within breeds. In Miles City Line 1 Herefords, a polymorphism in the kappa-casein gene accounted for 15 and 8% of the variability in expected progeny differences for birth weight and 180-d gain from birth to weaning, respectively, indicating that kappa-casein may be a useful marker for these traits in future marker-assisted selection programs in Hereford cattle.
583 Clearly, the result for IGF-I could be put to one side or treated with caution at the least. All that was said to be left in the frame as a suitable predictor was the K-Cas polymorphism.
584 I would also note at this point that Moody contained no express disclosure of:
(a) using the combination of 3 SNPs to predict a trait;
(b) limb (a) of claim 1 of the 253 Application; or
(c) limb (b) of claim 1 of the 253 Application.
585 And I would also note that Moody appears to have reported on a potentially useful result in relation to one SNP, contrary to the evidence of some of MLA’s witnesses (at least as I understood it) that one SNP (or even three SNPs) could not be a useful predictor of phenotype. However, even that result seems to have been spurious, as I will come to later.
MLA’s contentions
586 MLA contends that claims 1, 6 and 14 are anticipated by Moody. MLA also contends that claims 3 to 5 are anticipated. MLA seeks leave to amend its Notice of Appeal to also assert lack of novelty in respect of those claims in light of Moody. I will allow such an amendment, although the assertion fails in any event for reasons that I will later explain.
Claim 1
587 MLA says that Moody evaluates the effects of polymorphisms in seven genes with respect to their genetic potential for growth characteristics in cattle. It is said that the purpose of Moody is to identify polymorphisms in genes that are associated with various growth characteristics in cattle and to identify useful markers for these traits in future marker-assisted selection programs in cattle.
588 MLA contends that the growth characteristic traits observed in Moody included direct EPD (expected progeny differences) for birth weight (BW-d), maternal EPD for birth weight (BW-m), direct EPD for 180-day gain from birth to weaning (PR-d), maternal EPD for 180-day gain from birth to weaning (PR-m), and direct EPD for gain from weaning to yearling (PO-d) (see at 1785, right column).
589 MLA contends that the seven genes referred to in Moody included κ-casein (K-Cas or kappa casein), β-lactoglobulin (B-Lac), growth hormone (GH), pituitary transcription factor 1 (PIT-1), growth hormone receptor (GHR), insulin-like growth factor I (IGF-I) and prolactin (PRL).
590 Further, MLA contends that with respect to the polymorphisms themselves, the polymorphisms in the IGF-I and PRL genes were microsatellite markers, whereas the polymorphisms in the K-Cas, B-Lac, GH, PIT-1 and GHR genes were PCR-based bi-allelic restriction fragment length polymorphisms within the genes (see at 1786, left column). Accordingly, MLA says that the polymorphisms in the K-Cas, B-Lac, GH, PIT-1 and GHR genes are therefore SNPs.
591 MLA contends that Moody describes SNPs in each of the coding regions of three of those genes. Further, MLA says that regardless of what definition of “gene” is applied, all definitions include the exons (coding regions) of a gene.
592 Accordingly, MLA says that its analysis of the position of genes in which the SNPs referred to in claim 1 of the 253 Application are located shows that at least the K-Cas and PIT-1 genes fall within 500,000 nucleotides of a specified SNP. Therefore, so MLA says, the SNPs within those genes will fall within 500,000 nucleotides of a specified SNP as required by limb (b) of claim 1 of the 253 Application.
593 Further, MLA says that the left column of 1786 of Moody indicates that genomic DNA (that is, a nucleic acid sample within the meaning of claim 1 of the 253 Application) was used to genotype the five SNPs in each of the K-Cas, B-Lac, GH, PIT-1 and GHR genes.
594 Further, MLA says that Table 3 of Moody (at 1790) provides the results of the effects of polymorphisms in the K-Cas, B-Lac, GH, IGF-I and PIT-1 genes on growth characteristics of bovine. Table 3 shows several statistically significant associations (a p-value in column 4 of <0.01) between 5 genes and various traits. With respect to the K-Cas gene as an example, the SNP in this gene is shown to have a significant association with the BW-d, BW-m, PR-d and PR-m traits. Furthermore, each SNP in the K-Cas, B-Lac and PIT-1 genes has a significant effect on the BW-d trait, and each SNP in the K-Cas, GH and PIT-1 genes has a significant effect on the BW-m trait. Further, it says that throughout Moody, and in particular at 1786, left column and Table 3, the authors describe the effect of a single SNP in each of 5 genes.
595 Accordingly, so MLA contends, by performing the method disclosed in Moody, one would be performing the method defined by claim 1 of the 253 Application. To support that conclusion MLA relies upon, inter-alia, Professor Visscher’s evidence.
596 But MLA had to concede that Moody goes on to doubt the statistical significance of the associations reported at 1791 second full paragraph. But MLA points to the conclusion at the end of the paper that “kappa-casein may be a useful marker for these traits in future marker-assisted selection programs in Hereford cattle”. Accordingly, so MLA contends, that is a clear direction to use the K-Cas SNP in future marker-assisted selection programs.
597 Further, MLA relied upon the explanation of Professor Visscher that:
Whilst Moody does not specify how the reported association is to be used in marker assisted selection, Moody reports on statistically significant associations between polymorphisms and a trait of economic importance across Hereford cattle that could be used in marker assisted selection to identify (or make a prediction as to the genetic potential) cattle for that trait.
598 So, MLA says that the information disclosed in Moody provides a “method for identifying [the BW-m] trait of a bovine subject from a nucleic acid sample of the bovine subject” by the use of the SNP in K-Cas (cf the opening words of claim 1 of the 253 Application). In any event, it says that all that is required is suitability for that purpose. Further, MLA says that given that the number of SNPs used in claim 1 is arbitrary, it matters not that Moody only specifically recommends one SNP for future use. Again, I have rejected this assertion of arbitrariness and, accordingly, that conclusion fails.
Claims 3 to 5
599 MLA’s case concerning claims 3 to 5 in essence rises or falls with the strength of its arguments concerning claim 1.
Claim 6
600 As to step (a) of claim 6, MLA says that these features are identical to those of claim 1. As to step (b) of claim 6, MLA relies, inter-alia, upon the evidence of Professor Visscher who explained that:
any application of the disclosure in Moody in marker assisted selection would involve identifying animals having those traits and the sorting or ranking of those identified animals from others as part of a breeding program.
Claim 14
601 As to claim 14 of the 253 Application, MLA says that the methods disclosed in Moody for identifying the SNPs in the K-Cas, B-Lac, GH, PIT-1 and GHR genes were PCR-based bi-allelic restriction fragment length polymorphism detection methods, again relying upon the evidence of Professor Visscher. This required the use of isolated genomic DNA and PCR amplification of a fragment of DNA containing each SNP. Each PCR amplified fragment is “isolated” in the sense that it is an artificial sequence. According to the evidence of Professor Visscher, the PCR amplified fragments in Moody would be at least 20 contiguous nucleotides of the bovine genome, or a complement thereof, as this is typical for primers used for PCR amplification. These fragments would then be used according to the methods of at least claims 1 and 6 of the 253 Application as Professor Visscher explained. Further, MLA says that the remaining features of claim 14 are identical to those of claim 1.
Other matters
602 MLA says that the claims of the 253 Application do not require any validation of an association identified between the SNPs and a trait. Likewise Moody in Table 3 identifies such a non-validated association. Further, it says that notwithstanding that Moody goes on to doubt the validity of the associations between the SNPs and traits, it recommends at least one SNP for marker assisted selection. MLA says that in relation to the 253 Application, three SNPs (and on at least 2 genes) are arbitrary features of the claimed invention, with one SNP being enough, a proposition that I have indicated I reject.
Analysis of Moody
603 In my opinion, MLA has not clearly established that the said claims fail for lack of novelty based upon Moody.
604 First, Moody analyses the extent of genetic variation between three populations of Hereford cattle by looking at 7 polymorphic loci. The 7 polymorphisms include 5 bi-allelic Restriction Fragment Length Polymorphism (RFLP) markers and 2 microsatellites. But the only polymorphism which the authors speculate may be useful as a marker is the polymorphism within K-Cas (at 1791, column 2). But the relevant claims of the 253 Application require that the SNPs be associated with a trait. Now MLA has heavily relied upon Table 3 to contend that it discloses an association between 3 or more SNPs and a trait. But Moody upon further analysis concluded that there were no significant effects of genotype on the tested traits (at 1791 left column paragraph 3). Moreover, Professor Taylor gave evidence before me that all of the associations in Table 3 were spurious. Moreover, Professor Goddard ultimately gave evidence that his inclination was that “probably none of the effects are real”. I am not clearly satisfied that there was any clear and unmistakeable disclosure of any associations. And as I have said, I will require claim 1 of the 253 Application to be amended to deal with the appropriate stipulation of statistical significance concerning “association”; in other words claim 1 does not embrace or should not embrace spurious associations, which is all that Moody may be said to disclose.
605 Second, let it be accepted for the moment that Moody discloses that the K-Cas genotype may be a useful marker for 2 traits. And let it be accepted that Moody discloses 1 SNP associated with 2 traits, there is no clear and unmistakable disclosure of the number of SNPs in claims 1 to 5. And as I have said, the number of SNPs, and the requirement of more than one gene, are not arbitrary.
606 Third, none of the SNPs disclosed in Moody have been shown on compelling evidence to be in linkage disequilibrium with any of the specified SNPs in the 253 Application.
607 Accordingly, I reject MLA’s contention that claim 1 has been anticipated by Moody. The submissions in respect of claims 6 and 14 fail for the same reasons.
608 Further, to the extent that MLA contends that claims 3 to 5 of the 253 Application are anticipated, as is also apparent from my reasoning, Moody does not disclose the use of at least 5, 7 or 10 SNPs, all associated with a trait.
609 MLA’s lack of novelty case based upon Moody is rejected.
(d) Grosse
Introduction
610 Grosse describes the identification of 31 SNPs as markers for a set of seven bovine cytokine loci and their use in linkage mapping. It was hypothesised that allelic variation in cytokines (or more correctly genes that encode for cytokines, a group of proteins) may be associated with variation in response to infection in beef cattle populations.
611 The authors explained by way of introduction (at 1062), the following:
Infectious diseases are a significant source of economic loss to the cattle industry (NASS 1996). In addition, cattle infected with foodborne pathogens such as Escherichia coli O157:H7 and Salmonella typhimurium DT104 have become an emerging human health issue (Glynn et al. 1998; Mead and Griffin 1998). A potential method for reducing the impact of infectious disease is to increase host genetic resistance to infection. Among the genes likely to influence the magnitude, duration, and transmission of infection are those that encode cytokines. This group of proteins plays a pivotal role in mobilizing inflammatory cells in response to infectious challenges (Krakauer et al. 1999). Moreover, specific allelic variants of cytokines and their receptors profoundly influence infection phenotypes in humans (Smith et al. 1997; Altare et al. 1998; Martin et al. 1998; Winkler et al. 1998; Knight et al. 1999).
In an effort to identify bovine genes that are modulated during infection, we previously reported two pro-inflammatory cytokine transcripts whose abundance increased in epithelial cells upon exposure to E. coli O157:H7 lipopolysaccharide (Heaton et al. 1999). Both cDNAs encoded members of the CXC (or α) chemokine subfamily, specifically interleukin (IL)-8 and epithelial cell inflammatory protein (ECIP)-1. We hypothesize that allelic variation in chemokines, cytokines, or their receptors may be associated with variation in response to infection in beef cattle populations. Testing this hypothesis requires genetic markers for identifying allelic variants at each gene locus. In humans, there are 11 known CXC chemokines and more than 150 cytokines and receptors (Vaddi et al. 1997; Thomson 1998). This paper describes the identification of 31 single nucleotide polymorphism (SNP) markers for a set of seven bovine cytokine loci and their use in linkage mapping.
612 The authors concluded in part at 1068:
Because developing bovine amplification primers from human sequence was sometimes difficult, bovine genomic DNA sequence made available via public databases (e.g. IFN-γ and TNF-α) was an important resource in the SNP discovery process. IFN-γ is a potent activator of macrophages and monocytes and greatly augments their antibacterial activities (Boehm et al. 1997). Though the IFN-γ gene has been localized to BTA 5q22-5q24 by fluorescence in situ hybridization (Chaudhary et al. 1993), genetic markers for linkage analysis have not been described. TNF-α is considered a major inflammatory mediator and, in humans, variation in the TNF-α gene promoter has been associated with several infectious diseases (Hill 1998; Knight et al. 1999). SNP markers for these genes may be useful for evaluating whether variants are associated with similar traits in cattle.
In addition to determining linkage map position and marker order, analysis of SNP segregation patterns in reference families provided three important results. First, it allowed definition of SNP haplotype alleles within an amplicon. Unlike the individual SNPs that are biallelic, the set of SNP alleles constituting a haplotype is not limited to two alleles. The SDF1 amplicon, for example, contained six SNP haplotypes in the reference population. The use of SNP haplotypes in linkage analysis typically reduced the number of cases in which parents and progeny had identical heterozygous genotypes. These cases (termed “switches”) present computational difficulties for linkage analysis algorithms like CRI-MAP. A second advantage of analyzing segregation patterns in reference families was the detection of apparent heterozygous sites that result from sequencing “artifacts”. These artifacts were occasionally apparent even when a high-quality sequence was obtained from both DNA strands. The artifacts were revealed when non-Mendelian inheritance patterns were observed in reference progeny. A third benefit of segregation analysis was the discovery of unamplified alleles in certain families. This phenomenon caused animals with heterozygous haplotypes to appear homozygous. Unamplified alleles are postulated to occur when SNPs were present in the primer binding region. Mismatches between the primer and template may destabilize DNA duplex formation and reduce the amplification of specific haplotype alleles. For these reasons, it is imperative that SNP haplotype loci be extensively evaluated in families with defined pedigrees prior to their application in populations with poorly defined genetic backgrounds.
613 Clearly, the authors were reporting that the SNPs identified could be useful in mapping traits in cattle, particularly resistance to infectious disease. But an association study was necessary.
614 I would note at this point that unlike claim 1 of the 253 Application, there is no express disclosure of:
(a) the need for 3 SNPs;
(b) limb (a) or limb (b); or
(c) the method of claim 1, which requires an association.
615 But on the question of association, I would also note at this point that Professor Goddard gave evidence that Grosse was no different to claim 1, as even for claim 1 an association study was necessary for your population.
616 I should also note that Dr Whitmore gave the following evidence concerning Grosse:
Grosse reports the discovery of 31 SNPs in 7 bovine chemokine genes (namely GRO3, GRO1, GROX, IL8RB, SDF1, IFNG, TNFA), and the use of those SNPs to linkage-map each of the genes in a “reference population” of cattle maintained by the USDA Meat Animal Research Center at Clay Center, Nebraska.
Some of the discovered SNPs are within an intron of the relevant gene; others are within an exon of the relevant gene. The number of SNPs discovered within each gene are: 3 in GRO3, 2 in GRO1, 6 in GROX, 4 in IL8RB, 6 in SDF1, 7 in IFNG, and 3 in TNFA (giving a total of 31).
With respect to the IL8RB gene (which at the time of the Grosse publication was also known as the CXCR2 gene - Ensembl database reference ENSBTAG00000038042), a number of the research articles citing Grosse disclose an association between the c735C>G SNP in the gene (as disclosed in Grosse) and resistance to mastitis (a debilitating disease with respect to the dairy cattle industry). The c735C>G SNP changes an amino acid (glutamine to histidine) at position 245 of the encoded protein. This SNP has also been referred to as +777 in a number of the research articles citing Grosse, which I have read. As set out below, the c735C>G SNP was subsequently annotated to occur within the CXCR1 gene (Ensembl database reference ENSBTAG00000026753).
One article citing Grosse is Youngerman et al., 2004, J. Dairy Sci., 87: 2442-2448. Youngerman discloses that Holstein cattle expressing the GG genotype at the +777 SNP had a decreased incidence of subclinical mastitis.
A thesis publication by Russell in 2013 discloses at page 160 that the GC genotype at the c735C>G SNP was associated with a higher incidence of mastitis in Mayfield Holstein-Fresian herds. At page 140 of Russell, the author points out that the c735C>G SNP, originally thought to be located in the CXCR2 gene, is actually located in the CXCR1 gene. Therefore, the IL8RB (CXCR2) SNPs disclosed in Grosse (including the c735C>G SNP) are actually SNPs of the CXCR1 gene. This thesis also shows that other SNPs in the gene are significantly associated with mastitis. For example, analysis of the singleton SNP -205C>T revealed the heterozygote CT genotype significantly associated (P<0.05) with an increased rate of mastitis (0.69 cases/cow/year) compared with the most abundant CC genotype. However, this SNP is not one disclosed in Grosse. The c816C>A SNP disclosed in Grosse (i.e. AH6-1) was also studied in the Russell thesis. Although no significant differences between genotypes were identified, the data trend suggested that heterozygous CA cows had a higher mastitis incidence (0.62 cases/cow/year), than CC and the rarer AA populations.
When reviewing the Russell thesis above, one of the references cited in Russell was Chen et al., 2011, Anim. Biotech., 22: 133-142. In the Chen paper, the authors identified SNPs in the CXCR1 gene and tested for their associations with somatic cell score (SCS - an important index of the bovine mastitis trait) in Chinese Holstein cattle. Three 5'UTR SNPs were found to be associated significantly with mastitis. A coding SNP (i.e. c783C>A - corresponding to the AH6-2 SNP identified in Grosse) also correlated significantly with SCS (AA allele showing highest susceptibility to mastitis). Therefore, this paper shows that another one of the SNPs of Grosse has been linked to mastitis in Chinese Holstein cattle.
Furthermore, in the research article by Bagheri et al., 2016, J. Applied Genet., 57: 107-112, the authors show that the c735C>G SNP is significantly associated with mastitis in Holstein herds. The authors state that the G allele is associated with resistance to mastitis (see page 110, right column).
With respect to the TNFA gene, the research article by Konnai et al., 2006, Microbes Infect., 8: 2163-2171 discloses that a SNP (-824G/A) in the 5' region of the gene (a SNP which is disclosed in Grosse as AH9-3) is associated with disease progression (i.e. progression of bovine leukemia virus-infection). The A allele appears to be associated with disease resistance.
With respect to the IFNG gene, a number of the research articles provide evidence of an association between SNPs in this gene and disease (i.e. tick) resistance. For example, Maryam et al., 2012, Mol. Biol. Rep., 39: 4565-4570 discloses the identification of 9 SNPs in the IFNG gene (3 in the coding region) and testing for an association between these SNPs and tick resistance in Fresian and Sahiwal cattle. The SNPs in this paper do not correspond to those identified in Grosse. Nevertheless, the paper shows that in the Sahiwal herd, resistance to ticks was imparted by three SNPs (P2051, P562 and G319V), and in the Fresian herd, resistance to ticks was imparted by two SNPs (P562 and P2051).
Furthermore, in a study by Abatepaulo et al., 2008, Anim. Genet., 39: 328-332 the authors analysed SNPs in the IFNG and TNFA genes (as well as SNPs in 4 other genes) for an association in Nelore (tick-resistant) and Holstein (tick-susceptible) cattle. Statistical associations were confirmed for SNPs in both of these genes with resistance to ticks. None of the SNPs in the IFNG gene or the TNFA gene are those disclosed in Grosse.
In light of the matters set out above, I consider that a number of the research articles citing Grosse have shown that a number of the genes disclosed in Grosse, and a number of the SNPs disclosed in Grosse in these genes, have subsequently been confirmed to be associated with disease resistance.
MLA’s contentions
617 MLA contends that Grosse is a study that identified 31 SNPs in genes (chemokine genes) that were thought to be involved in disease resistance based on their functional characteristics.
618 It points out that the SNPs represented markers that could subsequently be used to test for an association between each marker and infection phenotypes, such as disease resistance. It refers to Dr Sonstegard’s evidence that Grosse directed the reader to conduct an association study between the SNPs and traits of disease resistance identified in Grosse.
619 MLA further refers to Professor Hayes’ evidence to the effect that the genes described in Grosse as GRO3, GRO1 IL8R, SDF1 and TNFA were within 500 kb of a specified SNP referred to in claim 1 of the 253 Application. Accordingly, MLA contends that at least one SNP within those genes disclosed in Grosse was within 500 kb, satisfying limb (b) of claim 1. Furthermore, MLA says that the requirement that at least three SNPs be in more than one gene is satisfied given that each of the SNPs is said to be within a gene.
620 As to the requirement that the at least three SNPs be associated with a trait, MLA says that that is inherently so in light of the evidence of Dr Whitmore. MLA points out that for the IL8R gene, Grosse discloses four SNPs in IL8R, namely AH6-1, AH6-2, AH6-3 (i.e. c735C>G) and AH6-4 (at 1063, 1065 and the Tables). MLA says that Dr Whitmore was able to conclude that certain SNPs disclosed in Grosse (namely c735C>G) in that gene “are significantly associated with mastitis”, i.e. disease resistance. Then, for the TNFA gene, Dr Whitmore was able to show, by a review of one article, that it had identified an association between a SNP disclosed in Grosse for that gene and disease resistance. Finally, it was said for the IFNG gene, that a number of SNPs in that gene, though not corresponding to SNPs disclosed in Grosse, were shown to be associated with disease (tick) resistance. Generally, MLA contends that there is no requirement in claim 1 of the 253 Application that the relevant trait be something more specific than “disease resistance”. Accordingly, it contends that each of the SNPs of Grosse can be said to be associated with that trait.
621 MLA contends that the evidence discloses that the position of the IL8R and TNFA genes, and therefore the SNPs in those genes as disclosed in Grosse, fall within the scope of claim 1 because each gene lies within 500,000 bp of a specified SNP. Accordingly, it says that each SNP in those genes will be within 500,000 nucleotides of a specified SNP.
622 Finally, MLA contends that although Grosse does not expressly recommend using the method disclosed to infer a trait or sort cattle for traits, it implicitly discloses this. In any event, it says that Grosse still anticipates at least claims 1 and 6 because each only requires that the method disclosed therein be capable of such use. Now I have already disposed of that construction question.
Analysis of Grosse
623 I also reject MLA’s lack of novelty case based upon Grosse.
624 First, as is apparent, Grosse describes a SNP discovery and comparative mapping study in cattle, where the authors sequenced 7 genes from 24 animals and identified 31 SNPs distributed among those genes. But in my view Grosse does not describe an association between any of the 31 SNPs and a trait, let alone the use of at least 3 SNPs to identify a trait from a nucleic acid sample of a bovine subject.
625 Second, Professor Goddard also accepted that it was not possible to be certain that any subsequent association study with the 31 SNPs would identify any associations with any particular trait.
626 Third, Professor Goddard accepted that even if an association were identified between any one of those SNPs and a trait, that would not necessarily lead to the method of claim 1 which requires at least three SNPs that are associated with the trait.
627 Fourth, Dr Whitmore despite his searches was only able to find subsequent studies that were undertaken well after the priority date and many of them do not even cite Grosse. Further, Dr Whitmore’s evidence demonstrates that of the 3 SNPs for which a subsequent association was found, only 2 SNPs were associated with a single trait. 1 SNP (c735C>G or +777) was subsequently associated with subclinical mastitis, although the SNP was not in IL8RB/CXCR2 gene as suggested by Grosse, but in CXCR1. Another SNP disclosed in Grosse (AH6-2), also in CXCR1, mistakenly located in IL8RB/CXCR2 according to Grosse, was subsequently associated with mastitis resistance. AH9-3, a third SNP disclosed in Grosse, was subsequently shown to be associated with resistance to bovine leukaemia virus.
628 Fifth, there is no compelling evidence to establish that any of the SNPs disclosed in Grosse would be in linkage disequilibrium with any of the specified SNPs in limb (a) of claim 1 of the 253 Application.
629 Generally, the skilled person following the teachings of Grosse would not inevitably produce something falling within the relevant claims of the 253 Application. If a skilled person had done an association study between the 31 SNPs disclosed in Grosse and any particular bovine trait, the study would not inevitably have led to a method of identifying that trait by using at least 3 SNPs in at least 2 genes.
630 In summary, MLA’s lack of novelty case based upon Grosse is not clearly established in relation to claims 1, 3, 4, 5, 6, 9, 14 and 15 which all have as an integer the requirement to have 3 SNPs in more than one gene associated with a trait.
(e) The Gengler Patent
Introduction
631 The Gengler Patent contained 23 claims. It is appropriate, and only necessary, to set out claims 1 and 2 (dealing only with advantageous milk production traits), which were expressed in the following terms:
1. A method for identifying a mammal having a genotype that is indicative of advantageous milk production traits, comprising the following steps:
a) obtaining a biological sample comprising the DNA of said mammal;
b) analyzing the polymorphism of the PIT-1 and κ-casein genes of said mammal, in which the simultaneous presence of allele A and/or T of the PIT-1 gene and allele B of the κ-casein gene is indicative of high potential for milk production in said mammal.
2. A method for identifying a mammal having a genotype that is indicative of advantageous milk production traits, comprising analyzing in a biological sample from said bovine the polymorphism of the PIT-1 and κ-casein genes of said mammal, in which the simultaneous presence of allele A and/or T of the PIT-1 gene and allele B of the κ-casein gene is indicative of high potential for milk production and protein production in said mammal.
632 Clearly claims 1 and 2 embrace the use of SNPs. And clearly, there is a reference to one SNP from each of the two genes. The question is whether what is disclosed is at least 3 SNPs from 2 genes. In other words, do the claims disclose the presence of allele A and allele T of the PIT-1 gene, and allele B of the κ-casein gene? Or just either of allele A or allele T of the PIT-1 gene, and allele B of the κ-casein gene? Branhaven contends for the latter construction. MLA contends for the former construction.
633 But there is no example in the Gengler specification disclosing both alleles of the PIT-1 gene, together with allele B of the k-casein.
634 It is necessary to set out Examples 6 and 7 (at 14 to 18).
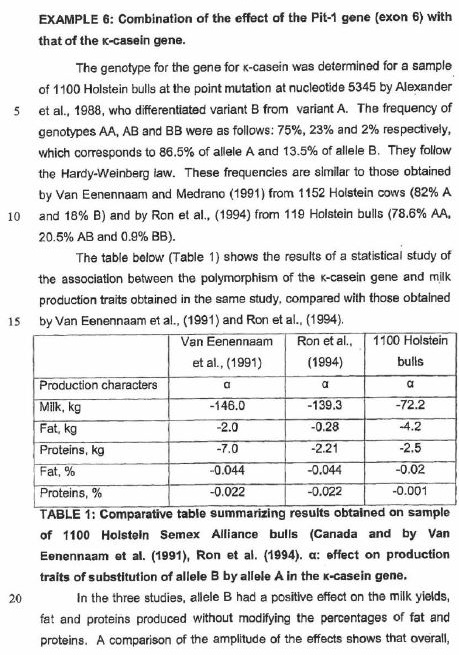
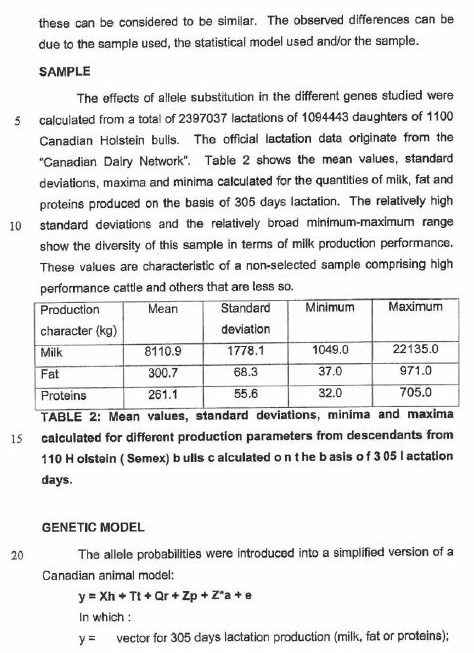

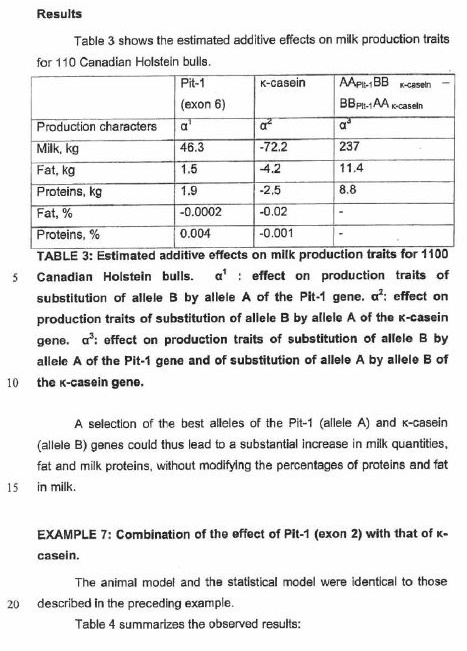
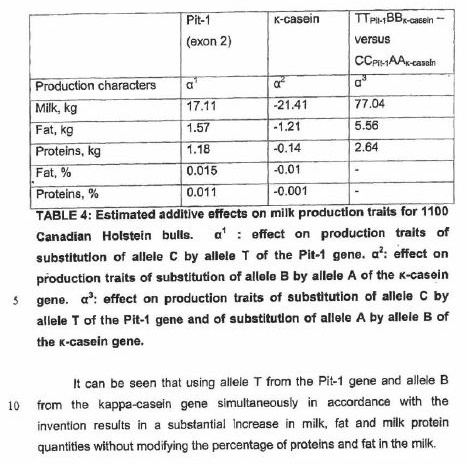
635 These do not disclose or discuss the effect of 3 SNPs.
636 Further, there is no reference to limb (b) and the question of linkage disequilibrium save that it may be accepted that it would appear that the two SNPs identified for the PIT-1 gene are in linkage disequilibrium with each other.
637 There is another matter. The following passage taken from the Gengler Patent (at 5) is not precisely set out in the French priority document:
Of course, the above steps 1 and 2 can be carried out by 2 different persons. For example, the sample can be obtained by the breeder and then sent to a laboratory that will carry out the analysis. Therefore, the present invention also concerns a method for identifying a mammal having a genotype that is indicative of advantageous milk production traits, comprising analyzing in a biological sample from said bovine the polymorphism of the PIT-1 and κ-casein genes of said mammal, in which the simultaneous presence of allele A and/or T of the PIT-1 gene and allele B of the κ-casein gene is indicative of high potential for milk production and protein production in said mammal.
638 But the latter part of the phrase is set out in the French priority document. I should also note that for completeness, the French priority document is FR0203026 filed on 11 March 2002. Claims 1 and 2 of the French priority document read:
1. Méthode d’identification d’un mammifère qui possède un genotype indicatif de traits de production laitière avantageux, comprenant les étapes suivantes:
a) obtention d'un échantillon biologique comprenant de l’ADN dudit mammifère,
b) analyse du polymorphisme des gènes PIT-1 et κ-caséine dudit mammifère, dans laquelle la présence simultanée de l’allèle A et/ou de l’allèle T de PIT-1 et de l’allèle B de la κ-caséine est indicative de potentialités élevées de production laitière et de production de protéines du lait chez ledit mammifère.
2. Méthode d’identification selon la revendication 1, dans laquelle le mammifère est un bovin.
639 On 5 June 2017, MLA provided to me a translation of the French priority document by Alexandre Gouget, a scientific and technical translator. He translated claims 1 and 2 as follows:
1. A method for identifying a mammal having a genotype that is indicative of advantageous milk production traits, comprising the following steps:
a) obtaining a biological sample comprising the DNA of said mammal;
b) analyzing the polymorphism of the PIT-1 and κ-casein genes of said mammal, wherein the simultaneous presence of allele A and/or T of the PIT-1 gene and allele B of the κ-casein gene is indicative of high potential for milk production and protein production in said mammal.
2. The identification method according to claim 1, wherein the mammal is a bovine.
640 Branhaven objected to the late translation, but in my view there is no substance to its objection. Moreover, it has had plenty of time to provide a competing translation, but has failed to do so.
MLA’s contentions
641 MLA contends that claims 1, 6, 8, 9, and 12 to 14 are anticipated by the Gengler Patent, filed 11 March 2003, but claiming priority from the French priority document (11 March 2002).
642 MLA also says that if the number of SNPs used in the claims is an arbitrary limitation, then claims 3 to 5 are also anticipated by the Gengler Patent. Accordingly, MLA seeks leave to amend its Notice of Appeal to also assert anticipation of those claims by the Gengler Patent. I will allow such an amendment. But as I have already rejected MLA’s arbitrariness submission, MLA’s case concerning claims 3 to 5 rises or falls with what I say concerning claim 1.
643 The Gengler Patent is relied upon by MLA as a “whole of contents” anticipation because, although it was not published before the earliest priority date (31 December 2002) of the 253 Application, its claims (so MLA contends) are entitled to an earlier priority date from the French priority document. Therefore, the claims of the Gengler Patent have a priority date before the earliest priority date asserted for the 253 Application.
644 Now MLA also says that the 253 Application is not entitled to claim priority from the US priority document filed on 31 December 2002, as that document does not disclose the specified SNPs claimed. MLA says that the information regarding the specified SNPs was first included in the parent application filed on 31 December 2003. And each claim of the 253 Application expressly relies on the specified SNPs as part of the invention. Therefore MLA says that the claims of the 253 Application are not entitled to claim priority before 31 December 2003: see s 43 of the Act, and regulations 3.12(1)(b) and (c), (2)(c), (2A) and (2C) of the Patents Regulations 1991 (Cth) (as they then were). It is said that if this is the case, the claims are not novel over the Gengler Patent.
645 Further, MLA also says that if the priority date is deferred to 31 December 2003, MLA can rely on the Gengler Patent as a novelty document without the need to rely on the French priority document and apply the “whole of contents” approach, as the Gengler Patent was published on 22 September 2003. The Australian copy was apparently published on 22 September 2003, but the international version was, on its face, published on 18 September 2003.
646 Before proceeding further I should say now that none of this matters given my conclusion, as I will later explain, that the Gengler Patent does not make a clear and unmistakeable disclosure of each and every integer of the relevant impugned claims of the 253 Application. But in case I need to express my views on these subjects I would say two things. First, in my view the Gengler Patent is entitled to an earlier priority date from the French priority document. Second, given that conclusion, I do not need to decide whether the 253 Application has the earlier priority date of 31 December 2002. If the Gengler Patent had made the requisite clear and unmistakeable disclosure, then it would have anticipated the 253 Application even with the earlier priority date. Now the 253 Application may have had the earlier priority date, but if it does not it certainly has the deferred priority date of 31 December 2003; but for present purposes I do not need to decide between the two. For completeness, I would reject the other MLA contention of a 17 October 2013 priority date if it is persisted with.
Claims 1 and 3 to 5
647 MLA says that the Gengler Patent is directed to the development of methods for genotyping animals for DNA-based markers in two candidate genes for milk production, and their use in identifying (selecting) animals with desirable phenotypes for various milk-production traits. At 2, lines 27 to 29, the Gengler Patent states: “[t]he aim of the invention is to provide a method for identifying a mammal having a genotype indicative of advantageous milk production traits.” The Gengler Patent specifically refers to cattle (Examples 5 and 6).
648 MLA relies upon Professor Visscher’s explanation that:
[t]he DNA-based markers referred to in Gengler comprise a previously-known polymorphism (with alleles A and B) in the κ-casein gene, a previously-known polymorphism (with alleles A and B) in exon 6 of the pituitary transcription factor 1 (PIT-1) gene, and another polymorphism (with alleles C and T) in exon 2 of the PIT-1 gene that appears to be novel to this document.
649 Professor Visscher also said that all three of the polymorphisms disclosed in the Gengler Patent are SNPs.
650 MLA points to the fact that the authors of the Gengler Patent stress that the novelty of their invention lies in selecting animals based on combined consideration of their genotypes in both the K-Cas and PIT-1 genes. For example, at 3, lines 24 to 26 it is stated that: “the method is more effective than known prior art methods based on the use of the PIT-1 marker alone or on the kappa-casein marker alone”.
651 MLA also relies upon Professor Visscher’s explanation that the Gengler Patent:
discloses a method to “identify a trait in a bovine subject” from 3 SNPs in 2 genes [i.e. A and T alleles in PIT-1, and B allele in kappa-casein] that are associated with the trait as described by claim 1 of the [253 Application].
652 Further, as to Branhaven’s assertion that the Gengler Patent does not teach the use of all three SNPs together, MLA points to claim 1 of the Gengler Patent which claims:
A method for identifying a mammal having a genotype that is indicative of advantageous milk production traits, comprising the following steps:
(a) obtaining a biological sample comprising the DNA of said mammal;
(b) analyzing the polymorphism of the PIT-1 and κ-casein genes of said mammal, in which the simultaneous presence of allele A and/or T of the PIT-1 gene and allele B of the κ-casein gene is indicative of high potential for milk production and protein production in said mammal. [emphasis added]
653 MLA says that the abstract, claim 2 and the body of the Gengler Patent (at 4, lines 1 to 5), are all in similar form. That is, they all disclose genotyping animals for all three SNPs (i.e. 2 SNPs in the PIT-1 gene and 1 SNP in the κ-casein gene) satisfying the “at least three SNPs in more than one gene” requirement of claim 1 of the 253 Application. MLA says that it does not matter that the examples in the Gengler Patent do not disclose use of all three SNPs together since they are merely experiments leading to the invention of all three SNPs and, in any event, the disclosure is not limited to the experiments that are performed.
654 Now MLA refers to the fact that Professor Plastow originally gave evidence that there was no point in using a SNP which was in high LD with another SNP but that he conceded that the Gengler Patent is drafted to cover the use of 3 SNPs together. Further, Professor Visscher explained why that would be useful and it is said that Professor Plastow did not refute that. Accordingly, MLA says that in the circumstances, not only does the Gengler Patent direct the use of three SNPs at once to infer a trait, but that it would be understood by a person skilled in the art as logical to do so.
655 Further, MLA contends that on the basis of the information provided by Professor Hayes, the PIT-1 and K-Cas genes on the bovine genome fall within 500,000 bp of specified SNPs of the 253 Application. In particular, based on the mapping results presented by Professor Hayes, the SNP disclosed in SEQ ID No: 20015 of the 253 Application (“SNP20015”) is located at base 35,186,040 of chromosome 1, and therefore the entire PIT-1 gene lies within 500 kb of SNP20015. Given that the two SNPs in PIT-1 are located in the gene (one in exon 6 and the other in exon 2), the two SNPs must also lie within 500 kb of SNP20015. Furthermore, the SNP disclosed in SEQ ID No: 20139 of the 253 Application (“SNP20139”) is located at base 87,471,038 of chromosome 6, and therefore the κ-casein gene lies within 500 kb of SNP20139. Given that the SNP in κ-casein is located in the gene (at nucleotide position 5345), the SNP must also lie within 500 kb of SNP20139. Accordingly, MLA says that the Gengler Patent claims the features of claim 1 of the 253 Application, with an earlier priority date. Further, MLA contends that if the number of SNPs used is an arbitrary parameter, then claims 3 to 5 are also anticipated. As I have said, I have rejected that submission of arbitrariness.
Claim 6
656 With respect to claim 6 of the 253 Application, MLA contends that by carrying out the method claimed in claim 1 of the Gengler Patent, animals are identified as having advantageous milk production qualities. That implicitly provides a method for sorting of those animals. In any event, it is said that the method in the Gengler Patent is suitable for that purpose.
Claims 8 and 12
657 With respect to claims 8 and 12 of the 253 application, the Gengler Patent states at 7, lines 23 to 24 that “… other techniques can be used to determine polymorphism in the genes under consideration …”. MLA contends that these other techniques would be understood by the skilled addressee to include standard nucleic acid hybridisation as set out in claims 8 and 12 of the 253 Application, as Professor Visscher explained.
Claim 9
658 Claim 1 of the Gengler Patent concerns the “high potential for milk production and protein production”. MLA contends that this falls within the list of traits in claim 9 of the 253 Application (e.g. milk production and milk quality).
Claim 14
659 As to claim 14, MLA contends that the methods disclosed in the Gengler Patent for identifying the SNP in the K-Cas gene (Example 1) and the SNP in exon 6 of the PIT-1 gene (Example 3) each involve isolation of genomic DNA and PCR amplification of a fragment of DNA containing the SNP.
Analysis of the Gengler Patent
660 I would also reject MLA’s lack of novelty case concerning the Gengler Patent.
661 In my view, even if the Gengler Patent forms part of the relevant prior art base, it does not anticipate.
662 The Gengler Patent discloses a candidate gene approach to identify animals possessing certain milk production traits; I accept though that the claims of the 253 Application are not limited by the approach taken to get to the 3 SNPs. There are two candidate genes disclosed, PIT-1 and K-Cas. The Gengler Patent describes 2 different SNPs in the PIT-1 gene and 1 SNP in the K-Cas gene. The authors test the effect of the K-Cas SNP alone, or in combination with either the exon 2 SNP of PIT-1 or the exon 6 SNP of PIT-1.
663 The method of the invention is described in terms of a single method using both the PIT-1 marker and the K-Cas marker:
[T]he inventors have developed a method for identifying a mammal that has a genotype indicative of advantageous milk production traits, in which both the PIT-1 gene and the kappa-casein gene are used as genetic markers. They have shown that this method is more effective than known prior art methods based on the use of the PIT-1 marker alone or on the kappa-casein marker alone.
664 As to exemplification, the inventors show in example 6 that one PIT-1 SNP combined with the K-Cas SNP gives a better result than either SNP alone. Further, in example 7, they show that the other PIT-1 SNP combined with the K-Cas SNP gives a better result than either SNP alone. But only in the relevant claims and the consistory clauses do the inventors use the term “and/or” to encompass use of one or other or both of the SNPs in the PIT-1 gene together with the SNP in the K-Cas gene.
665 First, I agree with Branhaven that the inventors are not disclosing 2 separate methods to identify certain milk production qualities, but a single method that is capable of being carried out using 2 SNPs (1 of the 2 PIT-1 SNPs and the K-Cas SNP) following all the examples in the Gengler Patent.
666 Now I would note that in relation to the evidence before me, Professor Goddard accepted that in the examples in the Gengler Patent, the inventors only carry out the method with 2 SNPs, one or other of the PIT-1 SNPs and the K-Cas SNP. He accepted that the inventors do not exemplify the method using both PIT-1 SNPs.
667 Indeed, Professor Taylor explained that the reason that only one of the two SNPs in the PIT-1 gene is used is because the PIT-1 SNPs are in tight linkage disequilibrium with each other. Professor Plastow also explained that because it was well apparent from the disclosure in the Gengler Patent that the 2 SNPs in the PIT-1 gene were in linkage disequilibrium, one would only use one or the other. Dr Sonstegard also confirmed that because the two PIT-1 SNPs are so tightly linked, there would be no reason to test for both.
668 Now I accept that Professor Visscher gave evidence that “it would not always be the case that the one [SNP] at exon 6 predicts perfectly the one on exon 2”, so you would get some extra information from using both the PIT-1 SNPs. But the inventors for the Gengler Patent apparently did not think this necessary. Further, I agree with Branhaven that even if Professor Visscher’s evidence is accepted, it only rises as high as supporting the proposition that there was an alternative way of carrying out the method to the way the method was carried out in the examples in the Gengler Patent, which was to use 3 SNPs instead of 2. In other words, using 3 SNPs is only one option in carrying out the method disclosed in the Gengler Patent. Alternatively expressed, the Gengler Patent contains a direction which is at least as likely to be carried out using 2 SNPs as it would be using 3 SNPs. Indeed, the evidence of Professor Plastow, Professor Taylor and Dr Sonstegard supports the conclusion that the method of the Gengler Patent would be more likely to be carried out using 2 SNPs, rather than 3 SNPs.
669 Second, the Gengler Patent does not disclose nor is compelling evidence given of an implicit disclosure that any of the 3 SNPs disclosed in the Gengler Patent are in linkage disequilibrium with any of the 2,510 SNPs of the 253 Application.
670 In summary, in my view there is not a clear and unmistakable direction to use 3 SNPs, an integer in claim 1 of the 253 Application. For at least this reason, the Gengler Patent is not an anticipation of the relevant claims. It is not inevitable that a skilled person following the Gengler Patent would necessarily follow a method that would infringe claim 1 of the 253 Application. A similar conclusion follows concerning claims 3 to 5, 6, 8, 9 and 12 to 14 which have the 3 SNPs (or indeed greater number) requirement.
671 Finally, given my conclusion I do not need to resolve priority date questions as I have said.
(f) Other matters
672 At one stage, MLA relied upon the Gelhaus publication (Gelhaus A and Förster B, “Cattle MHC Genes DOA and DOB: Sequence Polymorphisms and Assignments to the Class IIb Region” (2001) 28 European Journal of Immunogenetics 429), but this now appears to have been abandoned on this aspect at least. Nevertheless as my attention was drawn to it, it is convenient to dispose of it.
673 Gelhaus reported on a study of the genetic polymorphism of second exons of the cattle DOA and DOB genes. As it stated, two and four allelic variants were detected. The SNPs detected enabled the relevant loci to be used as markers in trait analysis. The context was a consideration of genetic variants that might influence disease resistance in cattle; unsurprisingly therefore, Gelhaus was reported in the European Journal of Immunogenetics.
674 The conclusion of Gelhaus was distinctly stated at 432 in the following terms:
The polymorphisms detected in the BoLA-DOA and -DOB genes allow these loci to be used as markers in genetic studies on disease resistance and related traits. Assuming functionality as in sheep, mouse and human, further characterization of the gene products is likely to contribute to a better understanding of the genetic control of immune responses in cattle.
675 Now no association study had been carried out although clearly Gelhaus was teaching that one could test a SNP(s) with an association study.
676 But there is no express disclosure of the method of claim 1 of the 253 Application or limb (a) or limb (b) thereof.
(g) Conclusion
677 MLA’s ground of opposition concerning a lack of novelty fails. Its arguments did not even approach satisfying the high threshold for establishing such a ground on this appeal, as I discussed at the outset of my reasons.
Lack of inventive step
(a) Some legal principles
678 The question is whether the claimed invention lacks an inventive step over the prior art base. An invention is taken to involve an inventive step when compared to the prior art base unless it would have been obvious to a person skilled in the relevant art in light of common general knowledge as described in Minnesota Mining and Manufacturing Co v Beiersdorf (Australia) Ltd (1980) 144 CLR 253 at 292 per Aickin J (Minnesota Mining) as it existed in the patent area (the then s 7(2) of the Act) before the priority date, whether that knowledge is considered separately or together with information of the kind described in the then s 7(3).
679 For the present context, the applicable provisions of ss 7(2) and (3) are the following:
(2) For the purposes of this Act, an invention is to be taken to involve an inventive step when compared with the prior art base unless the invention would have been obvious to a person skilled in the relevant art in the light of the common general knowledge as it existed in the patent area before the priority date of the relevant claim, whether that knowledge is considered separately or together with the information mentioned in subsection (3).
(3) The information for the purposes of subsection (2) is:
(a) any single piece of prior art information; or
(b) a combination of any 2 or more pieces of prior art information;
being information that the skilled person mentioned in subsection (2) could, before the priority date of the relevant claim, be reasonably expected to have ascertained, understood, regarded as relevant and, in the case of information mentioned in paragraph (b), combined as mentioned in that paragraph.
680 The term “obvious” means “very plain” (Aktiebolaget Hässle v Alphapharm Pty Ltd (2002) 212 CLR 411 at [34] per Gleeson CJ, Gaudron, Gummow and Hayne JJ (Aktiebolaget Hässle) and Lockwood (No 2) at [51]). The inventive element needed to sustain a patent can be small. A “scintilla of inventiveness” will be sufficient and “no smallness or simplicity will prevent a patent being good” (Meyers Taylor Pty Ltd v Vicarr Industries Ltd (1977) 137 CLR 228 at 249 per Aickin J). Relevant further content has been given to determining obviousness in The Wellcome Foundation Ltd v VR Laboratories (Aust) Pty Ltd (1981) 148 CLR 262 at 286 per Aickin J in stating:
whether the hypothetical addressee faced with the same problem would have taken as a matter of routine whatever steps might have led from the prior art to the invention, whether they be the steps of the inventor or not.
681 Further, in relation to experiments, his Honour said (at 280 and 281):
In the present case it was admitted by the respondent that the test of obviousness was an objective one, but it was argued that evidence of a subjective character was admissible. That is no doubt true in some cases because expert witnesses are often properly asked whether they found a particular invention “surprising” to them. That however does not answer the question whether evidence of the steps which the patentee took is relevant and therefore admissible. Evidence of what was in the patentee’s mind may be admissible as evidence of the state of the art, but could seldom be otherwise admissible. Evidence of what he did by way of experiment may be another matter. It might show that the experiments devised for the purpose were part of an inventive step. Alternatively it might show that the experiments were of a routine character which the uninventive worker in the field would try as a matter of course. The latter could be relevant though not decisive in every case. It may be that the perception of the true nature of the problem was the inventive step which, once taken, revealed that straightforward experiments will provide the solution. It will always be necessary to distinguish between experiments leading to an invention and subsequent experiments for checking and testing the product or process the subject of the invention. The latter would not be material to obviousness but might be material to the question of utility.
682 The question to be posed was whether putative experiments leading from the prior art to the invention as claimed were part of the inventive step or were of a routine character to be tried as a matter of course. That question has an affinity with the Cripps question posed in Olin Mathieson Chemical Corporation v Biorex Laboratories Ltd [1970] RPC 157 and paraphrased by French CJ in AstraZeneca AB v Apotex Pty Ltd (2015) 257 CLR 356 at [15] (AstraZeneca) in the following terms:
Would the notional research group at the relevant date, in all the circumstances, which include a knowledge of all the relevant prior art and of the facts of the nature and success of [the existing compound], directly be led as a matter of course to try [the claimed inventive step] in the expectation that it might well produce a useful alternative to or better drug than [the existing compound]?
683 Now the question does not import as a criterion that the inventive step claimed would be perceived by the hypothetical addressee as “worth a try” or “obvious to try” (AstraZeneca per French CJ at [15]). Further, Aktiebolaget Hässle (at [72]) rejected the adoption of a criterion expressed in terms of “obvious to try” or “worth a try”.
684 Let me say something further about common general knowledge.
685 Section 7(2) first requires consideration of what would have been obvious to a person skilled in the relevant art in the light of the common general knowledge as it existed before the priority date, putting to one side for the moment s 7(3) information. Common general knowledge is knowledge “generally known and accepted without question by the bulk of those who are engaged in the particular art” (British Acoustic Films Ld v Nettlefold Productions (1936) 53 RPC 221 at 250 per Luxmoore J). Information cannot be treated as part of the common general knowledge unless there is evidence of its general acceptance and assimilation by persons skilled in the art. Information does not constitute common general knowledge merely because it might be found, for example, in a journal, even if widely read by such persons. Further, as Luxmoore J said at 250:
In my judgment it is not sufficient to prove common general knowledge that a particular disclosure is made in an article, or series of articles, in a scientific journal, no matter how wide the circulation of that journal may be, in the absence of any evidence that the disclosure is accepted generally by those who are engaged in the art to which the disclosure relates. A piece of particular knowledge as disclosed in a scientific paper does not become common general knowledge merely because it is widely read, and still less because it is widely circulated. Such a piece of knowledge only becomes general knowledge when it is generally known and accepted without question by the bulk of those who are engaged in the particular art; in other words, when it becomes part of their common stock of knowledge relating to the art. Whatever else common general knowledge may be, it has never in my judgment included public knowledge of particular documents reports or scientific papers and the like. The knowledge of a number of individuals that a particular suggestion or particular suggestions has or have been made for the use of biasing in a particular apparatus, or a number of particular apparatus, cannot be held to be common general knowledge. It is certainly difficult to appreciate how the use of something which has in fact never been used in a particular art can ever be held to be common general knowledge in the art.
686 Further, and as stated in Minnesota Mining at 293 per Aickin J, the notion of common general knowledge:
involves the use of that which is known or used by those in the relevant trade. It forms the background knowledge and experience which is available to all in the trade in considering the making of new products, or the making of improvements in old, and it must be treated as being used by an individual as a general body of knowledge.
So it must be knowledge that is known and available to all in the trade or at least the bulk of those who are engaged in the relevant art. Accordingly, information ascertainable by a routine literature search is not of itself taken to be common general knowledge. And patent specifications do not form part of common general knowledge without evidence that they have been absorbed into common general knowledge.
687 And as further elucidated by Jagot J in Gilead Sciences Pty Ltd v Idenix Pharmaceuticals LLC (2016) 117 IPR 252; [2016] FCA 169 at [216] (affirmed on appeal in Idenix Pharmaceuticals LLC v Gilead Sciences Pty Ltd [2017] FCAFC 196 per Nicholas, Beach and Burley JJ), it is erroneous to treat a document as being part of common general knowledge simply because skilled persons could readily locate and assimilate its contents. Her Honour went on to explain (at [217]):
It may be accepted that instant recall of an article is not required. This does not mean, however, that documents found by searching for a subject-matter, rather than by some form of recall or reminder of what is already known to exist, are common general knowledge. This is so irrespective of the fact that experts in the field read widely. Further, it is not the case that mere publication and republication proves that a document and its contents have entered the common general knowledge. Nor is it the fact that a document and its contents necessarily form part of the common general knowledge merely because one expert knows or has managed to locate it and assimilate its contents. Such a document may or may not form part of the common general knowledge. The relevant inferences are to be drawn on the basis of the whole of the evidence.
688 There is also a further point to be made concerning common general knowledge. There is no general principle permitting admissions in the specification of a patent to be used to establish in and of themselves that information is common general knowledge. Whether information has become so widely assimilated that it forms part of common general knowledge must be determined on the evidence, although admissions can be considered as part of that evidence. But I would note in this context that [0211] of the 253 Application provided that:
Reference to any prior art in the specification is not, and should not be taken as, an acknowledgment, or any form of suggestion, that this prior art forms part of the common general knowledge in Australia or any other jurisdiction or that this prior art could reasonably be expected to be ascertained, understood and regarded as relevant by a person skilled in the art.
689 Now in addition to using common general knowledge on a stand-alone basis, common general knowledge can be aggregated with s 7(3) information. I have already set out s 7(3) in its then applicable form. That part of the prior art base which is common general knowledge and the information referred to in s 7(3) are considered for the purpose of looking forward from the prior art base to see what the skilled person is likely to have done when faced with a particular problem. In a case where the problem is known and is part of the common general knowledge, the problem may be similar to that which the patentee claims to have solved with the claimed invention. But where the problem addressed by the patentee does not form part of common general knowledge, the relevant starting point is the prior art base, but not including the problem as identified in the patent specification. As Besanko, Foster, Nicholas and Yates JJ noted in AstraZeneca AB v Apotex Pty Ltd (2014) 226 FCR 324 at [203]:
If the problem addressed by a patent specification is itself common general knowledge, or if knowledge of the problem is s 7(3) information, then such knowledge or information will be attributed to the hypothetical person skilled in the art for the purpose of assessing obviousness. But if the problem cannot be attributed to the hypothetical person skilled in the art in either of these ways then it is not permissible to attribute a knowledge of the problem on the basis of the inventor’s “starting point” such as might be gleaned from a reading of the complete specification as a whole.
690 The purpose of the inquiry is to determine whether the invention is obvious to solve the perceived problem, looking forward from the prior art base. But of course this may not have been the patentee’s starting point.
691 Now impermissible hindsight should be avoided in determining whether a claimed invention lacks an inventive step. The plurality in Aktiebolaget Hässle said (at [21]):
The defendant to an infringement action who cross-claims for revocation on the ground of obviousness bears the onus of establishing that case. This obliges the defendant to lead evidence looking back to the priority date, sometimes, as here, many years before trial. In those circumstances, the warnings in the authorities against the misuse of hindsight are not to be repeated as but prefatory averments and statements of trite law. The danger of such misuse will be particularly acute where what is claimed is a new and inventive combination for the interaction of integers, some or all of which are known. It is worth repeating what was said by Lord Diplock in Technograph Printed Circuits Ltd v Mills & Rockley (Electronics) Ltd [[1972] RPC 346 at 362] …
692 Further, for a combination invention, the question is whether the combination, not each integer, is obvious. It is simply impermissible to take any one integer or take each integer seriatim and ask whether each integer involved an inventive step. The invention is the combination. The combination is what is claimed as the monopoly. And the relevant question is whether the combination involves an inventive step (see Lockwood (No 2) at [69]). As the plurality in Aktiebolaget Hässle (at [41]) said:
… The claim is for a combination, the interaction between the integers of which is the essential requirement for the presence of an inventive step. It is the selection of the integers out of “perhaps many possibilities” which must be shown by Alphapharm to be obvious, bearing in mind that the selection of the integers in which the invention lies can be expected to be a process necessarily involving rejection of other possible integers. This expression of the issue follows what was said by Aickin J in Minnesota Mining. (citation omitted)
693 Further, the misuse of hindsight is most common in relation to combination claims (Minnesota Mining at 293 per Aickin J).
694 Further as to combination claims, Lord Davey stated in In the matter of Klaber’s Patent (1906) 23 RPC 461 at 469 (in terms approved by Dixon J in Palmer v Dunlop Perdriau Rubber Co Ltd (1937) 59 CLR 30 at 73):
A proper combination for a patent is the union of two or more integers, every one of which elements may be perfectly old, for the production of one object which is either new, or at any rate is for effecting an old object in a more convenient, cheaper or more useful way. But the point in a combination patent must always be that the elements of which the combination is composed are combined together so as to produce one result.
(b) Common general knowledge
695 MLA contends, which I of course accept, that common general knowledge constitutes part of the mental equipment of those concerned in the art under consideration being the background knowledge and experience which is available to all in the trade in considering the making of new products, or the making of improvements in old.
696 MLA further contends that where the specification itself describes matters as being known, I should proceed on the basis that such an admission is correct (see Bristol-Myers Squibb Co v F H Faulding & Co Ltd (2000) 97 FCR 524 at [30]) and indeed accept that to be common general knowledge. That submission overreaches. I would make two brief observations consistently with what I have already intimated. First, for the patentee to say that something is known as at the priority date does not necessarily entail that it was common general knowledge in the context of applying the statutory concept in s 7(2). Second, if an admission does arise, it is only part of the evidence for me to consider, attributing to the admission what I consider to be its appropriate weight. Statements in a specification cannot be definitive of the question.
697 MLA contends that the following eight matters were common general knowledge as at the earliest priority date of 31 December 2002 (and deferred priority date of 31 December 2003) of the claims of the 253 Application. It is convenient to deal with each matter and my response in the order advanced by MLA. As I have previously stated, where I use the expression “priority date”, it is to be taken as a reference to either the earliest priority date or the deferred priority date unless I state otherwise.
Need for improved methods for predicting genetic potential in animals
698 MLA says that there was a need for improved methods for predicting genetic potential in animals for numerous quantitative traits which were known to be desirable. Further, it points to the fact that the specification states at [0022] that “three different experimental approaches have been used with limited success to identify genes, chromosomal regions or DNA markers that account for a large proportion of the genetic variation observed in economically important traits in livestock species” and that the results “have not been widely utilized to date because they do not account for enough of the total genetic variation to allow accurate prediction of an animal’s performance for a specific trait.” MLA says that this accords with the evidence of the experts that the approaches at the time were of limited value for predicting an animal’s performance with respect to the majority of quantitative traits.
699 MLA says that by the late 1990s/early 2000s, there was a lot of interest in using molecular genomics in cattle breeding and that was where research and the industry was heading. It contends that ultimately the evidence of all experts was that before December 2002, DNA markers “were the way to go”.
700 MLA also says that in the review published in February 2001 of Andersson L, “Genetic Dissection of Phenotypic Diversity in Farm Animals” (2001) Nat Rev Genet 130 (cited in Wiggans GR, Cole JB, Hubbard SM and Sonstegard TS, “Genomic Selection in Dairy Cattle: The USDA Experience” (2017) 5 Annu Rev Anim Biosci 309), the author said that “[i]t is likely that large-scale marker analysis will be used routinely, as soon as the cost for genotyping has been reduced by a factor of around ten” (at 136 and 137, second column, second last line). The review also said that “[l]inkage disequilibrium mapping will be a very powerful approach for mapping and finding trait loci in domestic animals once dense SNP maps become available and the cost for genotyping has been reduced such that genome scans using thousands of SNPs can be done” (at 137, first column, second full para). The review also observed that “[i]t is also only a matter of time before initiatives will be taken to sequence the genomes of farm animals. This will most probably be carried out using a whole genome shotgun approach …” (at 137, second column, first full para). MLA says that this review was generally consistent with the views expressed by Dr Sonstegard in his 2001 review of dairy cattle genomics (Sonstegard TS, Van Tassell CP and Ashwell MS, “Dairy Cattle Genomics: Tools to Accelerate Genetic Improvement?” (2001) 79 J Anim Sci E307).
701 Further, MLA points to the fact that prior to the priority date, Professor Taylor had set up Genomic FX to do something which was similar to what is described in the 253 Application. It asserts that his company’s aim was to develop a simple diagnostic test for genotyping animals for the purpose of improving their genetic potential. His plan was to scan the genomes of thousands of animals and to identify SNPs evenly spaced throughout the genome and find SNPs that correlated with useful traits. It is said that his company did not proceed with the plan because it was forced to “shut its doors”. MLA says that the only difference between the work of Genomic FX and the 253 Application that Professor Taylor could identify was that he was looking at within family QTLs, not the whole bovine genome. But MLA says that the only difference between the within family QTL approach and the genomic selection approach is that GS looks at markers across the whole genome, whereas the within family QTL approach only looks at segments of the genome. Otherwise, so MLA contends, Professor Taylor’s proposed approach before the priority date was the same as that of the 253 Application.
702 I would make the following observations in response to these contentions.
703 It is not in dispute that the state of the art as at the priority date was that there were a variety of methods being developed and used to determine the genetic potential of bovines for economic traits, including progeny testing, the candidate gene approach and QTL mapping.
704 It is also not in dispute that as at the priority date, traditional phenotypic selection techniques such as progeny testing were commonly used in practice and that those techniques were sophisticated and being continuously improved. Further, as at the priority date there was limited use of molecular genetic techniques. And at that time the laboratory use of molecular genetic techniques was directed more to the candidate gene approach and the within family QTL mapping approach.
705 As I have previously summarised, the candidate gene approach is based on knowledge of the gene’s biological function and its relationship to the trait of interest. It is typically based on knowledge of analogous genes in other species. As Dr Sonstegard said, it was a popular approach amongst researchers at the time to identify genes responsible for quantitative traits. Further, as I have already previously summarised, QTL mapping typically involved genotyping and trait-association experiments conducted within families to identify particular chromosomal regions associated with a trait of interest. Further, Dr Sonstegard also gave evidence that when the candidate gene approach was combined with QTL mapping, the approach was referred to as a “positional candidate gene approach”. The search for candidate genes was limited to a specific region of the genome that was known to be associated with a trait (i.e. the QTL).
706 It is also not in dispute that as at the priority date neither QTL mapping nor the candidate gene approach involved the use of genome wide dense markers. Such approaches were rather focused on identifying the location of a causative gene(s) within a specific region of a chromosome.
707 Now what I have just said was reflected in the research work at the time, indeed even after the priority date, being conducted by the experts who were called before me. Professor Goddard’s publication closest to the priority date involved the fine mapping of a QTL locus for twinning rate in cattle which had all bulls genotyped for 15 markers located on a specific region of chromosome 5, but did not disclose the construction or use of a dense genome-wide panel of markers or map (see Meuwissen T, Karlsen A, Lien A, Olsaker I and Goddard ME “Fine Mapping of a Quantitative Trait Locus for Twinning Rate Using Combined Linkage and Linkage Disequilibrium Mapping” (2002) 161 Genetics 373 (accepted for publication on 11 February 2002 and published in May 2002)). Further, Professor Visscher’s work in bovines until at least 2005 predominantly involved family-based linkage studies using microsatellite markers. Further, Dr Sonstegard was involved in the fine-mapping and sequencing of a QTL on bovine chromosome 6 and identified osteopontin as a candidate gene affecting milk production in or around 2005, which research was published in the Proceedings of the National Academy of Sciences in 2005 (Schnabel RD, Kim JJ, Ashwell MS, Sonstegard TS, Van Tassell CP, Connor EE and Taylor JF, “Fine-Mapping Milk Production Quantitative Trait Loci on BTA6: Analysis of the Bovine Osteopontin Gene” (2005) 102 PNAS 6896). Further, in evidence before me reference was made to a well-known study in 2001 in which the positional candidate gene approach was successfully used to identify a causative polymorphism in a gene called DGAT1 which accounted for about 30% of the variation in particular milk traits (Grisart B, Coppieters W, Farnir F, Karim L, Ford C, Berzi P, Cambisano N, Mni M, Reid S, Simon P, Spelman R, Georges M and Snell R, “Positional Candidate Cloning of a QTL in Dairy Cattle: Identification of a Missense Mutation in the Bovine DGAT1 Gene with Major Effect on Milk Yield and Consumption” (2001) 12 Genome Research 222). Further, Professor Plastow gave evidence about his use of the candidate gene approach on the melanocortin receptor 4 gene (MC4R), published in 2000 (Kim KS, Larsen N, Short T, Plastow G and Rothschild MF, “A Missense Variant of the Porcine Melanocortin-4 Receptor (MC4R) Gene is Associated with Fatness, Growth and Feed Intake Traits” (2000) 11 Mamm Genome 131). The MC4R candidate gene explained variation in a polygenic trait, being growth and feed intake. And according to Professor Plastow, although this gene explained a very small amount of the variation in the trait, the ability to identify variants through the use of markers was commercially very valuable. Further, I note for completeness that Professor Plastow later discussed in a 2005 article the MC4R gene, the use of functional genomics studies to identify candidate genes and also high-density marker approaches (described as Phase 3), none of which was inconsistent with his evidence before me as to what was relevantly known or expected as at the priority date (van der Steen, H, Prall G and Plastow GS “Application of Genomics to the Pork Industry” (2005) J Anim Sci 83:E1).
708 Now MLA has made reference to Professor Taylor’s involvement in Genomic FX prior to the priority date. But the described objective of Genomic FX was to use within family QTL mapping with fine marker mapping to identify predictive variants or positional cloning, rather than to scan the genomes of thousands of animals and to identify SNPs evenly spaced throughout the genome. And in any event such work did not proceed. Now I accept that positional cloning was one of the methods being used in laboratories before the priority date, but this method was not a whole of genome approach.
709 Further, the evidence of Professor Taylor and Dr Sonstegard was that although there was interest in genetic markers, progeny testing was still a very common technique in the industry as at the priority date.
710 Further, and as I have endeavoured to explain previously, in contrast to the candidate gene and QTL mapping approaches, a genome wide approach does not require identifying any particular causal gene or region of the chromosome. And the possibility of using genome wide approaches in livestock breeding was theoretical as at the priority date, with it being unconfirmed whether such approaches could successfully be applied in cattle. Further, the tools required such as a genome-wide marker map, were not available. And the absence of a sufficient number of markers was a technical hurdle. Further, it was speculation whether a SNP chip would become available.
711 As at the priority date, to apply a genome wide approach it would have been necessary to identify a large number of genetic markers evenly spaced throughout the genome. And to design and carry out the necessary experiments would have presented many challenges, particularly when as at the priority date, work on sequencing the whole bovine genome had yet to commence. As Dr Sonstegard gave evidence of, which I accept, the challenges included the need to sequence and assemble a large proportion of the bovine genome, the need to identify a large number of informative markers, the need to determine the relative position of those markers, the need to generate a genome-wide panel of relatively evenly spaced markers, and the need to identify those markers that are associated with a trait. I would just interpolate at this point that MLA spent a considerable time attacking Dr Sonstegard’s evidence by reference to observations made in articles that he had co-authored which discussed in part genomic selection and the problems involved and resources required. But I do not consider that MLA established at all that Dr Sonstegard’s evidence was not reliable on what was known or expected as at the priority date on such matters.
712 Further, on the evidence before me, it is not seriously in doubt that as at the priority date, the relevant work was technically difficult, expensive and time consuming. The then existing techniques were inadequate. They did not provide the required specificity for SNP identification, and certainly not on a large scale. Further, SNP chips were not developed until the mid to late 2000s. I would note that they were not used by the inventors of the 253 Application. Further, as Professor Plastow explained, the necessary high throughput assay technology to screen multiple bovine markers was not developed until well after the priority date.
713 Further, the preponderance of the evidence also established that as at the priority date, microsatellites were the most widely used genetic marker in bovines. They had various advantages over other markers, including SNPs, as various of the experts accepted. Further, fragment analysers only became available in the early 2000s to enable the rate at which microsatellites could be genotyped to be substantially increased. Contrastingly, SNPs were seldom used as a genetic marker, and as at the priority date very few had been discovered in cattle. Moreover, it was only after the priority date that a significant transition towards the use of SNP markers occurred when the BovineSNP50 SNP chip was commercially released.
714 Further, according to Professor Plastow, even by 2005 the consensus was that there were still significant hurdles to be overcome before a genome wide approach could be utilised in the breeding selection and management of cattle.
715 Further, I would also note that Professor Visscher for MLA accepted that the main practical limitation to implementing a genomic selection approach as at December 2002 was the identification of a sufficient number of markers. Now although Professor Visscher said that there was no technical difficulty at the time in assaying or genotyping each individual marker, that is different to what was considered to be the main practical limitations that I have outlined. In any event, this is a reference to genotyping individual markers, rather than a genome-wide panel of markers. Now Professor Visscher accepted that the information in Vignal (Vignal A, Milan D, San Cristobal M and Eggen A, “A Review on SNP and Other Types of Molecular Markers and Their Use in Animal Genetics” (2002) 34 Genet Sel Evol 275) was consistent with his understanding of what was known prior to December 2002 about the use of genetic markers in animal genetics. But in my view Vignal does not assist MLA. Vignal does not describe the use of genomic selection as the direction of research in 2002. And I note that it does not cite the Meuwissen paper; MLA asserted that Meuwissen was well known at the time. Vignal does not suggest producing a genome wide map of evenly spaced markers. And it does not suggest how such a map could be constructed and applied. Further, Professor Visscher accepted the reference to “high densities of markers” being needed was in the context of a fine QTL mapping approach. It is well apparent that the expressed desire for more markers within the region of a QTL is different from a genome wide dense marker map necessary to carry out a GWAS or genomic selection. The purpose of identifying more markers for a fine QTL mapping approach is to more accurately locate the causal gene. But contrastingly, the GWAS and genomic selection approaches do not involve determining the identity or location of any causal mutation.
716 Further, what is apparent from Vignal, consistently with the expert evidence led before me, is that at the priority date, to the extent that genetic markers were being used, they were microsatellites. Now it might have been accepted that in the future SNPs might produce equivalent information, but that uncertainty does not suggest any expectation that SNPs could be used successfully (as claimed in the 253 Application). It is convenient to set out some extracts from Vignal.
717 Vignal, which was published in 2002 (after 8 March 2002 (the date of acceptance)), stated in section 2.2:
What is the reason for the increasing popularity of SNPs, whereas in terms of genetic information provided, as simple bi-allelic co-dominant markers, they can be considered as a step backwards when compared to the highly informative multi-allelic microsatellites? Are we not only putting a new name on what has just been considered until now as a common polymorphism and originally studied as RFLPs? In fact, the more recent SNP concept has basically arisen from the recent need for very high densities of genetic markers for the studies of multifactorial diseases, and the recent progress in polymorphism detection and genotyping techniques.
718 Further, Vignal from section 6.3 and in conclusion stated:
Several approaches can be taken for fine QTL mapping, such as increasing the number of meiosis events by increasing the size and/or the number of families for genotyping, selecting recombination events in recurrent backcrosses, using advanced intercross lines (AIL) or performing linkage disequilibrium and haplotype-based studies in outbreed populations. However, whatever the approach taken, high densities of markers will be needed. In some instances, when the populations studied are closely related, even the microsatellite markers may not be heterozygous for the F1 animals. Also, for some species, such as chickens, the density of microsatellites will be low.
Testing of candidate genes and candidate polymorphisms in exons, promoters or other important regions such as splice sites, promoters or other regulatory regions, will have to be done using the SNP approach, since this will be the most common polymorphism and the more likely responsible for phenotypic variation.
When testing for the association between complex phenotypic traits and candidate loci, single-loci SNP analyses present a loss of information due to the bi-allelic nature of the markers, as compared to the multi-allelic microsatellites. However, by performing haplotype frequency estimations over several SNPs from a locus, this can be overcome and even possibly improved, due to the fact that SNPs will more often be close to the site responsible for the variation than microsatellites.
7. CONCLUSION
Although in a strict molecular sense, SNPs are just what has been previously known as base substitutions, the fact of naming molecular markers by this acronym meaning single nucleotide polymorphism, is an indication of the new importance that this type of polymorphism has in molecular genetics. Indeed, if in some instances, the lack of information due to the bi-allelic nature of SNPs is a limitation, there are cases in which they can provide valuable data on associations between specific genes or other DNA structures and phenotypes, or on population and genome dynamics.
The very high density of SNPs in genomes, usually allows to develop several of them in a single locus of a few hundred base pairs. By reconstructing haplotypes, multi-allelic systems can eventually be defined for analyses, to overcome the limitations due to the low heterozygosity of SNPs. With increasing progress being made in the molecular techniques used to produce SNP data, in the automation of allele scoring and in the development of algorithms for genetic analyses, the effort needed to produce an equivalent amount of information as with microsatellites may some day be equivalent.
719 There is little doubt that the broader use of SNPs was expected to evolve.
720 In summary, although I readily accept that as at the priority date there had always been a desire to improve breeding methods in domesticated animals, it is readily apparent that at that time traditional non-molecular genetic approaches were the most widely used. Moreover, if genetic markers were used to identify causative genes, these markers were principally microsatellites.
Venter
721 Let me now deal with the next topic concerning common general knowledge.
722 MLA says that by the priority date, the Venter paper (Venter JC et al, “The Sequence of the Human Genome” (2001) 291 Science 1304) was well known. This paper was published in February 2001 in Science. Venter reveals that with the help of Celera, over a period of 9 months a 2.91 billion base pair consensus sequence of the human genome was generated by the shotgun sequencing method, which involved collating 14.8 billion base pairs of DNA sequence made up of 27 million high quality sequence reads from five individuals. Further, MLA says that although Professor Taylor appeared to suggest that the method taken by the inventors of the 253 Application was different to Venter because they took DNA from a number of individual cattle, whereas the human genome project did not, he was wrong. MLA says that both projects used multiple sources of DNA.
723 Further, MLA points to the fact that Venter states that 2.1 million single-nucleotide polymorphisms were identified, but less than 1% would result in variation in proteins.
724 Now in my view it has not been established that the information in Venter had become part of common general knowledge as at the priority date, although I accept that shotgun sequencing was well known. Venter was published before the priority date and the news that the human genome had been sequenced no doubt attracted some attention. But the relevant question is the nature of the information, if any, which had become generally accepted in the field of bovine genetics and had become relevantly common general knowledge in the context that I am considering. But even if the information in Venter had become part of common general knowledge, I do not consider that it significantly assists MLA.
725 Venter focuses on the human genome. It provides a general discussion of the identification of SNPs, but it does not describe an association between a SNP and a trait.
726 Further, the sequencing of the human genome published in Venter was done for the purpose of generating a reference assembly, and was not done to create a genome wide panel of informative SNPs. Moreover, the discussion of SNPs in Venter related to issues of human evolutionary history, rather than the potential use of SNPs in animal breeding programs.
727 Further, as compared with Venter, the genome sequencing and assembly methods used by the inventors of the 253 Application were different. Those inventors sequenced DNA from 4 diverse breeds of cattle. These were selected to be representative of relevant genetic diversity. This approach, which is different to the approach of Venter, had complicated contig generation and SNP identification. The inventors were able to identify 786,777 SNPs by comparing each of the sequenced DNA fragments from the four bovine subjects. The inventors then used the human genome as a scaffold to map the location of 242,181 SNPs. This step (also not described in Venter) presented technical challenges as explained by Professor Taylor; now MLA says that this is “pure speculation”, but I consider that there is an air of unreality to that assertion. The disclosure in the 253 Application was the first time that a genome wide panel of SNPs had been positioned for a non-human species using the human genome as a scaffold. Further, the inventors then identified which of the 242,181 SNPs would be highly informative across breeds. The inventors proceeded to identify which of the 242,181 SNPs were polymorphic across a majority of 210 animals from diverse breeds. SNPs that were not sufficiently polymorphic across the 210 animals were discarded and replaced with another SNP within the same region. This process was repeated until a map of 6,189 relatively evenly spaced SNPs was obtained; see Example 1 of the specification. The experts agreed that the scale of genotyping required to perform this process would have been very difficult; again, this process was not described in Venter. Then the inventors in order to identify SNPs that were associated with commercially valuable traits collected phenotypic data relating to marbling, tenderness, fat thickness, daily gain and retail yield from 4,791 bovine subjects. Animals at the extremes of each phenotype were separated and their DNA samples pooled. Each pool of DNA was genotyped for each of the 6,189 mapped SNPs to identify those SNPs that were statistically significantly associated with a trait. This genome-wide association study (GWAS) identified 2,510 SNPs that were associated with the five traits tested. Now Venter does not at all describe a genome-wide association study.
728 So, as I said above, even if Venter was part of common general knowledge, it hardly assists MLA.
729 Further, if it needs to be said at this point, the work carried out as part of the 253 Application could hardly be characterised as routine or involved “normal clinical trials, which utilise certain standards and procedures” (AstraZeneca, at [95] per Kiefel J) or trials that “would conventionally be carried out” (AstraZeneca, at [123] per Nettle J). A careful review of the 253 Application suggests the very opposite.
SNPs as DNA markers
730 Let me deal with another topic concerning common general knowledge.
731 MLA also contends that it was also known that DNA markers, including SNPs, were useful for identifying traits in animals. It is said that at the least, the utility of SNPs (and indeed a dense array of SNPs) was described and known through Venter. In any event, MLA says that regardless of Venter, it was known that SNPs would be useful markers to conduct genome wide association studies and infer/identify traits in bovines.
732 Further, MLA says that although Branhaven seeks to make much of the fact that microsatellites were in greater use than SNPs as at December 2002, that is not the relevant question. It is said that both kinds of markers were available and being used. Further, MLA says that while both were suitable for genotyping animals, the utility of SNPs was not in doubt. MLA says that it was known that there was a far greater prevalence of SNPs in the genome than microsatellites and that such prevalence accounted for much more than the greater polymorphism inherent in microsatellites. MLA also says that it was known that SNPs were easier to genotype than microsatellites and that statistical studies using SNPs were easier to perform than for microsatellites.
733 MLA also says that the methods of identifying SNPs in the bovine genome were also well known. It submitted that such techniques had been used in relation to the human genome (i.e. shotgun sequencing) and were equally applicable to the bovine genome. For example, before the priority date and according to Professor Taylor’s evidence, Genomic FX was using a computer to identify differences in sequences and therefore the presence of SNPs.
734 In response to these contentions I would observe the following.
735 As at the priority date, and indeed for a time thereafter, if genetic markers were being used in cattle breeding they were likely to be microsatellites rather than SNPs. The markers that Dr Sonstegard used during 1997 to 2003 when mapping QTL were microsatellites. His experience was that microsatellites were the most widely used up until the mid-2000s, even though other types of markers may have been available. Professor Taylor also used microsatellites in his own research and Professor Visscher continued to rely on data generated from microsatellites in his quantitative research work in cattle up to 2005.
736 The widespread use of microsatellites as at the priority date meant that they were the most common marker present on the bovine linkage map. They were therefore a valuable resource for QTL mapping experiments as they produced results that were more reproducible across laboratories. Further, microsatellites being more polymorphic were also considered by some of the experts to be more informative than SNPs.
737 Further, as at the priority date very few SNPs had been discovered in cattle. Moreover, the technology that was available for genotyping them was either expensive or low throughput. Indeed, many more bovine microsatellites had been identified as compared to SNPs. As Dr Sonstegard said, as at the priority date researchers were discovering new microsatellites and developing denser linkage maps.
738 Further, as I have already said when discussing the issue of novelty, the Meuwissen simulation was based on markers which more closely resembled microsatellites or behaved more like microsatellites. Vignal also illustrated a practical preference for microsatellites.
739 Now although MLA has sought to diminish the significance of the fact that microsatellites were in greater use than SNPs, I disagree. In assessing what steps the notional skilled person would have taken as at the priority date, the practical advantages (actual and perceived) of microsatellites over SNPs are important to consider. Indeed, Professor Hayes gave evidence that as at the priority date it was possible to implement genomic selection using microsatellites. Professor Goddard also said that although there were only about 4,000 bovine microsatellites known as at December 2002, this would have been sufficient to implement genomic selection. Now I accept that it was disputed by other experts whether there were in fact at the time a sufficient number of microsatellites across the entire genome to carry out a genomic selection approach. But it is apparent from the evidence that even assuming that the notional skilled person had decided to use a genome wide approach, they would have been likely to have used microsatellites.
740 But notwithstanding these observations, I would accept that as at the priority date, and generally speaking, it was part of common general knowledge that SNPs could be used to identify traits in animals. There were examples in the evidence where this had been done, although such examples were not themselves shown to be part of common general knowledge. For example, the Rothschild patent no 5,550,024 (1996) discloses genetic markers for pig litter size, methods for identifying such markers and methods for screening pigs based upon various polymorphisms, preferably a restriction fragment length polymorphism. One can see from the discussion of Figure 11, reference to the Pvu II polymorphism with underlined bases corresponding to polymorphisms between the two sequences generating the Pvu II recognition site. In other words, the patent identifies and uses SNPs as relevant markers.
Comparative genome mapping
741 Let me address another topic. MLA says that comparative genome mapping of species, for example the bovine genome against the human genome, was a well-known technique. It says that it was well known that such mapping allowed for spacing markers evenly along a putative genome designed with the human genome as a scaffold. But MLA says that in any event, it was also known that it was not necessary to have a genomic map in order to do genomic selection in some species, because the screening of many SNPs would inherently involve a spread of SNPs throughout the genome.
742 In my opinion, it may be accepted that comparative mapping between species was known as at the priority date. But it was typically used in relation to discrete loci to identify the specific location of a gene or the order of multiple genes.
Techniques for determining association between SNPs and traits
743 MLA also says that techniques for determining if SNPs were associated with traits were well known. It is said that as at the priority date this involved genotyping animals using known nucleic acid hybridisation techniques and then using standard statistical techniques to assess whether an association between a marker and a trait existed. As noted by Professor Taylor, the least squares approach had been used in a GWAS as early as 1996, and BLUP had been known for a considerably longer time. I accept that these techniques were well known as at the priority date.
Genotyping animals
744 Let me deal with another topic. MLA submitted that as the 253 Application explained at [0152], there were various “[m]edium to high-throughput systems” available for genotyping animals for identified SNPs. In this context, MLA said that Professor Taylor explained that as at the priority date the then high-throughput devices could genotype up to 50 SNPs at a time. In my view it is more accurate to say that Professor Taylor said that devices as at the priority date would generally genotype fewer than 50 SNPs at a time. Professor Taylor also gave evidence that in 2002, DNA samples could be sent to laboratories (e.g. Geneseek) and genotyped for fairly small numbers of SNPs.
745 MLA also says that by December 2002, people in livestock research knew that very high-throughput genotyping tools to use in DNA marker selection for cattle “were coming”. MLA says that SNP chips “as a concept” were not only known as at the priority date but were being used for markers found in the human genome.
746 Further, MLA says that Meuwissen anticipated using this technology. MLA says that the two statistical approaches in Meuwissen that used all markers for genomic selection (being BLUP and Bayes A) were based on the premise that rapid genotyping of SNPs would become possible with new technology. MLA has also pointed out that Professor Visscher referred to this in the 1998 paper by Haley and Visscher (Haley CS and Visscher PM, “Strategies to Utilize Marker-Quantitative Trait Loci Associations” (1998) 81 J Dairy Sci 85). Professor Visscher has also cited a 1999 paper by Lipshutz et al (Lipshutz RJ, Fodor SPA, Gingera TR and Lockhart DJ, “High Density Synthetic Oligonucleotide Arrays” (1999) 21 Nature Genetics Supplement 20), which said it would be possible to have up to 50,000 nucleotides on a chip. I would interpolate at this point though that the first commercial bovine SNP chips (being the 10,000 SNP Chip) that enabled very high throughput genotyping of the bovine genome were not commercially available until 2005. In any event, MLA says that the inventors of the 253 Application did not invent any new method of genotyping animals, but simply used the available devices at the time.
747 Further, MLA says that once Dr Sonstegard became aware of Meuwissen, he immediately set out to make a tool to prove the method of the paper; I would note though that Dr Sonstegard did not become aware of Meuwissen until 2005. Further, Professor Taylor stated that he was so excited by the method in Meuwissen that he spent 5 or 6 years trying to build a SNP chip to test the method once he had heard about it.
748 Now as to these submissions, I accept that as at the priority date, techniques for genotyping animals were known. However, SNP genotyping could only be accomplished in small numbers as high-density SNP genotyping assays had not been developed for any species. Further, in my view the anticipation that very high throughput genotyping tools “were coming” does not necessarily qualify them as forming part of common general knowledge as at the priority date.
Sorting animals and cloning animals
749 MLA contends that it was commonly known how to sort and select animals once the presence of DNA markers for desirable traits were identified, and how to clone animals once they were selected as being desirable. I accept that these methods were part of the common general knowledge.
Meuwissen and the concept of genomic selection
750 Let me deal with the final aspect of common general knowledge concerning Meuwissen that I touched on earlier and discussed in greater detail in the novelty section.
751 MLA says that Meuwissen, the concept of genomic selection and, in particular, the four statistical approaches to genomic selection described in Meuwissen, were well known as at the priority date. Nevertheless, MLA had to accept that the molecular geneticists, Professor Plastow and Dr Sonstegard, had no knowledge before the priority date of Meuwissen. But the quantitative geneticists all knew about the paper before the priority date and considered it to be significant. It is said by MLA that this case is a good example of how the hypothetical skilled addressee might have a greater breadth of common general knowledge than any one particular type of expert.
752 Now although I have discussed Meuwissen previously when discussing the question of novelty, it is appropriate to repeat some aspects.
753 Meuwissen described how to use genetic markers, preferentially microsatellites, throughout the genome of a livestock species (cattle being one example) to predict the genetic potential of an animal for a quantitative trait. The four statistical methods described used differing numbers of markers and differing approaches to detecting an association, which led to different degrees of accuracy of prediction for the trait. Two methods used every genetic marker that was available and were the most accurate (BLUP and Bayes A). The least squares and Bayes B methods chose the most statistically significant number of markers to proceed with, with the least squares method (using the least number of markers) being the least accurate method.
754 MLA says that it was well known at the priority date that the initial genotyping and phenotyping of animals for a dense map of genetic markers as recommended by Meuwissen would be costly and time-consuming. Now the genotyping systems at the time as described by Dr Sonstegard were limited to about 50 SNPs at a time, but MLA says that such work was “simply a matter of cost and time”. It is said that that is what the inventors did in relation to the 253 Application in that they genotyped a dense map of markers in populations of animals which had been phenotyped. But MLA says that they did not invent any new way of doing that work.
755 Further, MLA says that in terms of making a commercial genotyping tool that could be used to select animals as at the priority date, two of the statistical approaches in Meuwissen involved the use of a selection of the most significantly associated markers. That is, while the initial work in selecting the best markers out of the dense map of markers would have been costly and time consuming, it was possible to choose a much smaller number of markers. So, for the Bayes B method, you would discard 95% of the markers. It is said that that could provide some accuracy for prediction of the trait using existing genotyping devices as described by the specification. Further, MLA says that while SNP chips, which could handle thousands of SNPs, were not at the priority date commercially available for bovine species, they were in use in humans and were known to enable the use of all or most of the markers used for the BLUP and Bayes A methods disclosed in Meuwissen.
756 Now in my view, MLA has not established that Meuwissen had become part of the common general knowledge (although this may not matter as it can be brought in under s 7(3)). The fact that the publication Genetics may have been widely read, and the information presented and discussed at some conferences, is not sufficient to establish that Meuwissen had become common general knowledge. Professor Visscher did not know how widely accepted Meuwissen had become in December 2002. And it is not to be forgotten (as I have said) that Vignal, a reasonable summary of what was known at the relevant time, did not even cite Meuwissen.
757 Indeed, there was some evidence before me suggesting that Meuwissen (or the information therein) only became widely known after 2004 when the first genome reference assemblies and SNPs were deposited in the public domain and the first results were produced based on the analysis model disclosed in Meuwissen.
758 Further, the practical application of the approach suggested in Meuwissen referred to, inter-alia, a dense genome wide panel of markers which did not become available until the mid-2000s; but I do accept that it disclosed that a subset of markers could be used. Further, as Professor Taylor noted, it was also not clear how marker density would influence the accuracy of the approach suggested in Meuwissen or whether there was a marker density below which the approach would not work.
(c) Obviousness in light of common general knowledge
759 Having addressed in some detail what was common general knowledge, it is appropriate to now consider the question of obviousness in that light.
760 MLA has contended that before the earliest priority date, the hypothetical skilled person (or team) armed with the common general knowledge at the time, would have desired to produce a molecular approach for the selection of cattle for traits of economic significance. The team would also know that such traits were generally quantitative traits with a large number of genes each accounting for a small amount of the trait.
761 MLA contends that in addressing that desire, the team would know of the Meuwissen genomic selection approach. The team would know that the approach required at the outset a large set of genetic markers spread throughout the genome to take into account average LD in the target population.
762 MLA says that the team would be aware that LD not only changes from population to population, but even within the genome of a single animal. Accordingly, the team would pick an average LD when deciding how dense the marker map needed to be. MLA relies upon Professor Taylor’s evidence that at the time, for bovine it was known that there was widespread disequilibrium within the cattle genome and that it was anticipated that LD would be in the order of 500,000 bp. Further, MLA says that in any event, it did not matter if the estimate of LD was wrong. It says that reducing the number of markers would not have resulted in a large loss in accuracy, and relies accordingly on the evidence of Professor Goddard and Professor Visscher. It was simply a matter of degree.
763 MLA says that the skilled team would also be aware of the Venter paper and so was aware that it was possible to map the entire bovine genome within at most 9 months and, by the priority date, within 26 days, by shotgun sequencing (assuming that you had the number of machines that Celera had). The team would also know from Venter that the approach would disclose a dense set of SNPs throughout the genome which would be useful for association studies. Alternatively, the same skilled team was aware that it could use a less complete shotgun sequence and map it to the human genome.
764 MLA says that the skilled team would also be aware of how to phenotype animals. It had been practised as part of the progeny selection approach since well before 2002. The skilled team would also know how to genotype animals in a test population for the dense set of SNPs spread throughout the genome for associations between the SNP markers and a trait, or indeed, numerous traits. The only barrier to these things was time and cost.
765 MLA also says that the skilled team would have been aware that it could use two of the approaches disclosed in Meuwissen immediately on existing genotyping devices (i.e. Bayes B and least squares), though such approaches would be less accurate than the approaches using all markers (i.e. BLUP and Bayes A). Alternatively, the skilled team would appreciate that SNP chips were being used in humans and, once made commercially for bovine, could be used to employ the more accurate approaches disclosed in Meuwissen. MLA says that the skilled team was thus aware of how to correlate the SNP markers with traits using any one of the statistical methods reviewed in Meuwissen, with varying degrees of accuracy. Accordingly, in addressing the desire to arrive at a new or improved method for selecting bovine as at the priority date, the skilled team would directly be led as a matter of course to try each of the approaches in Meuwissen in the expectation that it might well produce such a method. Each of the Meuwissen statistical methods would involve the use of a number of SNP markers spread throughout the genome, being associated with a quantitative trait, and used to identify the genetic potential of a bovine subject for the trait. MLA says that on the balance of probabilities, at least three of those markers would inevitably be within 500,000 nucleotides of a specified SNP and at least two of the markers would be within a gene.
766 Further, MLA says that the skilled team would know that it could use the predictive power of any of the approaches in Meuwissen to sort animals, select the best animals for breeding or clone them if so desired.
767 Further, MLA has contended that there would have been no relevant hurdles or difficulties as at the priority date. But that if there were difficulties and hurdles, they only “boiled down” to time and cost.
768 The first difficulty advanced by Branhaven is said to be identifying a sufficient number of markers. But MLA says that at the time, the shotgun sequencing method used in the human genome project as described in Venter was available and would produce a significant number of SNPs in bovine. It says that that was the method in fact used by the inventors (253 Application at [0053], [0191] and [0192]) and the 253 Application does not identify any particular technical difficulty or hurdle overcome or experienced in applying that method, nor does it describe the method other than by reference to Venter.
769 The second difficulty advanced by Branhaven is said to be the task of evenly spreading the SNPs throughout the bovine genome. But MLA says that that had already been done with the human genome. Also, it could either have been done against an existing bovine linkage map (as was in fact done in Grosse and Gelhaus & Förster (Gelhaus A and Förster B, “Cattle MHC Genes DOA and DOB: Sequence Polymorphisms and Assignments to the Class IIb Region” (2001) 28 European Journal of Immunogenetics 429) or using the human genome as a scaffold, which is what the inventors of the 253 Application did. MLA accepts that both those methods would be inaccurate to some degree, but not useless. MLA says that otherwise, the SNPs could be mapped against the bovine genome once it was created, which was just a matter of time and cost (26 days for Celera), but not technical difficulty. But, in any event, MLA says that this difficulty would not in reality exist if sufficient SNP markers were used because they would necessarily be spread throughout the genome. That is, it was not in fact necessary to map the markers at all. MLA relies upon the evidence of Professor Visscher in support of such propositions. MLA says that the above analysis “does away with” the evidence from Professor Taylor regarding the “non-trivial” exercise of collating and mapping the shotgun sequences.
770 The third difficulty advanced by Branhaven is said to be genotyping thousands of SNPs. But MLA says that the ability to genotype up to at least 50 SNPs was not difficult and so the difficulty in genotyping many more was simply a matter of time and cost. Furthermore, SNP chips (array of SNPs) was the obvious solution, which approach was being taken for markers found in the human genome. In any event, so MLA contends, the 253 Application does not itself suggest that there is any such technical difficulty. The 253 Application describes various “known in the art” medium to high-throughput systems for analysing SNPs (253 Application at [0152]). Accordingly, MLA says that even if Branhaven was right to assert that it was speculation whether a SNP chip would become available, that does not answer the inventive step case because that was not the only way of achieving the invention.
771 Generally, MLA says that if a path is known or an obvious one to take, that is not altered by the fact that it might take a long time and be expensive. MLA accepts that it would have taken a long time and cost a lot of money to apply the claimed method to thousands of SNPs until the bovine SNP chip became available. But MLA says that does not mean that it involved an inventive step to seek to apply the method without a SNP chip, or to consider using one in the future. It says that a lack of resources is simply not relevant to the inventive step question.
772 In summary, MLA says that the evidence shows that the skilled team, faced with the problem of improving the methods for predicting the genetic potential of animals and armed with sufficient resources, would have been led directly as a matter of course to something falling within each claim of the 253 Application in the expectation of producing a better method than the existing approaches. It says that a finding of obviousness must follow. It contends that there is no inventive step in identifying naturally occurring SNPs and their natural associations with traits as a result of following an obvious path. Once the obvious path was followed, the result was inevitable.
773 Further, to Branhaven’s assertion that as at the priority date no one was practicing genomic selection or GWAS because there were nowhere near enough markers available to apply either and that this was a profound technical hurdle, MLA says that the 253 Application neither describes a new way to identify such markers nor claims one. It simply sets out the use of known methods (Venter) to find them and claims not only the result of their work, but any SNP falling within the region of about +/- 500,000 bp of a specified SNP.
774 Further, MLA says that to the extent that Meuwissen can be said to have made the concept of genomic selection well known in livestock breeding, at most the inventors reduced that idea to practice using known techniques. It is asserted that Dr Sonstegard agreed that “all” the inventors did was reduce an existing concept to practice and that reduction to practice involved no new techniques. I do not consider that this assertion fairly reflects the totality of Dr Sonstegard’s evidence (or for that matter Professor Plastow’s evidence) or the appropriate concessions they were prepared to make under cross-examination.
775 MLA also says that no inventive step is given to the method by the addition of the requirement that at least three SNPs occur in more than one gene, because this is merely an arbitrary parameter. As I have said, I have rejected that contention.
776 In summary, MLA says that for these reasons, all of the claims lacked an inventive step in light of the common general knowledge as at the priority date. Further, the additional features of claims other than claim 1 were also all well known.
Analysis
777 In my view MLA has not established that common general knowledge would have directly led the skilled person to the claimed invention. Common general knowledge did not suggest using SNPs identified by a genome wide association study to identify a trait, and a fortiori the particular method claimed in the 253 Application.
778 Now the Venter paper (assuming in favour of MLA that it was part of common general knowledge) disclosed the shotgun sequencing of the human genome, but it was not concerned with the construction of an informative genome wide dense panel of markers of SNPs to identify commercially important traits in bovine. Further, and contrary to MLA’s assertion (as I understood it), the task of evenly spreading the SNPs throughout the bovine genome had not already been done with the human genome. Further, Venter did not disclose any processes for selecting and refining SNPs to ensure that the SNPs were informative for traits.
779 Further, Meuwissen (assuming in favour of MLA that it was part of common general knowledge) simulated genomic selection assuming that a dense genome wide panel of markers already existed. As Professor Goddard said: Meuwissen “assume[d] that there are these markers known and can be used, so we assume that there are a panel of genome-wide markers available. We don’t describe how you would actually discover them”.
780 Further, it is not apparent why a skilled person would choose the least accurate approach (least squares) in Meuwissen, particularly if, as asserted by Professor Goddard, there was a need for high levels of accuracy.
781 Further, although MLA has contended that the skilled person would appreciate that SNP chips were being used in respect of humans, and once made commercially for bovine could be used to employ the more accurate approaches disclosed in Meuwissen, as at the priority date genome wide SNP chips were not being widely used in respect of humans.
782 Further, I would also reject a central proposition of MLA’s inventive step case that the construction of an informative SNP map for the entire bovine genome at the priority date, involved only routine steps.
783 Let me begin by saying that MLA did not lead evidence from a molecular geneticist on this issue and it was accepted by all experts that the construction of a SNP map for the entire bovine genome was work done exclusively by a molecular geneticist. MLA’s witnesses accepted that they had no relevant laboratory experience. Professor Goddard has never performed molecular genetic techniques and did not know how to. Neither Professor Hayes nor Professor Visscher had ever worked in a laboratory save for Professor Hayes saying that he was once “marginally involved”.
784 I agree with Branhaven that little weight should be accorded to the assertions of MLA’s witnesses that the steps that a molecular geneticist would have had to take to construct and characterise an informative SNP map for the entire bovine genome was entirely a matter of routine steps. I accept the evidence of both Professor Plastow and Dr Sonstegard, both highly experienced molecular geneticists, that the construction of a SNP map for the entire bovine genome was anything but routine.
785 As Professor Plastow and Dr Sonstegard said, the development of a sufficient number of markers would have presented a significant technical hurdle. Moreover, the development of a dense panel of markers, which was adequately spaced throughout the genome in order to adopt the genomic selection approach, presented even greater difficulty. In my view, and contrary to MLA’s assertions, it was not just a matter of “time and expense” or a mere matter of “scale”.
786 As Branhaven correctly pointed out, one key objective touchstone of the practical difficulties in constructing an informative SNP map is the time that it actually took for researchers in both the fields of human and bovine genetics to produce a sufficient number of SNPs to construct a dense genome wide panel of markers.
787 Professor Visscher referred to the 1996 publication of Risch and Merikangas in relation to research into human disease (Risch N and Merikangas K, “The Future of Genetic Studies of Complex Human Diseases” (1996) 273 Science 1516) which proposed a GWAS approach in humans. It was apparent from Professor Visscher’s review article in 2012 titled “5 Years of GWAS Discovery” that the first results for a GWAS study in humans were not published until 2005 and 2006 and that the starting point was actually considered to be the 2007 Wellcome Trust paper (The Wellcome Trust Case Control Consortium, “Genome-Wide Association Study of 14,000 Cases of Seven Common Diseases and 3,000 Shared Controls” (2007) 447 Nature 661). This was all about 10 years after first being suggested by Risch and Merikangas. But Professor Visscher did say in his oral evidence that he was aware that in 2002 a GWAS study was carried out on humans where a SNP chip wasn’t used but a different experimental technique was undertaken.
788 And even in 2005, it was recognised with reference to studies of humans that whole-genome association studies had not generally previously been possible because dense enough marker maps and high-throughput genotyping platforms were lacking (see Uimari P, Kontkanen O, Visscher PM, Pirskanen M, Fuentes R and Salonen, JT, “Genome-Wide Linkage Disequilibrium from 100,000 SNPs in the East Finland Founder Population” (2005) 8 Twin Research and Human Genetics 185).
789 As for bovines, the evidence of Dr Sonstegard and Professor Taylor establishes that the first 10,000 SNP chip did not became available until 2005. Further, the evidence of Professor Plastow and Dr Sonstegard also establishes that the 50,000 SNP chip did not become available until 2008.
790 Now MLA has asserted that people in the field knew that something was coming, for example, high density SNP chips or high throughput genotyping tools. But in my view that anticipation is not a proper basis upon which to answer the statutory test imposed by s 7(2) of the Act. As Branhaven rightly contended, that the construction of a SNP map for the entire bovine genome was theoretically possible does not make its construction a matter of routine. Nor does the fact that there was interest in the use of molecular genetics in cattle breeding as at the priority date.
791 Further, as I touched on earlier in my reasons, there was a definitional debate amongst the experts as to the distinction between GS and GWAS. To the extent that anything turns on it, I accept the distinction identified in the Rabier article between GS and GWAS (Rabier C-E, Barre P, Asp T, Charmet G and Mangin B, “On the Accuracy of Genomic Selection” (2016) 11(6) PLoS ONE e0156086). The difference between GS and GWAS is that GS involves all the marker effects being estimated simultaneously. To adopt the language of Professor Goddard, in GS all markers are “taken into account” and “treated impartially”. Similarly, Professor Visscher suggested an approach that involved “estimating marker effects for all SNPs”. Contrastingly, GWAS involves establishing specific associations between individual SNPs and a trait. But it is also important to observe that this distinction was not necessarily apparent as at December 2002. But it is not in dispute that both GWAS and GS require a dense, genome wide, panel of markers.
792 Let me now proceed to make some general observations.
793 If one is applying the Cripps question, it must be shown that the notional research group would directly be led as a matter of course to try the combination of integers claimed in the expectation that it might well produce a useful result. The combination of integers claimed in the 253 Application require, inter-alia, the use of at least 3 SNPs, one which must be selected from the panel of 2,510 SNPs or SNPs within about 500 kilobases either side of these SNPs. And all of the at least 3 SNPs must be associated with a trait. Now even if it were accepted that the notional skilled person would have routinely identified and characterised SNPs across the bovine genome, MLA has not established that the skilled person would arrive at the actual SNPs of the 253 Application, let alone the use of at least 3 SNPs in the claimed method. And even if it were assumed that a sufficient number of SNPs had been identified, different researchers would make their selection from available SNPs in different ways as Professor Plastow explained. And that is irrespective of whether the claims are construed to require that the limb (b) SNPs be in LD with the limb (a) SNPs.
794 Further, although MLA has asserted that each of the various steps taken by the inventors of the 253 Application were individually known and therefore the invention as claimed was obvious, I agree with Branhaven that such an approach invites error. The invention is directed to a method for inferring a bovine trait using a number of identified SNPs all of which must be associated with the trait. Even if it were accepted that each of the individual methods used by the inventors were known techniques, that would not provide a proper basis for finding that the claimed invention was obvious.
795 Further, although MLA also relies on the fact that the 253 Application does not identify particular technical difficulties in the steps undertaken by the inventors, there is no requirement for any such disclosure.
796 Further, let me deal with another point raised by MLA concerning claim 1 and inventive step. MLA says that where one is dealing with a combination of elements, it is their interaction which is the essential requirement of invention (Aktiebolaget Hässle at [6] and [41]). So much may be accepted. But MLA seeks to draw too much from the notion of interaction in suggesting that the integers of claim 1 do not interact. One is dealing with a method claim which has “component” parts. But the parts do not need to interact like a physical locking mechanism in order for the component parts to interact in a relevant sense to form a new invention. Each component is necessary to, and the combination is sufficient for, the invention as claimed. In the relevant sense they interact.
(d) Section 7(3) documents
Meuwissen as a section 7(3) document
797 As I have said above, I have not accepted MLA’s contention that the information in Meuwissen was common general knowledge. But MLA also says that it can at least be relied upon under section 7(3) of the Act. Now as I have previously indicated, as the 253 Application was filed after 1 April 2002 and examination requested before 15 April 2013, the amendments to section 7(3) of the Act brought about by the Intellectual Property Laws Amendment (Raising the Bar) Act 2012 (Cth) do not apply to the 253 Application.
798 Under the applicable version of s 7(3), for a document to be relied on under that provision there must be some basis for concluding that the skilled person could be reasonably expected to have ascertained, understood and regarded as relevant the particular prior art in question. The opponent may need to adduce specific evidence to support a contention that a particular prior art document satisfies the requirements of s 7(3), although in some cases this may be readily apparent without direct evidence.
799 But even if information is able to be taken into account under s 7(3) of the Act, that is not the end of the matter. The opponent must still establish on the evidence, that the invention would have been obvious in the light of that information taken together with the common general knowledge.
800 In AstraZeneca, Kiefel J summarised the effect of s 7(3) as follows (at [68] to [70]):
Before a document containing prior art information can be used along with the common general knowledge for the purposes of the s 7(2) inquiry, it is necessary that it meet the requirements of s 7(3). In Lockwood Security Products Pty Ltd v Doric Products Pty Ltd (No 2) it was explained that prior art information which is publicly available in a single document is “ascertained” if it is discovered or found out, and “understood” means that, having discovered the information, the skilled person would have comprehended it or appreciated its meaning or import. The Court also explained that the phrase “relevant to work in the relevant art” is directed to publicly available information, not part of the common general knowledge, which the skilled person could be expected to have regarded as relevant to solving a particular problem, or meeting a long-felt want or need, as the patentee claims to have done.
Lockwood (No 2) also explains [at [127]] that, in answering the question of obviousness, the information referred to in s 7(3), like that part of the prior art base which is the common general knowledge, is considered for a particular purpose. That purpose is to look forward from the prior art base to see what the skilled person is likely to have done when faced with a problem similar to that which the patentee claims to have solved with the claimed invention. It is this aspect of the s 7(2) inquiry which assumes particular importance on these appeals.
In addressing s 7(2), it is to be borne in mind that the skilled person is an artificial construct, intended as an aid to the courts in addressing the hypothetical question of whether a person, with the same knowledge in the field and aware of the problem to which the patent was directed, would be led directly to the claimed invention. The statute’s creation of the skilled person construct for this purpose is not to be taken as an invitation to deal with the question posed by s 7(2) entirely in the abstract. Whilst the question remains one for the courts to determine, the courts do so by reference to the available evidence including that of persons who might be representative of the skilled person. (citations omitted)
801 French CJ made clear at [43] and [44] that data is not required in order for a prior publication to render an invention obvious, provided that experiments to create such data would be routinely carried out:
The question posed by this Court in Wellcome Foundation Ltd and AB Hässle does not require that, in order to sustain an obviousness case, a party has to lead evidence which echoes the terms of that question. A similar conclusion was open, as the primary judge found, on Patent 471.
AstraZeneca submitted in this Court that the “claimed invention” is a treatment using a once daily, 5-10 mg dosage of rosuvastatin. The only dosage expert, Dr Reece, confirmed that neither the Watanabe Article nor Patent 471 contained animal or human trial safety data. He had given evidence that such data were essential to determining what dosage should be tested in clinical trials. The person skilled in the art would never have chosen the dose to be tested simply by trying the doses that worked for other statins. That evidence was referred to by Jessup J, who said:
But that evidence also made it quite clear that such trials would conventionally be carried out. They would fall within the concept of working towards the invention with an expectation of success referred to in AB Hässle.
No error is disclosed in that reasoning. (citations omitted)
802 MLA says that that is the case here. It says that the evidence shows that a person skilled in the relevant art, faced with the task of improving existing methods to predict the genetic potential of cattle, even if not aware of Meuwissen before December 2002, would have found it in a search, understood it and regarded it as relevant, and would have been directly led by Meuwissen to practically apply each of the statistical approaches described therein as a matter of course, subject only to available resources. MLA says that the person skilled in the relevant art would have expected to improve existing methods by doing so.
803 MLA says that to the extent that Meuwissen does not defeat any of the claims of the 253 Application on the ground of lack of novelty, then it must defeat all of the claims on the ground of lack of inventive step (whether it is part of common general knowledge or relied upon under s 7(3)).
804 Now I accept that Meuwissen could reasonably be expected to have been ascertained by the skilled person at the priority date and accordingly can be relied upon under s 7(3). But the problem for MLA is that Meuwissen was a hypothetical simulation and accordingly did not involve any real data. Further the aim of Meuwissen was (at 1820, left hand column):
to compare least squares, BLUP and Bayesian analyses for their accuracy of predicting total breeding value of individuals in a situation where a limited number of recorded individuals are genotyped for many markers with many alleles per marker.
805 Now MLA has relied on the proposition from AstraZeneca (at [43] and [44] per French CJ) that data is not necessarily required to be provided in a prior art document to render a claimed invention obvious. But that observation was subject to the qualification that the experiments required to create the data would conventionally be carried out. But contrastingly, the experiments required to produce the necessary data based on the Meuwissen simulation could not sensibly be said to be merely routine work.
806 Further, Meuwissen taught away from the claimed invention. The least squares method discussed in Meuwissen performed worse than the three other statistical methods discussed. It greatly overestimated some haplotype effects and underestimated others. Professor Hayes accepted that the least squares statistical method discussed in Meuwissen was the worst performing method of the four discussed, although he said that it did “work”.
807 In my view it is likely that the skilled person would have been led by Meuwissen to use the BLUP method or Bayes A method which took into account the potential effect of all markers. He would not have been led to use the worst performing statistical method which only considered a sub-set of informative markers. So in my view he would have been led away from an approach which was closest to the claimed method of the 253 Application.
808 Further, it was accepted by both Professor Goddard and Professor Hayes that the simulation in Meuwissen was based on markers akin to microsatellites rather than SNPs. So even if it were accepted that Meuwissen would lead the skilled person to adopt a genome wide approach, he would have likely done so using microsatellites; this is fortified by the evidence that there were sufficient microsatellites available at the priority date to perform a genome wide analysis in cattle.
809 In summary, in my opinion there is nothing in Meuwissen that would lead to the use of at least 3 SNPs each of which is associated with a trait, as required by the method of claim 1 (and other claims) of the 253 Application.
Moody or Grosse as section 7(3) documents
810 As I have already explained, MLA’s primary contention is that either of Moody or Grosse anticipate certain claims of the 253 Application. But MLA says that if, contrary to that contention, none of those documents anticipates at least claim 1 of the 253 Application, then the claims are bad for lack of inventive step in light of common general knowledge in combination with any one of those documents.
811 MLA points to Professor Visscher’s explanation that in order to assemble a genome-wide panel of SNPs in bovines, he would have started by looking at what SNPs other researchers had already identified for inclusion in his panel of SNPs. Professor Visscher said that he would have ascertained, understood and regarded as relevant each of Moody and Grosse as disclosing SNPs that he could use in his SNP panel. Further, MLA points to the work by Professor Hayes that is said to confirm that a majority of those SNPs fall within the broad +/- 500,000 nucleotide ranges in the claim and all fall within “genes”.
812 So, MLA contends that I can be confident that the person skilled in the art would have hit upon a method falling within the claims of the 253 Application, as a matter of routine, in light of common general knowledge and either of Moody or Grosse, considered separately.
813 Now in the light of Professor Visscher’s evidence, I am prepared to accept that Grosse or Moody can be considered under s 7(3). But I am not sure where any of this takes MLA.
814 MLA’s witnesses were not asked to consider whether the invention as claimed would have been obvious in light of Grosse or Moody when combined with the common general knowledge. The issue was also not addressed by the experts in the conclave questions and answers relating to Grosse and Moody, which were directed to the novelty of claim 1. Further, as I have said, Professor Goddard accepted that Moody disclosed no associations between any SNP and any trait. I reject MLA’s assertions that either Grosse or Moody made out together with common general knowledge a lack of inventive step. Now MLA asserted that based upon the evidence of Professor Hayes, Moody or Grosse on his analysis gave you limb (b) SNPs. I must say that I was not sure whether all of this had been looked at carefully enough by the experts. But in any event I consider it a stretch to conclude from his findings and common general knowledge that this established a lack of inventive step.
The Gengler Patent as a section 7(3) document
815 In some respects it is unclear whether MLA is raising the Gengler Patent as a s 7(3) document. If it is it has not established the point. MLA has adduced no evidence that the skilled person would have ascertained the Gengler Patent. Professor Visscher gave evidence that as at 2002-2003 it was relatively rare for breeding companies to obtain patents in the field of animal breeding and genetics that directly affected his research. Professor Plastow agreed with this and did not consider patent literature to be a relevant source of information as to the state of knowledge in his field. Moreover, none of MLA’s witnesses gave evidence that they would have ascertained the Gengler Patent.
816 In any event, none of MLA’s witnesses were asked to consider whether the invention as claimed would have been obvious in light of the Gengler Patent when combined with the common general knowledge. The issue was also not addressed by the experts in the conclave questions and answers relating to the Gengler Patent, which was directed to novelty questions.
817 In my view, MLA has adduced no evidence to support its contention that the invention as claimed would have been obvious in light of the Gengler Patent when combined with the common general knowledge. And on my reading of the Gengler Patent, that would not be established in any event.
(e) Conclusion on inventive step
818 I reject MLA’s obviousness case. The evidence does not establish, and certainly not clearly, that the invention as claimed does not involve any inventive step.
819 Further, although the above discussion focuses on the subject matter of claim 1, I accept that each claim of the 253 Application is to be considered independently for the purposes of this ground (as with other grounds). But MLA has not clearly satisfied me that any of the remaining claims lack an inventive step based upon the test(s) discussed in Aktiebolaget Hässle at [50] to [53].
Lack of utility
820 Section 18(1)(c) of the Act requires that an invention be “useful”, with the onus of showing otherwise resting on MLA. Moreover, in the context of the present appeal, for MLA to succeed on this ground I must be clearly satisfied of a lack of utility.
821 Three questions may be posed. What has been promised (as gleaned from the specification) for the invention as delineated by the relevant claim? Is the promise useful? Has the promise been met? In this context the patent application (including the claims) must be construed from the perspective of a skilled person, and not in a way that such a person would appreciate would lead to unworkability when by construction it could be given a more limited meaning.
822 There is no need for the patent specification to provide support to demonstrate that a promise of the claims was satisfied (SNF (Australia) Pty Ltd v Ciba Speciality Chemicals Water Treatments Limited (2011) 92 IPR 46; [2011] FCA 452 at [296] per Kenny J). And as Jagot J accepted in Apotex Pty Ltd v AstraZeneca AB (No 4) (2013) 100 IPR 285; [2013] FCA 162 at [352], “[u]ltimately, an asserted lack of utility must be established by appropriate evidence, not by mere speculation that the invention will not work or meet the promise set out in the specification.”
823 Now a lack of commercial viability or commercial practicality does not establish lack of utility. This point was made (in part) in Lane Fox v Kensington and Knightsbridge Electric Lighting Co Ltd [1892] 3 Ch 424 at 431 per Lindley LJ, cited by the High Court in Advanced Building Systems Pty Ltd v Ramset Fasteners (Aust) Pty Ltd (1998) 194 CLR 171 at [24], with that Court stating:
It is no objection to the validity of a patent granted under the Act that it is commercially impracticable; its utility depends on whether, by following the teaching of the complete specification, the result claimed is produced.
824 Further, when construing the specification, it is important to consider the nature of any promises. Unless the specification would be understood to convey a clear assertion that the invention will achieve a particular outcome across the breadth of the claims, inutility will not be demonstrated by showing that in some cases that outcome may not be achieved. A claim may have utility even though the promised advantage is not achieved in all cases. In Rescare Ltd v Anaesthetic Supplies Pty Ltd (1992) 111 ALR 205; [1992] FCA 811, the claimed method of treatment utilised a nose-piece apparatus applied to patients suffering from snoring sickness so as to maintain air pressure sufficient to keep the nasal passages open. It was argued that the claim was not useful as not all patients would achieve a beneficial result. Gummow J said (at 232):
One looks at the claim to see whether there is a failure to fulfil that promise. It is not necessary to show utility that the promise be fulfilled in every case. On the evidence, the claimed invention plainly is of considerable practical utility in the treatment of substantial numbers of persons who are “patients” within the meaning of claim 1.
825 See also Sanofi-Aventis Australia Pty Ltd v Apotex Pty Ltd (No 3) (2011) 196 FCR 1 at [245] per Jagot J.
826 Moreover, there are limits on any broad principle, if there be one, that everything within the scope of a claim must be useful (cf H Lundbeck A/S v Alphapharm Pty Ltd (2009) 177 FCR 151 at [217] per Bennett J). Lehane J in Aktiebolaget Hässle v Alphapharm Pty Ltd (1999) 44 IPR 593; [1999] FCA 628 at [227] observed in respect of such a broad principle that “[a] degree of caution, however, is required”. If qualifications and expedients necessary to make the invention work are left to the reader to supply, that does not necessarily equate with inutility.
(a) MLA’s contentions
827 MLA says that if a claim, properly construed, includes within its scope means that will not produce the promised result, even if a skilled addressee would recognise which means to avoid, then the claim will lack usefulness.
828 Now MLA says that the promise of the invention is the provision of a very accurate method of predicting genetic potential of animals. MLA places heavy emphasis on that part of the specification that says at [0101]: “This invention identifies animals that have superior traits, predicted very accurately, that can be used to identify parents of the next generation through selection.”
829 MLA says that Branhaven has attempted to “run away” from the clear words of the specification at [0101] by asserting that, to the extent there is a promise, it is merely “to allow the identification and selection of cattle with superior genetic potential for desirable characteristics”. MLA says that even if there was such a broad promise, it does not take away from the clear promise at [0101].
830 Now on the assumption that the claims do not require a statistically significant association, a proposition that is now contestable as I will require this by way of amendment as I have already said, MLA says the following.
831 MLA says that as the claims only require SNPs to be “associated” with traits, with no degree of association specified, the claims will include at least three SNPs that will not, collectively, accurately predict genetic potential in relation to such traits.
832 MLA says that the claims include specified SNPs that are asserted to have an association with a trait in the discovery bovine population only. But as Professor Hayes has pointed out, none of those has been validated as associated in any other bovine population. Moreover, the claim includes non-specified SNPs for which no association has been shown, but is only assumed based on LD. Accordingly, so MLA says, it is not possible to say with respect to any specified SNP or non-specified SNP that it will accurately predict genetic potential in relation to any trait. Nor is it possible to say that a combination of at least three SNPs wherein at least one SNP is a specified SNP or a non- specified SNP will accurately predict genetic potential in relation to any trait.
833 MLA says that the experts agreed that there will be a number of false positive results, which means that there will be SNPs within the claim that will not predict association with a trait. I agree that the evidence establishes that there are likely to be false positives. But MLA had to accept that the experts disagreed as to the number of false positive results. But MLA pointed to the evidence of Professor Goddard that it is likely to be higher as a result of the way in which Example 2 was conducted. MLA says that it is also likely to be very significant as no degree of association between a SNP and a trait is required by the claims.
834 Further, MLA says that even the addition of an integer requiring that the non-specified SNPs be in LD with a specified SNP could not save the claims from a finding of lack of utility, as one SNP being in LD with another SNP does not necessarily indicate that both SNPs are associated with the same trait. Professor Goddard gave evidence that you can only be certain of this if both SNPs are in 100% LD. But in my view, if LD is injected into limb (b) and each of the 3 SNPs has to have a statistically significant correlation with the trait, this may not be as much of an issue.
835 Now on the alternative assumption that the claims require a statistically significant association, MLA says the following.
836 MLA has contended that even if the claims require that there must be a statistical significance in the association between at least three SNPs and the trait in question with a p value of equal to or better than 0.01, that does not resolve the issue.
837 First, it is said that a mere statistical significance in a test population is not useful until it is validated in the working population. MLA says that even if the word “associated” in claim 1 required statistical significance, it does not require validation. Accordingly, it says that given the number of false positives that would exist in the specified SNPs (Professor Hayes indicated there would be about 305) it is possible to choose at least three SNPs which will not work (and more SNPs as required in the later claims).
838 Second, MLA says that even if a claim required not only a statistically significant association but also a validated association, that still does not assist Branhaven because the degree of statistical significance it attributes is too low to very accurately predict multigene quantitative traits from just three SNPs. Professor Goddard gave an example of a study of human height. The study found that in 250,000 people, 700 markers were identified that were statistically significant. All of those markers explained 16% of the variation in height. Therefore three of those markers chosen at random would explain only about 1% of the genetic variation. Professor Goddard said that this was typical of virtually every trait that had been looked at. Further, Professor Hayes stated that traits in cattle are similar to the height trait in humans. Accordingly, MLA submits that 3 or even 10 of the specified 2510 SNPs are not enough to make an accurate prediction.
839 I must say that this particular issue, as well as various aspects of construction that I have discussed earlier that go to clarity and a description of the invention, has troubled me the most.
(b) What is the promise?
840 It is convenient to again set out [0022] and [0101] of the specification which are potentially relevant to identifying the promise:
[0022] In summary, three different experimental approaches have been used with limited success to identify genes, chromosomal regions or DNA markers that account for a large proportion of the genetic variation observed in economically important traits in livestock species. The results achieved from research programs utilizing these methods have not been widely utilized to date because they do not account for enough of the total genetic variation to allow accurate prediction of an animal’s performance for a specific trait. Furthermore, even when successful these approaches are only capable of identifying additive genetic components while ignoring non-additive genetic components such as dominance (i.e. dominating trait of an allele of one gene over an allele of another gene) and epistasis (i.e. interaction between genes at different loci) which are critical to the development of diagnostics that can be utilized by animal breeders to accurately predict genetic potential for economically important traits in livestock species.
…
[0101] This invention identifies animals that have superior traits, predicted very accurately, that can be used to identify parents of the next generation through selection. These methods can be imposed at the nucleus or elite breeding level where the improved traits would, through time, flow to the entire population of animals, or could be implemented at the multiplier or foundation parent level to sort parents into most genetically desirable. This invention provides a method for determining the optimum male and female parent to maximize the genetic components of dominance and epistasis thus maximizing heterosis and hybrid vigor in the market animals.
(c) Some of the expert evidence
841 It is also appropriate to record some of the expert evidence relevant to this aspect.
842 Professor Goddard did not believe that the inventors of the 253 Application accounted for possible statistical errors when they identified the 2,510 significantly associated SNPs. Consequently, the 2,510 SNPs would have included a number of false positive results. Referring to the process at [0195] to test each of 6,189 mapped SNPs for each trait, Professor Goddard explained that at least 60 (i.e. 1%) of the associated SNPs were likely to be false positives. As this process was carried out for each of five traits, it was likely that 300 of the 2,510 SNPs were false positives and these could not be identified or distinguished. Further, it was his opinion that, based on his experience, the statistical test performed by the inventors would have yielded a false positive result more often than the assumed rate of 0.01. Professor Goddard considered that there were other confounding factors that the experiments in the 253 Application did not account for, including differences between bovine breeds. These factors would have likely resulted in additional false positive results. In light of these factors, a p value of 0.01 was inappropriate and a more stringent statistic should have been applied. Further, Professor Goddard considered that the significance test applied to determine association undermined the utility of the 2,510 SNPs. It failed to assess whether one of the two alleles was consistently associated with a high value of a trait. This affected the results in Tables 1A and 1B, as it was not possible to know in which sort of cattle and under what circumstances one allele was better than another.
843 Further, Professor Goddard also noted that Tables 1A and 1B failed to report the effect sizes of the 2,510 SNPs, making it inappropriate to combine genotypes from multiple SNPs to predict the genetic value of an animal for a trait.
844 Further, Professor Goddard also considered that the 253 Application did not select enough SNPs to provide dense coverage of the bovine genome. In the absence of such dense coverage, one would not be able to obtain meaningful results that could be applied to bovine breeds not included in the Example 2 experiment. He considered that one would require greater than 300,000 SNPs in order to find SNPs that had consistent effects across breeds.
845 Further, Professor Goddard did not believe that a small number of SNPs would be useful for prediction of a trait, because each SNP may only predict a very small amount of genetic variance. For example, and as MLA submitted, Professor Goddard said that in relation to human height, 700 statistically significant markers explained only 16% of variation. Three markers would only explain 1%, a result that would be typical of “virtually every trait”. Professor Goddard also calculated that the predictive power of the 2,510 SNPs was 4%, and that assuming 500 SNPs are associated with each trait the predictive power of three SNPs was 0.024%, which was so low as to not be practically useful. And although Professor Goddard was aware of the study that identified a causative polymorphism of the DGAT1 gene that accounted for a large degree of genetic variance for fat content in milk, he considered such a finding to be very rare, especially for complex traits that rely upon a large number of genes of small effect.
846 Further, Professor Goddard considered that the 253 Application provided no guidance as to how SNPs other than the 2,510 would be useful for predicting a trait’s likelihood. He stated that the invention might have worked only if it was confined to the use of the 2,510 SNPs in respect of the five traits and in respect of a cattle population identical or very similar in breed composition to the population used in the 253 Application.
847 Professor Goddard considered that the specified SNPs would be little better than random SNPs for inferring bovine traits for the five disclosed traits. In respect of additional traits, he stated that the 2,510 SNPs were no better than random SNPs. In respect of the non-specified SNPs in limb (b), Professor Goddard considered that they would be no better than random SNPs for inferring any bovine trait. He stated that most SNPS within 500,000 bp of a specified SNP would be in low LD such that it would have little value as a surrogate for that SNP.
848 Further, Professor Goddard also considered it necessary that an association between a SNP and a trait be subjected to a validation study before concluding that it was useful for predicting genetic potential.
849 Professor Hayes stated that at a p value of 0.01, 305 results in the 253 Application would have been false positives. He said that this was simply a matter of statistics. He also recalled that in 2002, there were known problems with DNA pooling methods that could generate spurious results if the pools were not constructed well.
850 Further, Professor Hayes undertook work at the instruction of MLA’s solicitors. He recorded that he identified the location of position 300 for 2,447 of the 2,510 sequences. He could not identify unique locations in the bovine genome for the remaining 63 sequences. Of the 2,447 aligned sequences, Professor Hayes noted that 100 of them contained flanking sequences of between one and 20 nucleotides, i.e. the polymorphism did not occur at position 300. He said that this typically indicated that the polymorphism may have been an indel rather than a SNP. Of these 100 sequences:
(a) 57 had a polymorphism at either position 301 or 302. Professor Hayes said that an artefact may have been introduced by the way the relevant algorithm aligned the sequences, and that when the sequences were realigned, 54 of the 57 sequences aligned within one nucleotide of each other.
(b) For the 46 sequences where flanking sequences were more than one nucleotide apart, Professor Hayes considered that there was a high probability that these sequences had an indel at the relevant position. This was supported by his observation that many of these sequences contained multiple polymorphic nucleotides within 6 nucleotides of position 300, indicating a region with multiple insertions.
851 Further, Professor Hayes also took issue with the absence of a validation study in the 253 Application. He considered that SNPs would only be useful in a selection method once they were shown to be associated in both a discovery population and a further validation population. In the absence of validation, one could not assess which of the 2,510 SNPs would be useful in selecting an animal for a trait.
852 Further, regarding the utility of the limb (b) non-specified SNPs, Professor Hayes noted that the +/- 500,000 nucleotides encompassed 60.3% of the bovine genome. Given this wide coverage, he thought it was likely that some of these SNPs would be useful in inferring a bovine trait.
853 Professor Visscher stated that due to the way that the experiments were conducted in the 253 Application, the number of false positive results could be higher than 300. Further, he considered that the utility of the invention was undermined by the absence of a validation study, which would have sorted truly associated SNPs from false positives.
854 Professor Visscher raised several concerns in relation to utility. Despite the objective of the 253 Application being a method to make more accurate and earlier predictions, there was no evidence that the method using the 2,510 SNPs was better than or could make more accurate predictions than existing methods.
855 Further, Professor Visscher said that there was no disclosure of association between the 2,510 SNPs and any traits other than the five disclosed. Consequently, he was not confident that the SNPs could make useful predictions for a trait other than these five. Further, he said that any uncertainty about the predictive accuracy of the 2,510 SNPs for the five disclosed traits would only be amplified in relation to other traits.
856 Further, the 253 Application did not quantify the usefulness of any predictive accuracy of the method, such that Professor Visscher considered that the usefulness of the invention for the five disclosed traits could be very low or non-existent. Professor Visscher believed that three SNPs could not be a useful predictor if associated SNPs only accounted for a small amount of genetic variance.
857 Further, regarding the utility of limb (b) non-specified SNPs, Professor Visscher noted that the +/- 500,000 nucleotides covered most of the bovine genome. Given this wide coverage, he thought that the scope would capture most of the associations with trait loci.
858 Professor Plastow explained that as at December 2002, using the method of claim 1 he would have employed as many of the markers from the 2,510 SNPs for as many traits as he could, because at the time it was a “remarkable” jump forward. He would have done this even without knowing the size of the genetic effect of each of the markers.
859 Further, Professor Plastow also said that using the 2,510 SNPs he would have expected a “good chance of success”.
860 Professor Plastow considered that the specified 2,510 SNPs would be useful in identifying associations with bovine traits. In contrast to Professor Goddard, Professor Plastow thought that the SNPs disclosed in the 253 Application would be sufficiently robust to produce meaningful information from which trait associations could be inferred in other populations. In relation to limb (b) non-specified SNPs, he said that because the non-specified SNPs were to be in linkage disequilibrium with specified SNPs (his assumption concerning limb (b)), then they too would be useful in identifying associations with bovine traits. In relation to the necessity of conducting a validation study, Professor Plastow said he would expect that breeders would be able to calibrate the SNPs disclosed in the 253 Application against their own population of animals.
861 I would also note that Professor Plastow took issue with many of Professor Goddard’s propositions. For example, he said:
I do not share Professor Goddard’s belief that the 2,510 SNPs disclosed in the '253 Application would only work to identify cattle of superior genetic potential in respect of a population of cattle identical or very similar in breed composition to the ones used in the experiments described in the '253 Application. In my view, the series of SNPs disclosed in the '253 Application would be sufficiently robust to produce meaningful and useful information from which an inference of trait association may be made in other populations. As I have also discussed above, I understand that what the method of the '253 Application seeks to achieve is to provide SNPs close to causative mutations. In this way, the SNPs disclosed in the '253 Application provide a useful tool to, say, commercial breeders, who want to improve the genetic characteristics of their animals. I consider it would be reasonable to expect that breeders would be able to calibrate the SNPs disclosed in the '253 Application against their own population of animals and to select those with superior genetic potential. In my view, the process would be iterative, but that does not mean that the method disclosed in the '253 Application would not work as Professor Goddard states.
I disagree with Professor Goddard that the P-value selected was inappropriate. In my experience, a P-value of 0.01 is commonly used and considered perfectly acceptable as the benchmark by which the results obtained may be sorted. I do not consider that the occurrence of false positives, certainly at the low level asserted by Professor Goddard, would significantly undermine the usefulness of the 2,510 SNPs as suggested by Professor Goddard. I also disagree with Professor Goddard that it would take “a large amount of work” to identify which of the SNPs are false positives and which are not. As I have already indicated above, any “large amount of work” would have been carried out at the stage of identifying the disclosed SNPs and in determining that they can be used to infer a trait. Once provided with the disclosed SNPs, I do not consider that it would have been difficult or in any way burdensome to use those SNPs in accordance with the method claimed in the '253 Application.
Professor Goddard suggests that in order for “a useful estimate” based on genetic markers to be determined, it is necessary that the number of SNPs and the size of the population be “very large” – specifically, that it is “usual” to use >10,000 cattle and at least 300,000 markers. I completely disagree. As at December 2002, the number of SNP markers that had been identified was very limited, and indeed the 50K lllumina SNP chip did not become commercially available until 2008. Moreover, in my experience, even by today’s standards, a study involving 10,000 animals would be viewed as exceptionally large and certainly not usual. In my view, Professor Goddard has set a benchmark which even with the identified SNPs and SNP chip technology available today, the experimental design proposed by Professor Goddard would be extremely difficult to achieve. Professor Goddard’s views in relation to the stringency that ought to be applied in considering the method claimed by the '253 Application are also, in my opinion, contrary to his view (set out above in the First Goddard Affidavit) that it is possible to have as little as 20 markers to explain 50% of variation in traits. If this were the case, I cannot understand why Professor Goddard considers it necessary to use at least 300,000 markers in order to produce a useful or meaningful result.
Professor Goddard asserts that he is aware that a density of one SNP every 0.5 cM is not enough when studying breeds. In my view, this statement could not have been based on information known as at December 2002. I cannot see how Professor Goddard can state with such certainty, based on the information that was available in 2002, that the '253 Application does not disclose the selection of enough SNP markers to provide enough density of coverage of the bovine genome. As I have noted above, even the Meuwissen 2001 paper does not suggest that the density needs to be as great as Professor Goddard states is necessary.
For the reasons I have already explained above, I do not agree that the process of validating the SNPs disclosed in the '253 Application must involve the matters discussed by Professor Goddard. In my view, the process can be iterative and can involve sorting through SNPs that do not apply to your population (what Professor Goddard considers to be a false positive) from those that do. In my view, that process would provide robust results that enable the selection of animals based on their genetic potential.
Professor Goddard suggests that the LD of 500,000 nucleotides has resulted from “selective analysis”. I do not agree. The '253 Application clearly indicates that tests were carried out in order to determine the extent of LD, and the LD of 500,000 nucleotides is fully supported by the results reported. I cannot see any reason why the LD analysis would not be justified.
862 Dr Sonstegard acknowledged that statistically there would have been false positive results in the method of the 253 Application. He also acknowledged that a validation study was necessary in order to reach a definitive view of whether a set of markers was capable of accurate prediction.
863 Dr Sonstegard stated that the specified SNPs would be useful for inferring one of the five bovine traits identified. He also said that they would have some predictive value in respect of other traits based on the extent of LD in cattle genomes and the distribution of the specified SNPs across more than half the genome.
864 Dr Sonstegard said that non-specified SNPs would be useful for inferring one of the five bovine traits if you knew what they were. They would also be useful for inferring other bovine traits but that this would require testing.
865 Commenting on the 2009 study by Hayes and Sonstegard, Dr Sonstegard said that the conclusion of a validation study that 56 SNPs of the 125 initially thought to be associated were actually associated was “pretty good”. Further, he said that it was “pretty good too” when 20 of those SNPs were validated in a different bovine breed.
866 Professor Taylor considered that the 2,510 specified SNPs would be useful for inferring the five specified bovine traits. Further, he expected to find on average that 204 of those SNPs were associated with other bovine traits that possessed similar heritabilities to the five specific traits.
867 Professor Taylor considered that limb (b) SNPs would also be useful for inferring the five specific bovine traits. But he also said that not all SNPs within 500,000 nucleotides of the specified SNPs will be in LD with the QTL and that those SNPs not in LD with the QTL would not be predictive. This qualification also applied to inferring additional traits using the limb (b) SNPs.
868 Professor Taylor acknowledged the necessity of a validation study to identify issues around false positive results and a tendency in GWAS to overestimate the effect of SNPs.
(d) Analysis
869 Now there are some themes that can be distilled from the expert evidence, particularly MLA’s witnesses.
870 First, it seems to be said that many more SNPs than the number stipulated in the claims of the 253 Application would be necessary in order to establish the requisite association with a trait, and therefore for the invention to be useful.
871 Second, it seems to be said that absent a validation process, the invention would not be useful.
872 Third, it seems to be said that on any view some high statistically significant association between the 3 SNPs and the trait is necessary, and that the invention would not be useful absent this. Moreover, it seems to be said that there would be difficulties in identifying a limb (b) SNP, although in my view that would have been well within the skill of the calling.
873 Fourth, it seems to be said that even if a high statistically significant association was required by the claims, that in any event there would be a substantial number of false positives.
874 Fifth, it seems to be said that on any view the invention would only have utility for the 5 traits referred to in the specification.
875 Now many of these propositions were challenged by Branhaven’s experts. Given the conflict in the expert evidence and for other reasons, I am not clearly satisfied that the claims of the 253 Application lack utility in the dimensions asserted by MLA providing that amendments are made to deal with statistical significance and LD as I have indicated. Let me elaborate.
876 First, I agree with Branhaven that there is no promise in the specification that a definite degree of advantage will result from the use of the invention. MLA has referred to various passages in the specification, but these do not promise that any particular level of “accuracy” will be achieved by the use of the claimed method. Further, the specification must be construed as a whole and in context. Relevantly, [0101] cannot be taken out of context.
877 Second, if the amendments that I have suggested be made concerning “associated” and LD are made, then I agree with Branhaven that the claims of the 253 Application include (or will include), as limiting and characterising features, requirements that ensure the utility of the invention.
878 Third, matters such as the need to validate SNPs in the population of interest are qualifications and expedients of the kind readily left to the skilled reader to supply.
879 Fourth, I do not agree with MLA that many more SNPs than those specified in the claims would be necessary to make the invention work. Indeed, the specification and other evidence before me negates such a proposition. One SNP may be sufficient.
880 Fifth, as to traits other than the 5 stipulated traits, MLA has not adduced cogent evidence to support the proposition that the invention would not work for the other traits.
881 Sixth, if statistical significance is required (as I have said), the incidence of false positives does not prove a lack of utility. Any association established by statistical correlation will always have such an incidence. It is inherent in the very invention claimed.
882 In summary, in my view, MLA has not clearly established by evidence that the method would not achieve a useful result in any particular case. MLA has the onus which in my view was not discharged by its experts throwing up theoretical possibilities.
Lack of sufficiency
883 At the relevant time, s 40(2)(a) of the Act provided:
(2) A complete specification must:
(a) describe the invention fully, including the best method known to the applicant of performing the invention; and …
884 The “invention” for the purposes of s 40(2)(a) is the embodiment(s) which is described and around which the claims are drawn. Kimberly-Clark Australia Pty Ltd v Arico Trading International Pty Ltd (2001) 207 CLR 1 at [19] and [21] approved that formulation and its application to s 40(2)(a).
885 Only one embodiment within each claim need be enabled for sufficiency purposes. An invention is sufficiently disclosed if a skilled person could make a single embodiment of the invention which falls within the scope of the claims. The following test for sufficiency was formulated in Kimberly-Clark at [25]:
The question is, will the disclosure enable the addressee of the specification to produce something within each claim without new inventions or additions or prolonged study of matters presenting initial difficulty? (footnote omitted)
886 In Lockwood Security Products Pty Ltd v Doric Products Pty Ltd (No 1) (2004) 217 CLR 274 (Lockwood (No 1)), the High Court said at [60]:
For the purposes of s 40(2)(a), it is not necessary for the inventor to disclose all the alternative means; it is enough that there is disclosure in the sense of enabling the addressee of the specification to produce something within each claim without new inventions or additions or prolonged study of matters presenting additional difficulty. (footnote omitted)
887 In SNF (Australia) Pty Ltd v Ciba Speciality Chemicals Water Treatments Ltd at [234], Kenny J explained:
A specification is not insufficient merely because some experiment of a routine character (as distinct from prolonged study of matters presenting initial difficulty) is necessary in the particular case … Nor is a specification insufficient because it fails to give detailed instructions as to matters which a “practical person … would naturally settle, and expect to have to settle … himself”, provided he “would find no difficulty in so doing”. (citations omitted)
888 Further, the plurality in AstraZeneca AB v Apotex Pty Ltd (2014) 226 FCR 324 observed (at [205]):
although the complete specification must describe the invention fully, including the best method known to the inventor of performing the invention, it does not follow that the inventor must explain how he or she arrived at the invention. It is the invention itself that must be fully described, not the route that was travelled by the inventor to arrive at it. Again, the way in which the inventor came to the invention described and claimed may also have an evidentiary significance. However, an invention may be the result of chance or luck as much as long experiment and the question whether an invention, or an alleged invention, involves an inventive step is an objective one. (citation omitted)
(a) MLA’s contentions
889 MLA contends that the claims are insufficient because there is no clear teaching of how to produce something with each claim. In particular, it is said that the person skilled in the relevant art does not know which three SNPs to choose to be able to predict genetic potential. It is said that this was clear from Professor Plastow’s evidence about how he would put claim 1 into practice.
890 Further, MLA contends that where (as here) a claim in effect claims alternative methods, the specification must describe how to perform the invention for each such method, not just one method within the claim. It contends that a patentee should not be permitted to avoid a finding of lack of sufficiency by drafting a claim to cover two or more alternative methods and then only providing a sufficient description as to one. In support of this contention it cites Tramanco Pty Ltd v BPW Transpec Pty Ltd (2014) 105 IPR 18; [2014] FCAFC 23 at [206] to [208], per Nicholas J (referred to by me in GlaxoSmithKline Consumer Healthcare Investments (Ireland) (No 2) Ltd v Apotex Pty Ltd (2016) 119 IPR 1; [2016] FCA 608 at [693]). Nicholas J said at [207] (Allsop CJ and Greenwood J agreeing):
In Kimberly-Clark and Doric the High Court was concerned with various product claims each of which consisted of a number of features all of which were essential (in the sense of non-optional) integers of the claim. However, there are some types of claim that may need to be approached slightly differently including, in particular, claims to methods for producing one or more specified results. For example, a claim for a method of producing one or more of outcomes A, B or C might be infringed if the alleged infringer uses the method to produce outcome A, but not outcome B or C. Whether there is infringement in such a case will depend upon (inter alia) the proper construction of the claim and, in particular, whether it requires the use of the method to produce only one or more of outcomes A, B or C, as opposed to all three of them. Assuming the former construction (in the present case it appears to have been assumed by both parties at the trial and on appeal that claim 1 is to be construed in this way), it would seem to me to be wrong in principle to hold that the description of the invention is sufficient if the specification enables the use of the method to achieve outcome A, but not outcomes B or C. It would be inconsistent with the purposes of the Act to confer a monopoly on a patentee for a method of producing any of outcomes A, B or C, if the patentee’s disclosure only enabled the use of the method to produce some of those outcomes.
891 Further, MLA pointed out that in Apotex Pty Ltd v Warner-Lambert Company LLC (No 2) (2016) 122 IPR 17; [2016] FCA 1238 at [244], Nicholas J reinforced his observation that the Kimberly-Clark test for sufficiency may not be able to be applied to all forms of claim and said (at [245] to [247]):
Tramanco was not concerned with methods of medical treatment, but with a method for determining one or more parameters relating to the performance of the suspension system of a vehicle. In the present case the method claims, as I have construed them, are to methods of treating pain, or different types of pain, in all kinds of mammals including, in particular, humans.
As we are reminded from time to time, judgments are not to be construed as if they were statutes, and it is always necessary to read and understand them in context. Application of the test of insufficiency propounded by the High Court in Kimberly-Clark based upon a test originally formulated outside the context of methods of treatment claims cannot be sensibly applied to a case such as this. In my respectful opinion, it would be a nonsense to say that the invention was sufficiently described if it enabled the skilled addressee to perform the method of the invention on rats (or other mammals) but did not enable it to be performed on humans.
In my view, s 40(2)(a) requires that the Patent in this case enable the skilled addressee to perform the claimed invention in relation to humans without new inventions or additions or prolonged study of matters presenting initial difficulty. It is only by requiring this degree of enablement that the patentee could be sensibly understood to have given consideration for the grant of a patent which the skilled addressee would understand to be essentially directed to the treatment of pain in humans.
892 Now MLA has relied heavily upon his Honour’s observations. But as appropriate insurance, MLA has also reserved the right to argue in a suitable forum that Kimberly-Clark is wrong if I consider that contrary to what Nicholas J has said, Kimberly-Clark can not be so distinguished.
893 MLA says that in the present case, claim 1 specifies one method that requires one of the at least three SNPs to be a specified SNP, and another method that requires one of the at least three SNPs to be within 500,000 nucleotides of a specified SNP. Accordingly, MLA says that even if limb (a) is sufficiently described, which in any event it does not accept, that cannot provide an answer to limb (b) of claim 1. It says that its contention should apply, by parity of reasoning, to different traits. That is, the claims describe a different method for each trait to be identified or inferred. Therefore each method must be sufficiently described for each such trait.
894 Further, MLA says that the specification does not describe the following matters, which are required to perform the invention when construed as being to distinct methods. And the evidence shows that determining each of these matters would have required undue experimentation as at the priority date. It has essentially made four points.
895 First, MLA says that none of the specified SNPs has been shown in the 253 Application to occur in a gene. Accordingly, the person skilled in the art would need to work out if one of the specified SNPs, or a non-specified SNP, was in a gene. Further, given that the majority of genes in the bovine genome had not been identified by December 2002, this would have required significant experimentation. And even using comparative mapping with the human genome, the skilled addressee could not know whether a SNP was within a gene in the bovine genome where there was no equivalent gene in the human genome, as Professor Goddard explained. Further, Professor Taylor’s evidence was that this exercise would not necessarily be trivial. Further, as Professor Hayes explained, many of the specified SNPs do not occur in genes.
896 Second, MLA says that none of the specified SNPs was validated in any animals and many will not have any predictive value at all. Further, not all of the specified SNPs were shown to be associated with each of the five traits. Only about 600 were identified as associated for each trait. Further, even then, a mere association is not sufficient to be predictive of genetic potential with any degree of accuracy according to the evidence of Professor Goddard and Professor Visscher. Further, it was submitted that Professor Plastow accepted that the SNPs need to be validated in a different population as having the relevant association. Further, each of Professor Taylor and Dr Sonstegard also agreed that markers need to be validated in another population to be useful in that population.
897 MLA points out that the 253 Application does not validate any SNP / trait association, leaving it to the skilled addressee to do so. But it is said that such work would be time consuming and costly. Professor Goddard said that the amount of work required is comparable to starting from scratch to do a genomic selection project. It involves genotyping and phenotyping work. Further, MLA points to the fact that Professor Hayes was directly involved in the validation work in the study reported in Hayes BJ, Bowman PJ, Chamberlain AJ, Savin K, van Tassell CP, Sonstegard TS and Goddard ME, “A Validated Genome Wide Association Study to Breed Cattle Adapted to an Environment Altered by Climate Change” (2009) 4(8) PLoS ONE e6676. He said that it was a significant amount of work involving 12 to 18 months to run an analysis, and 12 months to extract the DNA and to carry out genotyping. MLA points out that this was carried out in 2007, and that importantly Professor Hayes was able to use a SNP chip. But without access to a SNP chip, Professor Hayes described a genotyping process involving only 6 to 8 SNPs as “just a pain”. I would just note at this point that this study tested 152 SNPs in the validation data sets.
898 MLA says that it is no answer to say that any use of the method will require validation. MLA says that if the inventors had, as they should have done, validated the associations for the specified SNPs in a working population of cattle, then the SNPs would have a much higher prospect of being associated with the trait of interest in any population.
899 Third, MLA says that none of the specified SNPs was tested for association with any trait other than the five traits tested. Accordingly, to the extent that the claims extend to other traits, they are not sufficiently described. It is said that that is because the skilled addressee would have to conduct his own association studies involving genotyping and phenotyping significant numbers of animals for the SNPs and trait of interest. That would effectively involve repeating the inventors’ work which required undue experimentation in terms of time and cost.
900 Further, MLA says that if the skilled addressee wanted to use one of the specified SNPs or a non-specified SNP for any trait other than the five traits identified in the specification, he would need to re-do Example 2 of the 253 Application. This would involve measuring phenotypes and genotypes for all SNPs to be used with more than 4,000 cattle. This would be a major undertaking that could take one or two years and cost millions of dollars. And after all that, it could show that many SNPs are not valid.
901 Fourth, MLA says that no non-specified SNPs (that is, SNPs within +/- about 500,000 nucleotides of a specified SNP) were identified as being associated with any trait. In this regard, it says that Branhaven accepted that it was very difficult and time consuming to identify a sufficiently large number of SNPs and identify associations between those SNPs and traits. MLA says that the same must apply to any non-specified SNPs. Furthermore, even the task of determining whether a non-specified SNP is in fact within +/-about 500,000 nucleotides of a specified SNP would be extraordinarily time consuming. MLA says that it effectively required the entire bovine genome to be known, as Professor Goddard and Professor Visscher suggested.
902 MLA also referred to Gilead Sciences Pty Ltd v Idenix Pharmaceuticals LLC, where Jagot J found certain claims lacked sufficiency and accepted Gilead’s submission (at [624]) that:
… a screening process of any significant number of nucleosides – which of course require synthesising before screening – is a research project akin to those in which Biota and Avexa were engaged and exceeds the level of work permitted for a patent to comply with s 40(2)(a).
903 MLA says that the same applies here, as the skilled addressee would have to repeat the work done by the inventors. I would note that her Honour’s decision was upheld on appeal (see Idenix Pharmaceuticals LLC v Gilead Sciences Pty Ltd [2017] FCAFC 196).
904 Further, MLA says that Branhaven, through its patent attorneys, has admitted that undue experimentation or invention was required to identify SNPs and determine their association with a specific trait. MLA relies upon the letter from Freehills Patent Attorneys to the Commissioner of Patents dated 11 October 2013 and in particular point 4. MLA says that Branhaven cannot have it both ways. If invention was required to identify relevant associated SNPs, then there was insufficient information in the specification for a person skilled in the relevant art to identify SNPs other than the specified SNPs (limb (a) of claim 1).
905 Generally, and applying the principle in Kimberly-Clark at [25], MLA says that the specification does not:
enable the addressee of the specification to produce something within each claim without new inventions or additions or prolonged study of matters presenting initial difficulty. (citation omitted)
(b) Analysis
906 Now as a preliminary matter, I would observe that the observations in Tramanco Pty Ltd v BPW Transpec Pty Ltd at [207] and Apotex Pty Ltd v Warner-Lambert Company LLC (No 2) at [244] have little application to the present case. The claim under consideration in Tramanco at [207] was in form and in substance a claim to a method for producing alternative results or outcomes. It is apparent that his Honour regarded the claim in Apotex v Warner-Lambert in similar terms. But claim 1 of the 253 Application is not a claim to such a method. It is not a claim to a method for producing alternative results or outcomes. Rather it is a claim to a general method that may be employed using any combination of features within the scope of the claim.
907 Further, I agree with Branhaven that MLA’s contentions fail to apply the Kimberly-Clark test. The skilled person could readily put the invention into practice, by performing an embodiment of the invention within the scope of each claim. Moreover, this could be done without undue experimentation. And this is all that is required. The evidence does not establish that the skilled person could not use an embodiment of the method within claim 1 of the 253 Application without undue experimentation. For example, he could take any three or more of the 2,510 specified SNPs and use them in the method of the invention to infer the potential for a trait with which they were shown to be associated. In my opinion, the evidence does not clearly establish otherwise.
908 Now MLA has pointed to four matters in respect of which it is said that the description in the specification is deficient. But none of these matters supports any finding of a lack of sufficiency.
909 First, MLA says that the skilled person would need to work out if the SNPs proposed to be used in the claimed method were in genes. But this is something that the skilled person could readily do, as Professor Plastow and Dr Sonstegard explained. At the priority date this could be done by comparing sequence homology with publicly available information concerning the human genome. And from the time of filing of the 253 Application, the bovine genome could be used as the reference point. The location of a SNP within a gene is for molecular geneticists well within the skill of the calling. Moreover, the difficulties and uncertainties said by MLA to be associated with this process are overstated. As a skilled person could employ a method within claim 1 using any three or more SNPs that meet the requirements of the claim, it is not the case that large numbers of SNPs would need to be assessed and located within genes in order to apply the invention. Moreover, it would have been well within the skill of the calling to identify a limb (b) SNP.
910 Second, although MLA says that the SNPs would need to be validated, this is something that the skilled person could readily do. As some of the evidence established, validation is something that would be done as a matter of course in any case where the population of interest is genetically different to the research population. Moreover, I agree with Branhaven that it is not to the point that the 253 Application does not show that the 2,510 specified SNPs were validated in a population other than the research population. There would have been little point to doing this for the purposes of the 253 Application. As a practical matter, the SNPs would have needed to be validated again in any event by the skilled person seeking to practise the invention in his population of interest. It would only be where the population of interest happened to be the same as or genetically similar to the validation population (if there had been validation) in the 253 Application that this could be avoided. Further, in my view the work involved in validating SNPs in a population of interest is routine, and not undue experimentation. I accept Branhaven’s submission that it is part of putting the method into effect and would be done as a matter of course.
911 Third, although MLA submits that it would be necessary to re-do at least example 2 for traits other than the five traits the subject of the association study in the 253 Application, such a proposition does not properly apply Kimberly-Clark. The method of claim 1 is not specific as to the trait of interest. It extends to any trait. I agree with Branhaven that enablement in relation to any one or more of the five identified traits is sufficient. Moreover, MLA’s characterisation of the work involved in practising the invention in relation to other traits is debatable. As Branhaven says, it would not be necessary to “re-do” example 2 of the 253 Application for this purpose. Example 2 was part of the inventors’ GWAS approach that resulted in the development of the panel of 2,510 SNPs associated with a variety of traits. The invention could be put into practice with far fewer SNPs. The method of claim 1 requires only three or more SNPs associated with a trait of interest. I am not satisfied that the skilled person could not identify a sufficient number of SNPs and their association with a trait in order to use the claimed method.
912 Fourth, MLA says that an association study would need to be conducted for non- specified SNPs to identify their association with a trait. But this again fails to apply Kimberly-Clark.
913 Finally, let me briefly deal with two other matters. First, the decision of Jagot J in Gilead Sciences Pty Ltd v Idenix Pharmaceuticals LLC does not assist MLA. Her Honour’s acceptance that a “research project” would be required was quite a different context. But in the present case, 2,510 SNPs have been identified as being associated with one or more of five traits. The skilled person can elect to use them in the claimed method. It is a matter for the skilled person if he or she chooses instead to use other SNPs to carry out the method. The fact that this might involve some additional time and effort does not establish insufficiency. Second, MLA’s reliance on correspondence from Branhaven’s previous attorneys does not carry the matter far. It adds little to my assessment of whether the claimed invention has been sufficiently described to refer to representations made during the course of prosecution. Further, the letter does not in terms admit that, having been provided with the information in the 253 Application, the skilled person would be unable to validate the specified SNPs without undue experimentation.
914 I am not clearly satisfied that MLA’s insufficiency ground has been made out.
Lack of fair basis
915 Section 40(3) requires a real and reasonably clear disclosure in the body of the specification of what is claimed. The language of s 40(3) points to a comparison between the claims and what is described in the body of the specification only. As stated by Barwick CJ in Olin Corporation v Super Cartridge Co Pty Ltd (1977) 180 CLR 236 at 240:
the question is a narrow one, namely whether the claim to the product being new, useful and inventive, that is to say, the claim as expressed, travels beyond the matter disclosed in the specification.
916 Section 40(3) does not use the word “invention”, but it requires that the claims “be fairly based on the matter in it that discusses the ‘invention’”; Lockwood (No 1) at [53]. This is the embodiment(s) which is described and around which the claims are drawn, but it does not mean the inventive step taken by the inventor or the advance in the art made by the inventor. Lockwood (No 1) described the relevant test in the following terms (at [68] and [69]):
Erroneous principles. The comparison which s 40(3) calls for is not analogous to that between a claim and an alleged anticipation or infringement. It is wrong to employ “an over meticulous verbal analysis”. It is wrong to seek to isolate in the body of the specification “essential integers” or “essential features” of an alleged invention and to ask whether they correspond with the essential integers of the claim in question.
“Real and reasonably clear disclosure”. Section 40(3) requires, in Fullagar J’s words, “a real and reasonably clear disclosure”. But those words, when used in connection with s 40(3), do not limit disclosures to preferred embodiments.
The circumstance that something is a requirement for the best method of performing an invention does not make it necessarily a requirement for all claims; likewise, the circumstance that material is part of the description of the invention does not mean that it must be included as an integer of each claim. Rather, the question is whether there is a real and reasonably clear disclosure in the body of the specification of what is then claimed, so that the alleged invention as claimed is broadly, that is to say in a general sense, described in the body of the specification.
Fullagar J’s phrase serves the function of compelling attention to the construction of the specification as a whole, putting aside particular parts which, although in isolation they might appear to point against the “real” disclosure, are in truth only loose or stray remarks. (footnotes omitted, save that the italicised phrase is drawn from Rehm Pty Ltd v Websters Security Systems (International) Pty Ltd (1988) 81 ALR 79 at 95 per Gummow J)
917 So, one should not use an “over meticulous verbal analysis”. Further, the focus is not on an identity of language between the claims and the disclosure in the body of the specification. Rather, one is looking for a generalised disclosure in the body that provides support for the claims in substance. Moreover, it is inappropriate to isolate in the body of the specification essential integers or features of an alleged invention and to ask whether they correspond with the essential integers of the claim.
918 Now fair basis can be established by a comparison with the consistory clause(s). But even if a claim is based on and mirrors the form of the consistory clause(s), it will not be fairly based if other parts of the specification show that the invention is narrower than the consistory clause(s). What has been described as a “coincidence of language” between a claim and part of the body of a specification does not per se establish fair basing if that part of the language of the specification does not reflect the description of the invention in the light of the specification as a whole.
919 Further, it is appropriate to restate that the complete specification is not to be read in the abstract, but is to be construed in the light of common general knowledge and the relevant art before the priority date.
920 Further, where a feature included in a claim is a limiting feature, there is no need for it to be the subject of an explicit disclosure in the body of the specification, if the subject matter of the claim falls within the scope of what is more broadly described in the specification. As explained in DSI Australia (Holdings) Pty Ltd v Garford Pty Ltd (2013) 100 IPR 19; [2013] FCA 132 at [240] per Yates J:
… the inquiry as to fair basis is directed to the question of claim width: see, for example, Olin Corporation at CLR 240 ... A claim may be fairly based for the purposes of s 40(3) of the Act where it adds a feature to a combination otherwise described in the specification and, by that addition, limits the described invention, as a matter of definition, to a more restrictive form than that to which the patentee might otherwise be entitled. In short, a claim may be fairly based for the purposes of s 40(3) of the Act even when all the characteristics by which the invention is defined in the claim are not described in the body of the specification itself, provided those characteristics are truly limiting ones in the sense that I have described.
921 Relatedly, the claims need not be restricted to precise embodiments described in the specification. As Gummow J said in Sartas No 1 Pty Ltd v Koukourou & Partners Pty Ltd (1994) 30 IPR 479; [1994] FCA 936 at 497:
it is no objection to any particular claim that it claims a monopoly for less than every feature described in the body of the specification. It cannot be the case that, for example, a claim is restricted to the precise embodiment which is depicted in the body of the specification.
(a) MLA’s arguments
922 MLA says that the claims essentially include methods of using any three SNPs from anywhere in the bovine genome that are associated with any trait to identify cattle with that trait. The only restrictions are that at least one of the SNPs is a specified SNP or a non-specified SNP i.e. within +/- about 500,000 nucleotides of a specified SNP, which MLA says covers two thirds of the bovine genome, and at least two SNPs occur on different genes. MLA says that the claims are extraordinarily wide and do not reflect what is described as the invention in the specification. MLA says that they travel beyond the matter described in the specification in at least the following two ways.
923 First, MLA says that there is no basis for a claim to any of the specified SNPs being associated with any trait other than those five traits examined in the association study, which are all carcass traits. It says that the 253 Application does not provide any basis for considering that the specified SNPs are associated with anything other than the five identified traits. Accordingly, insofar as the claims include SNPs that are associated with traits other than those five carcass traits, for example, disease resistance, they travel beyond the invention described in the specification.
924 Second, MLA says that there is no basis for any SNP other than those identified as associated with one of the five traits. It asserts that no association with any trait is shown with any non-specified SNP. The SNPs identified in Example 3 of the 253 Application concern specified SNPs. Further all of the SNPs in use in the LD analysis, and shown to be associated, are one of the specified SNPs. Further, in practice, the +/- about 500,000 nucleotide limitation is not a real limitation, as it covers more than two thirds of the bovine genome. MLA says that there is no basis for the claims to include any SNP that is not a specified SNP. Further, MLA asserts that Branhaven identified that the invention is in the panel of SNPs provided. MLA says that Branhaven’s experts also understood the invention as being to a subset of SNPs. MLA says that it is apparent from Professor Plastow’s characterisation of the 2,510 specified SNPs as a “windfall” and a “remarkable jump” that this is where he considered the true value of the invention to lie. Accordingly, MLA says that the claims lack fair basis in that they travel well beyond the panel of SNPs identified in the specification.
(b) Analysis
925 I would reject MLA’s fair basis attack.
926 First, although MLA contends that there is no basis for a claim to any of the specified SNPs being associated with any trait other than those five traits examined in the association study, I agree with Branhaven that this misapplies the principles of fair basis.
927 Claim 1 is commensurate with the description in the body of the specification. As Branhaven points out, the broad form of the invention described at [0027] is not limited to the use of any particular SNP associated with any particular trait. And I also agree that the embodiment described at [0030] is also not so limited. Further, later embodiments are similarly broadly expressed. And as for paragraphs of the specification that refer to particular traits, including the five traits that were the subject of the association study, these are expressed to be non-limiting, preferred embodiments of the invention: see [0027] and [0090].
928 In my view such disclosures provide ample fair basis for other traits. Further, I agree with Branhaven that there is no need for the description to establish, by experiment or otherwise, that any of the specified SNPs is in fact associated with any other trait. No authority discussing fair basis and binding upon me supports any such requirement.
929 Further, MLA’s argument focuses on particular integers of claim 1, not the claim as a whole. It is the claim that must be fairly based. The claim is to a method for identifying a trait in a bovine subject that comprises, relevantly, identifying in a nucleic acid sample of the subject at least three SNPs associated with the trait. The claim does not require that any of the 2,510 specified SNPs will necessarily be associated with any particular trait whether within or outside the five traits the subject of the association study. They may or may not be. Moreover, the evidence does not establish that the 2,510 specified SNPs would not be useful to infer an association with other traits outside the five traits.
930 Second, as to MLA’s contention that there is no basis for a claim to any SNP other than the 2,510 specified SNPs, I would also reject such a contention. There is support in the specification for a claim to a method that involves the use of SNPs other than the 2,510 specified SNPs; see, for example, [0030], [0035] and [0126] in the detailed description. The method described is not limited to the specified SNPs. The identified SNPs are examples of SNPs provided that can be used in the method. But the description teaches that the method can also be employed using other SNPs, provided that they satisfy the elements of claim 1, including being associated with a trait of interest, and for a limb (b) SNP being in relevant linkage disequilibrium (as I have explained elsewhere and as I will require by way of amendment) with the identified limb (a) SNP.
931 I reject MLA’s fair basis attack.
Lack of clarity and definition
932 A valid claim is required to define with sufficient certainty the scope of the monopoly being claimed (s 40(3)). Given that a patent is a public instrument, the claim must be defined in such a way that it is not reasonably capable of being misunderstood so that others know the “exact boundaries of the area within which they will be trespassers”: Electric & Musical Industries Ld v Lissen Ld (1939) 56 RPC 23 at 39 per Lord Russell of Killowen. A claim will lack clarity if a person skilled in the relevant art cannot ascertain whether what he proposes to do falls within the claim’s ambit.
933 But lack of precise definition will not be fatal to the validity of a claim as long as it provides a workable standard suitable to the intended use. But as stated in Kauzal v Lee (1936) 58 CLR 670 at 685 per Dixon and McTiernan JJ:
[v]agueness of description, want of particularity and evident indistinctiveness of thought may be the source of so much uncertainty as to the scope of the monopoly that the claim fails to fulfil the requirement of stating with definiteness to what the patentee is exclusively entitled.
934 A claim is clear if either there is no ambiguity or any ambiguity is resolvable by properly construing the claim applying the principles that I have set out previously. But I accept that a claim is bad if no reasonably certain construction can be given to it. And I also accept that I am not bound to find a meaning for a claim nor to approach a claim with the conviction that its language is capable of a reasonable construction when carefully examined.
935 In Flexible Steel Lacing Co v Beltreco Ltd (2000) 49 IPR 331; [2000] FCA 890, Hely J held that each of the method and product claims for pulling lagging used in conveyor systems was invalid for lack of clarity. In respect of the method claim, he said at [107]:
The method claim is fairly open to more than one meaning not because of grammatical problems but because, even to a skilled reader, it would not be clear which of two methods claim 13 describes.
936 In respect of the product claim, he held at [131]:
Thus, I conclude that the product claim is obscure; it is fairly and equally open to diverse meanings, namely that the sipes run at right angles across the strip, on the one hand, or that the sipes run along the length of the strip on the other. Another possibility is that the claim embraces both. Sometimes, ambiguity or insufficiency in description can be resolved by a skilled addressee through the application of commonsense and common knowledge: cf Innovative Agriculture Products Pty Ltd v Cranshaw (1996) 35 IPR 643 at 666. I do not think that this is such a case.
937 Finally, if a claim is clear, it is not to be made obscure or treated as obscure by taking elements of a preferred embodiment not referred to in the claim and artificially creating obscurity.
938 Let me turn to the question of definition.
939 Section 40(2)(b) requires the complete specification “to end with a claim or claims defining the invention”. In General Tire & Rubber Co v Firestone Tyre and Rubber Co Ltd (1971) 1A IPR 121, the Court stated at 167 that “the issue of definition is to be considered as a practical matter and little weight is to be given to puzzles set out at the edge of the claim which would not as a practical matter cause difficulty to a manufacturer wishing to satisfy himself that he is not infringing the patent”. The Court also observed that allowances should be made for any difficulties of the case, so that an alleged issue of want of definition should always be considered in relation to the particular facts. It concluded (at 167 and 168):
It is clear in our judgment that the question whether the patentee has sufficiently defined the scope of his claims is to be considered in relation to the facts of each case, that allowance is to be made for any difficulties to which the circumstances give rise, and that all that is required of the patentee is to give as clear a definition as the subject matter admits of. It is also clear in our judgment that, while the court is to have regard to all the relevant facts, the issue of definition is to be considered as a practical matter and little weight is to be given to puzzles set out at the edge of the claim which would not as a practical matter cause difficulty to a manufacturer wishing to satisfy himself that he is not infringing the patent. We accept also that definition of the scope of a claim is not necessarily insufficient because cases may arise in which it is difficult to decide whether there has been infringement or not provided the question can be formulated which the court has to answer in the issue of infringement.
940 A claim will be bad if it fails to define the monopoly claimed so that the skilled addressee of the patent can know the exact boundaries of the area within which they will be trespassers.
941 MLA contends that there is no basis for a claim to any of the specified SNPs being associated with any trait other than those five traits examined in the association study, which are all carcass traits. The 253 Application does not provide any basis for considering that the specified SNPs are associated with anything other than the five identified traits. Insofar as the claims include SNPs that are associated with traits other than those five carcass traits, for example, disease resistance, they travel beyond the invention described in the specification.
942 Further, MLA contends that there is no basis for any SNP other than those identified as associated with one of the five traits. No association with any trait is shown with any non-specified SNP. In practice, the +/- about 500,000 nucleotide limitation is not a real limitation, as it covers more than two thirds of the bovine genome. There is no basis at all for the claims to include any SNP that is not a specified SNP. At the priority date, many such SNPs were yet to be discovered (but they have since been discovered).
943 MLA says that as a consequence of the above two matters, each of the claims fails to define the invention described in the specification. As is apparent from my reasons I would reject these contentions.
944 In terms of lack of clarity and for the reasons that I have already expressed in the construction section, claim 1 (and analogous claims) will require amendment to:
(a) define “associated” in terms of statistical significance at the p value of equal to or less than 0.01 (or such other measure as I decide after hearing from counsel further);
(b) require each of the 3 SNPs to satisfy that level of statistical significance (to be discussed further with counsel); and
(c) require the limb (b) SNP to be in LD with the relevant limb (a) SNP and to the requisite degree (to be discussed further with counsel).
945 As presently formulated, claim 1 and analogous claims fail for lack of clarity and proper definition, and also give rise to aspects of inutility. But if such amendments are made, these matters may be rectified.
946 I should note that I have not made any final decision on whether amendment will be permitted and if so its form. I will hear further from the parties on that aspect.
Conclusion
947 As I have indicated in these reasons, aspects of the claims in their present form are deficient in terms of:
(a) lack of clarity;
(b) a failure to define the invention ; and
(c) related to some questions of construction, lack of utility.
948 Accordingly I would uphold MLA’s appeal to this extent. But otherwise I would dismiss MLA’s appeal. I will not, however, make any final orders until Branhaven has been given the opportunity to consider whether to apply to amend any of the claims to address the concerns that I have expressed in these reasons.
949 I will hear further from counsel as to the course that they suggest should be taken and the orders that they suggest should be made consequential upon these reasons.
I certify that the preceding nine hundred and forty-nine (949) numbered paragraphs are a true copy of the Reasons for Judgment herein of the Honourable Justice Beach. |
Associate:




